Search Count: 62
All
Selected
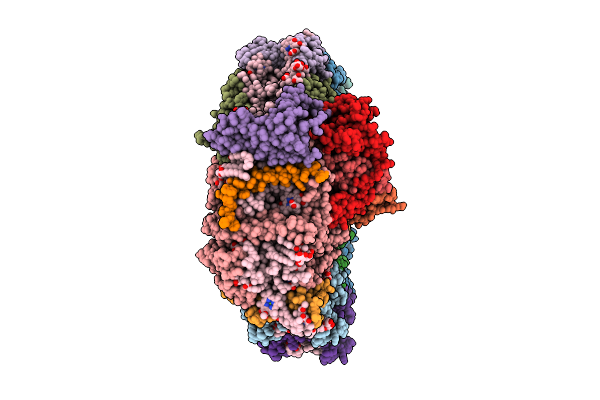 |
Structure Of Photosystem I From Chlamydomonas Reinhardtii At 1.83 A Resolution
Organism: Chlamydomonas reinhardtii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, CHL, BCR, LUT, LMG, LMU, XAT, LHG, NEX, CL0, PQN, SF4, DGD |
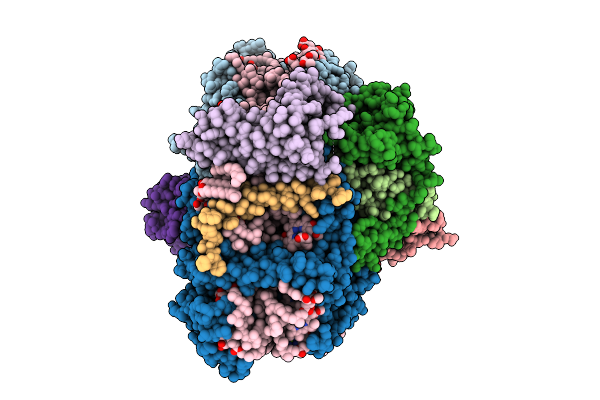 |
Structure Of Cytochrome C6 Bound Photosystem I From Chlamydomonas Reinhardtii At 2.07 A Resolution
Organism: Chlamydomonas reinhardtii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, SF4, LMU, LMG, DGD, LUT, HEC |
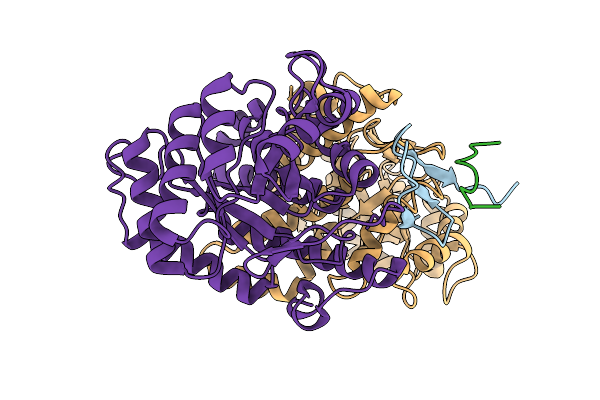 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN |
 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1JAP, L1P, A1JAO, NA, A1JAN |
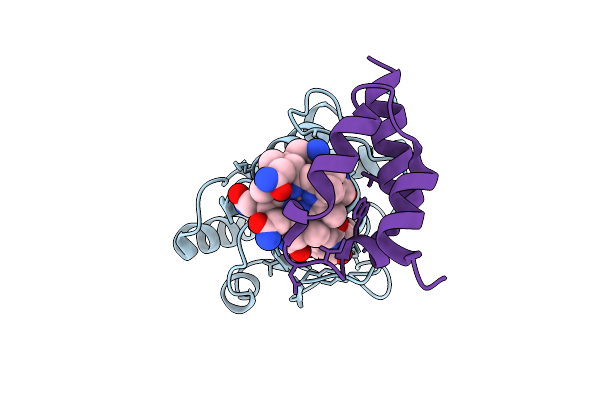 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13 |
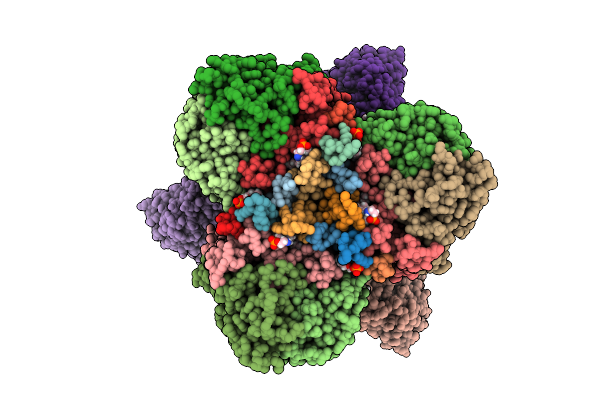 |
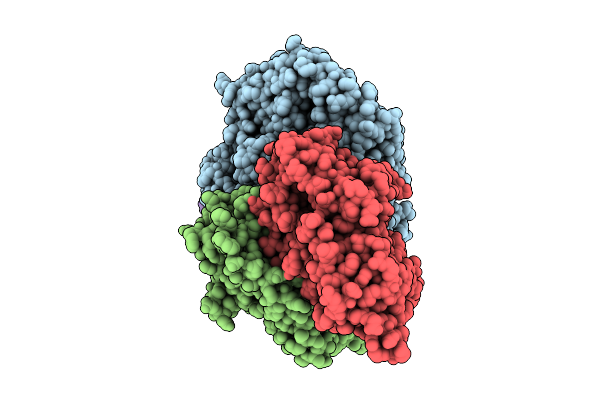
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13, A1JAP, L1P, A1JAO, NA, A1JAN |
 |
Organism: Rhodospirillum rubrum
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: OXIDOREDUCTASE Ligands: S5Q, CLF, ADP, MG, AF3, SF4 |
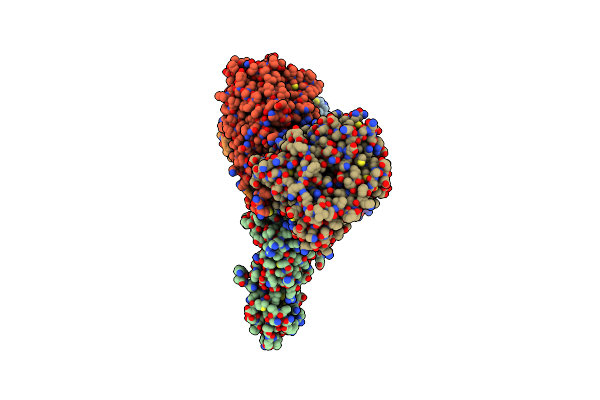 |
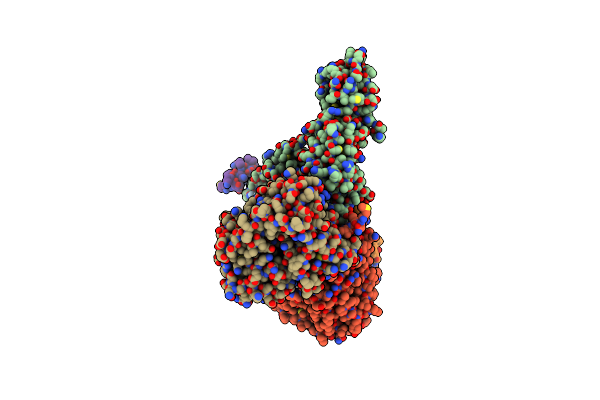
