Search Count: 100
All
Selected
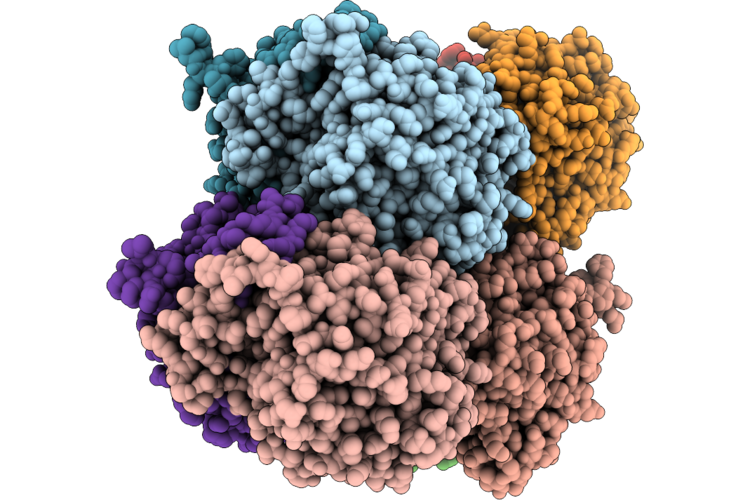 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-29 Classification: LIPID BINDING PROTEIN |
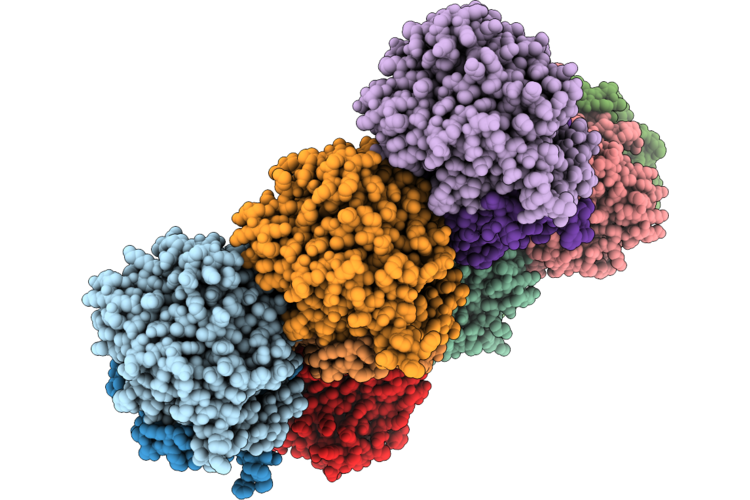 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.76 Å Release Date: 2026-04-29 Classification: LIPID BINDING PROTEIN Ligands: 4BW |
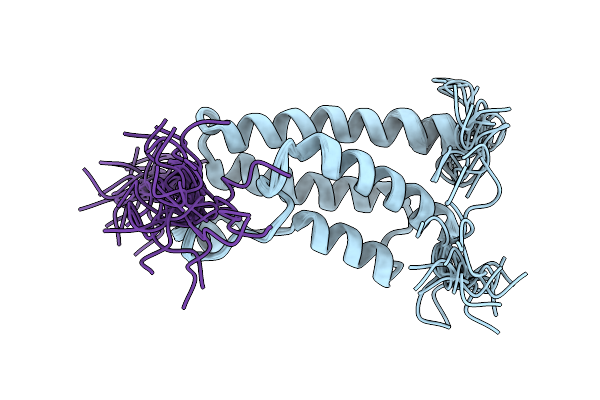 |
Solution Structures Of Brd9 Bromodomain In Complex With Histone H3 Acetyl-Lysine 18 (H3K18Ac) Peptide
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-03-11 Classification: PROTEIN BINDING |
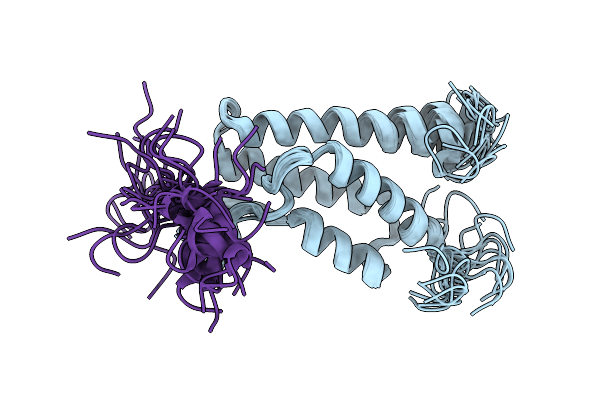 |
Solution Structures Of Brd9 Bromodomain In Complex With Histone H3 Lactyl-Lysine 18 (H3K18La) Peptide
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-03-11 Classification: PROTEIN BINDING |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: HYDROLASE Ligands: 4BW, PEE |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: HYDROLASE Ligands: 4BW |
 |
Organism: Epilithonimonas lactis
Method: ELECTRON MICROSCOPY Resolution:2.88 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: C2E, CA |
 |
