Search Count: 1,320
All
Selected
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 1996-08-01 Classification: RACEMASE Ligands: MG, APG |
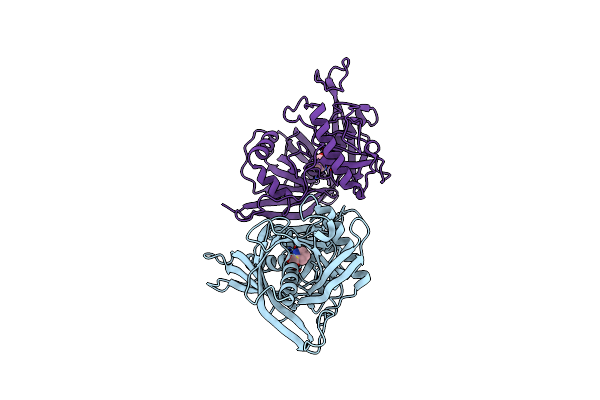 |
Proline Racemase In Complex With 2 Molecules Of Pyrrole-2-Carboxylic Acid (Holo Form)
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2006-02-03 Classification: RACEMASE Ligands: PYC |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 1996-10-14 Classification: RACEMASE Ligands: MG, APG |
 |
Mechanism Of The Reaction Catalyzed By Mandelate Racemase. 2. Crystal Structure Of Mandelate Racemase At 2.5 Angstroms Resolution: Identification Of The Active Site And Possible Catalytic Residues
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 1994-01-31 Classification: RACEMASE Ligands: MN, SO4 |
 |
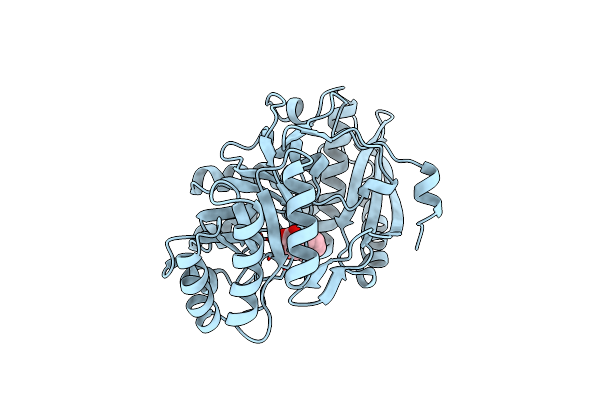
On The Role Of Lysine 166 In The Mechanism Of Mandelate Racemase From Pseudomonas Putida: Mechanistic And Crystallographic Evidence For Stereospecific Alkylation By (R)-Alpha-Phenylglycidate
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 1993-10-31 Classification: RACEMASE Ligands: MG, APG |
 |
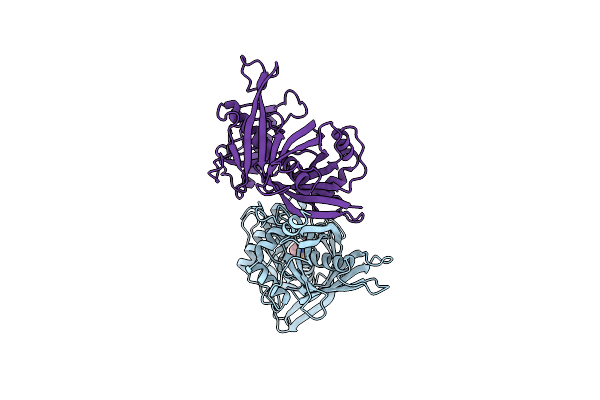
The Role Of Lysine 166 In The Mechanism Of Mandelate Racemase From Pseudomonas Putida: Mechanistic And Crystallographic Evidence For Stereospecific Alkylation By (R)-Alpha-Phenylglycidate
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 1994-08-31 Classification: RACEMASE Ligands: MG, APG |
 |
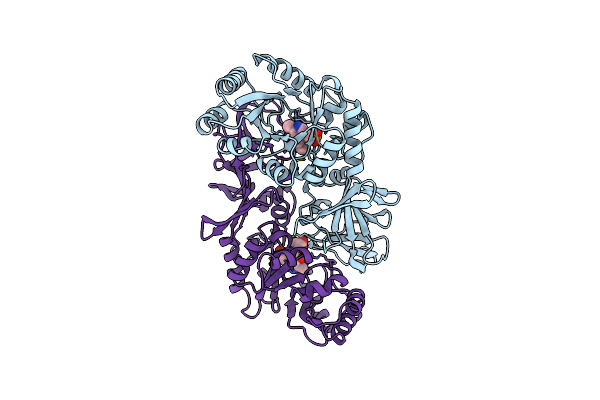