Cryo-Em Structure Of Sodium Pumping Rnf Complex From Acetobacterium Woodii Bound To Nadh
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: MEMBRANE PROTEIN Ligands: FES, SF4, FMN, NAD, RBF |
 |
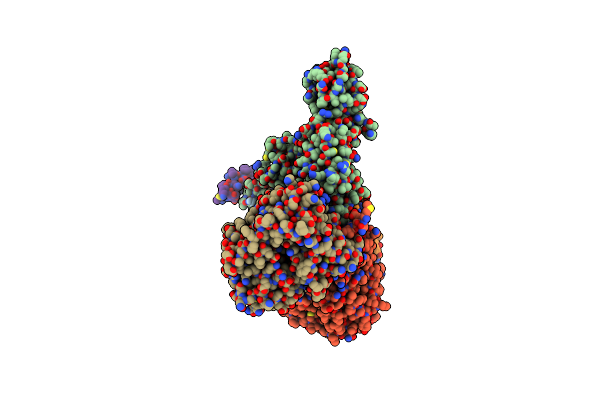
Cryo-Em Structure Of Sodium Pumping Rnf Complex From Acetobacterium Woodii Reduced With Low Potential Ferredoxin
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: MEMBRANE PROTEIN |
 |
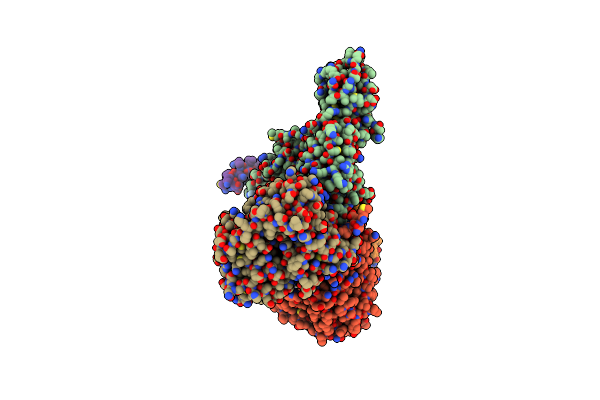
Cryo-Em Structure Of Sodium Pumping Rnf Complex From Acetobacterium Woodii Reduced With Low Potential Ferredoxin (Consensus Map)
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Structure Of Sodium Pumping Rnf Complex From Acetobacterium Woodii In Apo State
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: MEMBRANE PROTEIN |
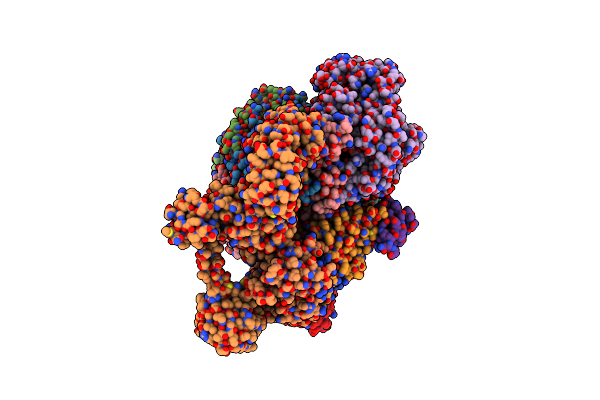 |
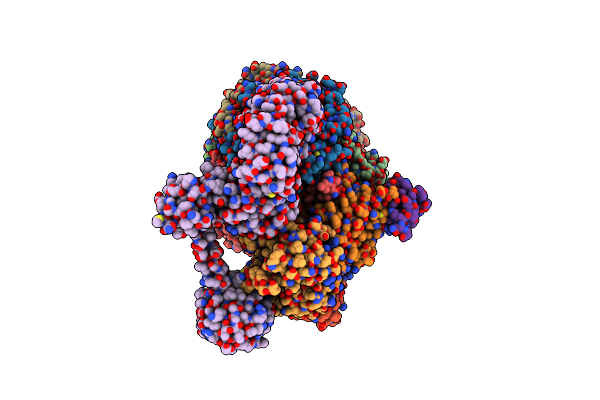
Organism: Methanococcus maripaludis
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: SHT, TP7, COM, F43, S5Q, ZN, ATP, MG |
 |
Organism: Methanococcus maripaludis
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: F43, SHT, COM, TP7, S5Q |
 |
Methyl-Coenzyme M Reductase Activation Complex Binding To The A2 Component After Incubation With Atp
Organism: Methanococcus maripaludis
