Search Count: 305
All
Selected
 |
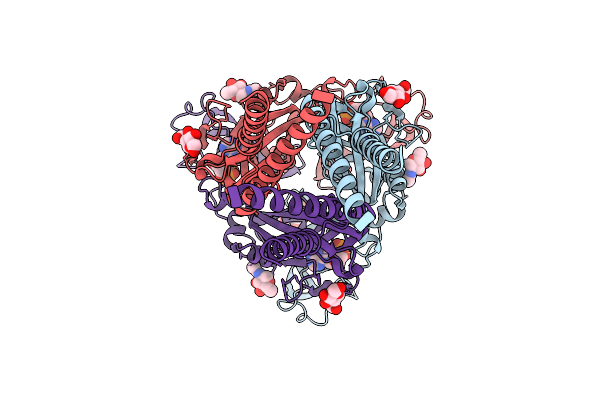
Cryo-Em Structure Of Integrin Alpha V Beta 6 Complex With A Bicyclic Inhibitory Peptide
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CELL ADHESION Ligands: CA, NAG, MAN, MN |
 |
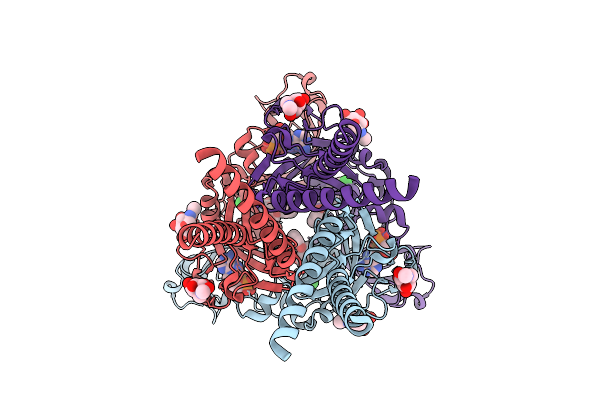
Cryo-Em Structure Of The Human P2X3 Receptor In The Atp-Bound, Desensitized State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: ATP, NAG |
 |
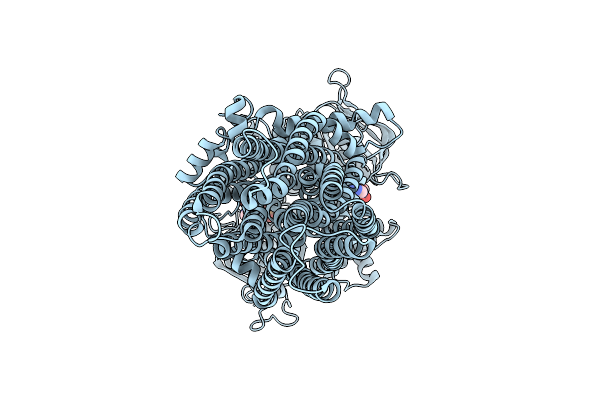
Cryo-Em Structure Of The Human P2X3 Receptor In The Atp- And Sivopixant-Bound Closed State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: A1E2N, NAG, ATP |
 |
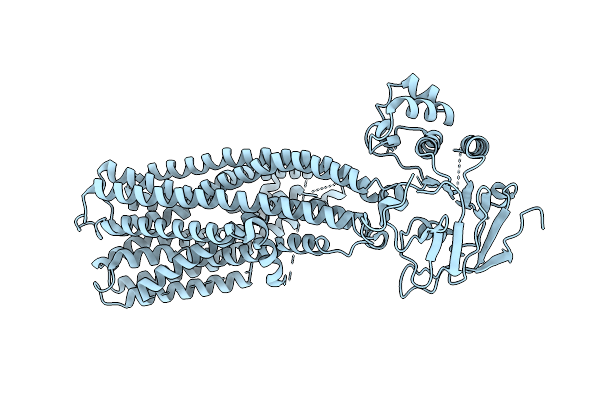
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Resolution:2.63 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: ATP, MG |
 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Resolution:3.21 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN |
 |
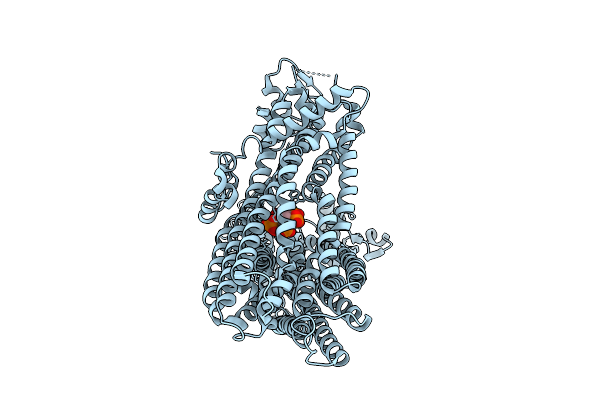
Cryo-Em Structure Of Rice Multidrug Resistance Protein 5 (Mrp5) With Insp6 In State A
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: IHP |
 |
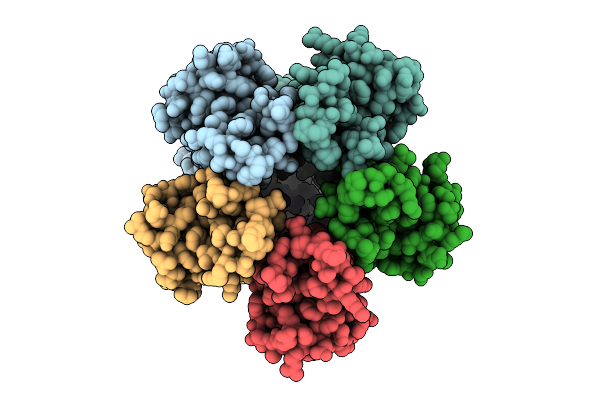
Cryo-Em Structure Of Rice Multidrug Resistance Protein 5 (Mrp5) With Insp6 In State B
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: IHP |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.92 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
 |