Organism: Epilithonimonas lactis
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: C2E |
 |
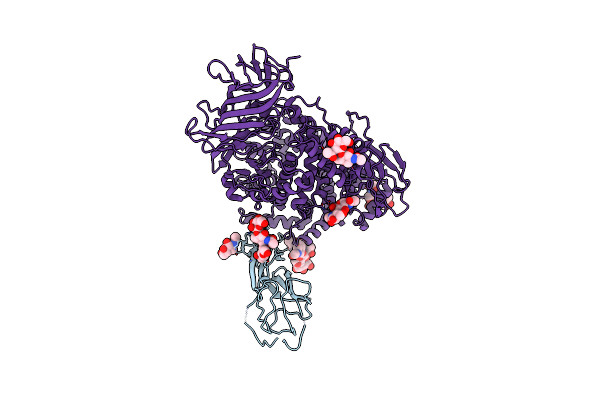
Organism: Porcine deltacoronavirus, Canis lupus familiaris
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: ISOMERASE Ligands: NAG, ZN |
 |
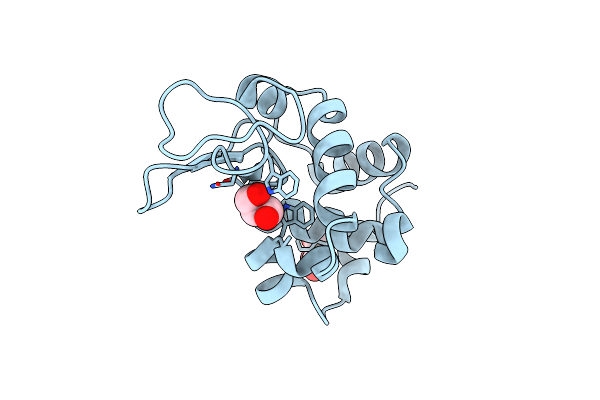
Organism: Tgev virulent purdue, Canis lupus familiaris
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: ISOMERASE Ligands: NAG |
 |
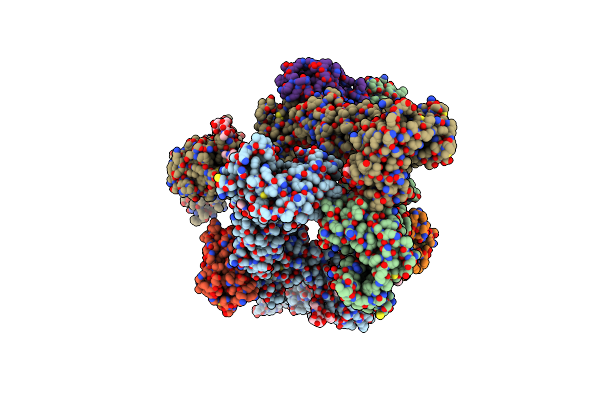
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2024-08-07 Classification: ANTIMICROBIAL PROTEIN Ligands: GOL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-07 Classification: CYTOKINE Ligands: NAG |
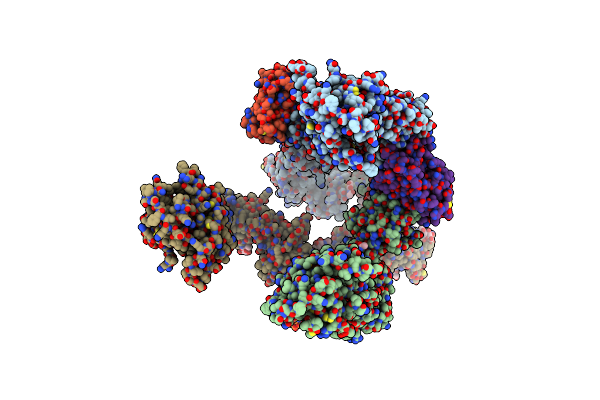 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-07 Classification: CYTOKINE Ligands: NAG |
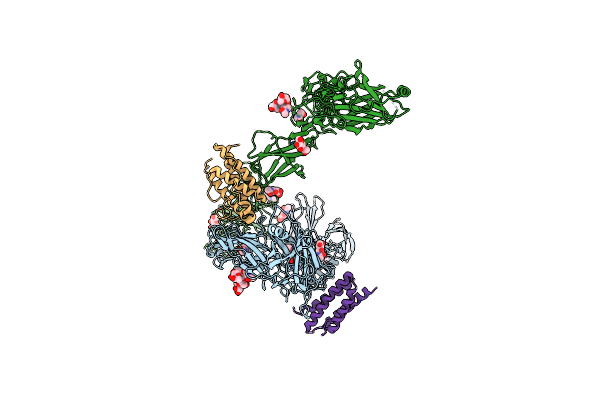 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-07 Classification: CYTOKINE Ligands: NAG |
 |
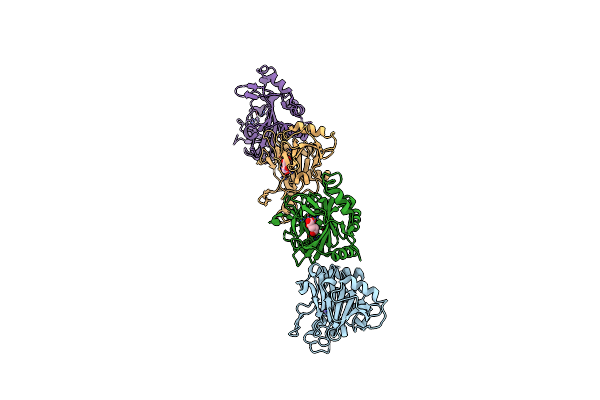
Crystal Structure Of Caenorhabditis Elegans Nmad-1 In Complex With Ligand I