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: F43, COM, TP7, SHT, S5Q, ZN, ATP, MG |
 |
Organism: Methylophaga sulfidovorans
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2024-07-31 Classification: TRANSFERASE |
 |
Organism: Ananas comosus
Method: ELECTRON MICROSCOPY Release Date: 2024-07-24 Classification: TRANSFERASE |
 |
Organism: Synechococcus elongatus pcc 7942 = fachb-805
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2024-04-24 Classification: TRANSFERASE Ligands: SO4, CIT, MG |
 |
Structure Of Hexameric Subcomplexes (Truncation Delta2-6) Of The Fractal Citrate Synthase From Synechococcus Elongatus Pcc7942
Organism: Synechococcus elongatus pcc 7942 = fachb-805
Method: ELECTRON MICROSCOPY Release Date: 2024-02-28 Classification: TRANSFERASE |
 |
Pseudoatomic Model Of A Second-Order Sierpinski Triangle Formed By The Citrate Synthase From Synechococcus Elongatus
Organism: Synechococcus elongatus pcc 7942 = fachb-805
Method: ELECTRON MICROSCOPY Release Date: 2024-02-28 Classification: TRANSFERASE |
 |
Structure Of A First Order Sierpinski Triangle Formed By The H369R Mutant Of The Citrate Synthase From Synechococcus Elongatus
Organism: Synechococcus elongatus pcc 7942 = fachb-805
Method: ELECTRON MICROSCOPY Release Date: 2024-02-28 Classification: TRANSFERASE |