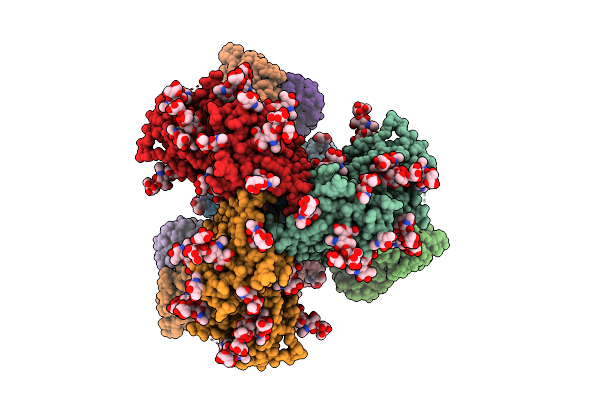
Cryo-Em Structure Of J601-1B2 Fab In Complex With Hiv-1 Bg505 Ds-Sosip Env Trimer
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, MAN |
 |
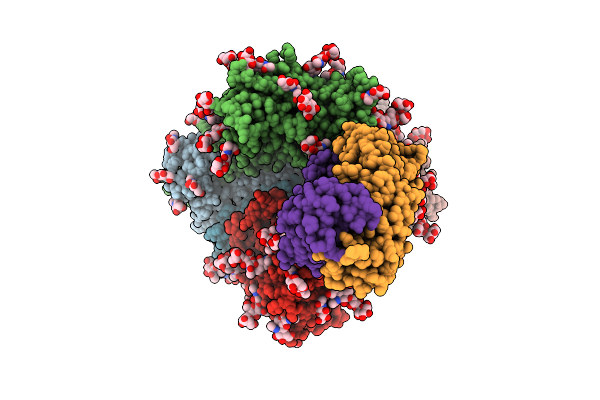
Cryo-Em Structure Of J601-A6 Fab In Complex With Hiv-1 Bg505 Ds-Sosip Env Trimer
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
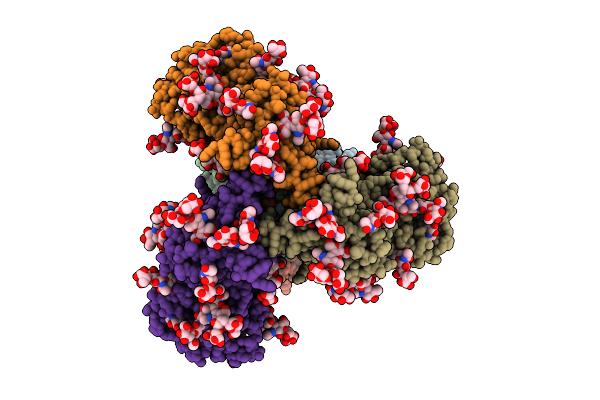
Cryo-Em Structure Of K001-A1 Fab In Complex With Hiv-1 459C-Opt Rns Ds-Sosip Env Trimer
Organism: Macaca mulatta, Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
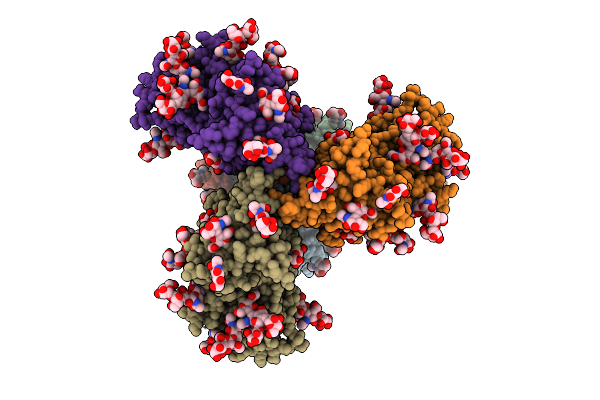
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
 |
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Resolution:3.52 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
 |
Organism: Drosophila melanogaster, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: ABU |
 |
Organism: Drosophila melanogaster
Method: ELECTRON MICROSCOPY Resolution:3.61 Å Release Date: 2025-10-01 Classification: MEMBRANE PROTEIN Ligands: 2BV |
 |
Organism: Human enterovirus d68
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: VIRUS LIKE PARTICLE |
 |
Cryo-Em Structure Of Ev-D68 Vaccine Candidate - A2 Subclade Virus-Like Particle
Organism: Human enterovirus d68
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: VIRUS LIKE PARTICLE |
 |
Organism: Human enterovirus d68
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: VIRUS LIKE PARTICLE |