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2024-02-07 Classification: OXIDOREDUCTASE Ligands: SO4 |
 |
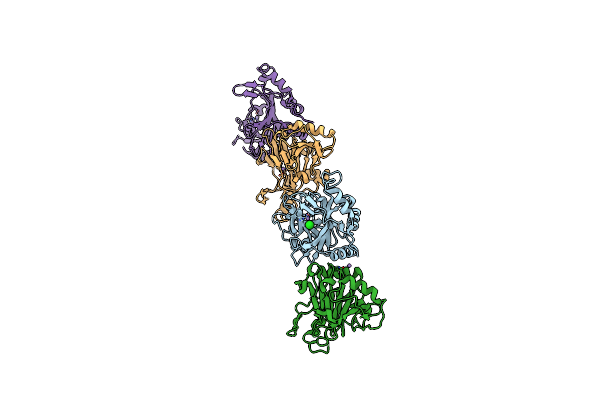
Crystal Structure Of Caenorhabditis Elegans Nmad-1 In Complex With Ligand Ii
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:3.06 Å Release Date: 2024-02-07 Classification: OXIDOREDUCTASE Ligands: MN, AKG |
 |
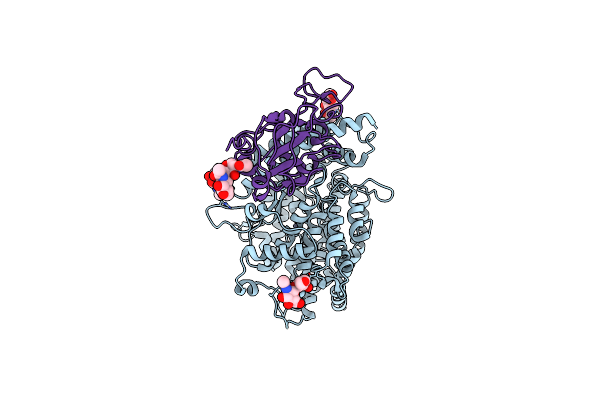
Crystal Structure Of Caenorhabditis Elegans Nmad-1 In Complex With Ligand Iii
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:3.09 Å Release Date: 2024-02-07 Classification: OXIDOREDUCTASE Ligands: MN, CL |
 |
Cryo-Em Structure Of Sars-Cov-2 Delta Rbd In Complex With Golden Hamster Ace2 (Local Refinement)
Organism: Mesocricetus auratus, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: VIRAL PROTEIN/HYDROLASE Ligands: ZN, NAG |
 |
Cryo-Em Structure Of Sars-Cov-2 Ba.3 Rbd In Complex With Golden Hamster Ace2 (Local Refinement)
Organism: Mesocricetus auratus, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: VIRAL PROTEIN/HYDROLASE Ligands: ZN, NAG |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-12-06 Classification: IMMUNE SYSTEM/VIRAL PROTEIN |