Proline Racemase In Complex With One Molecule Of Pyrrole-2-Carboxylic Acid (Hemi Form)
Organism: Trypanosoma cruzi (strain cl brener)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2006-02-03 Classification: RACEMASE Ligands: PYC |
 |
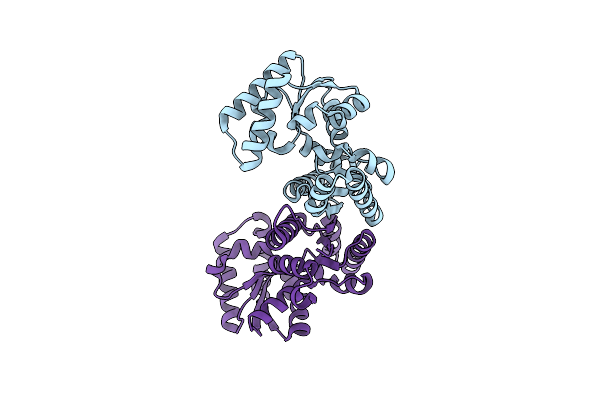
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 1999-02-24 Classification: RACEMASE Ligands: PLP, PPI |
 |
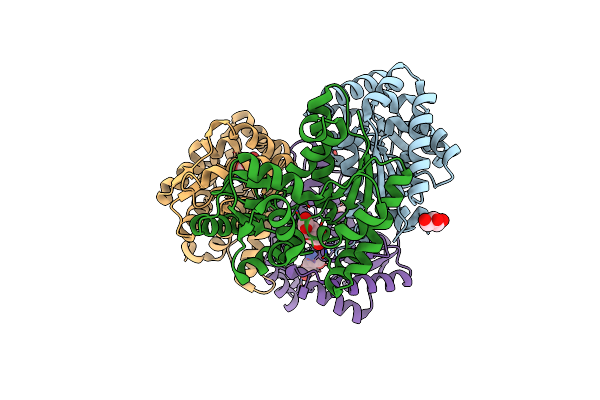
Crystal Structure Of L-Aspartate/Glutamate-Specific Racemase From Escherichia Coli
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2015-11-18 Classification: ISOMERASE |
 |
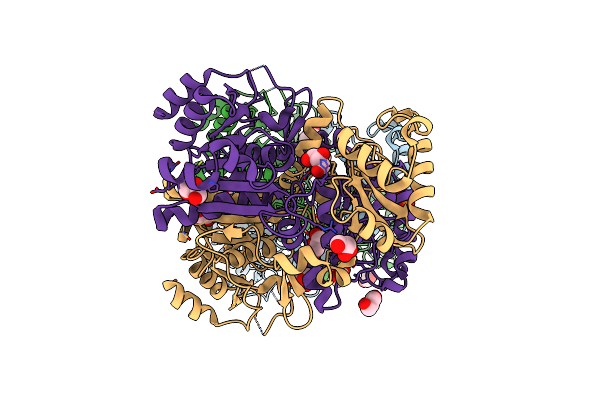
Crystal Structure Of L-Aspartate/Glutamate Specific Racemase In Complex With L-Glutamate
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2015-11-18 Classification: ISOMERASE Ligands: GOL, GLU, NO3 |
 |
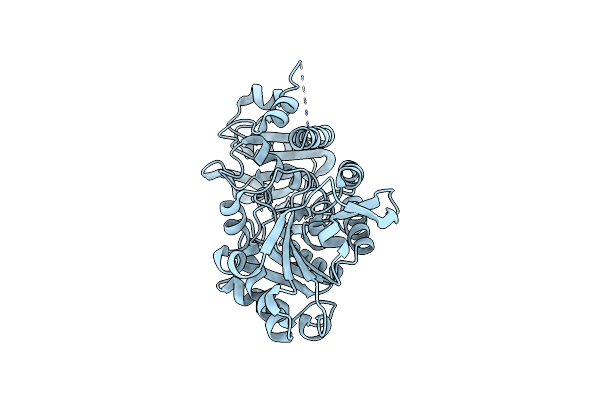
Alpha-Methylacyl-Coa Racemase From Mycobacterium Tuberculosis- Mutational And Structural Characterization Of The Fold And Active Site
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2005-01-18 Classification: ISOMERASE Ligands: GOL, PO4 |
 |
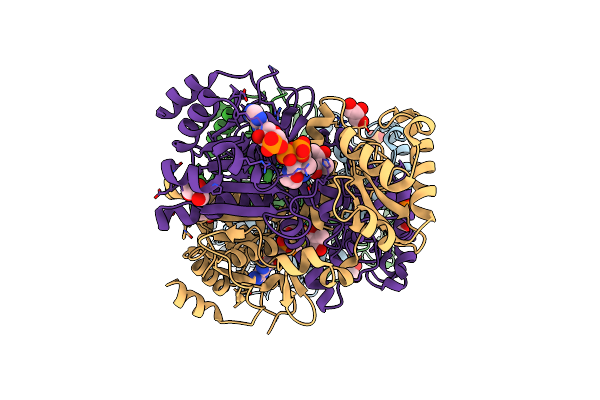
Structure Of The Malate Racemase Mar2 Apoprotein From Thermoanaerobacterium Thermosaccharolyticum At 2.25 Angstroms Resolution
Organism: Thermoanaerobacterium thermosaccharolyticum (strain atcc 7956 / dsm 571 / ncib 9385 / nca 3814)
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2021-10-13 Classification: ISOMERASE Ligands: CL |
 |
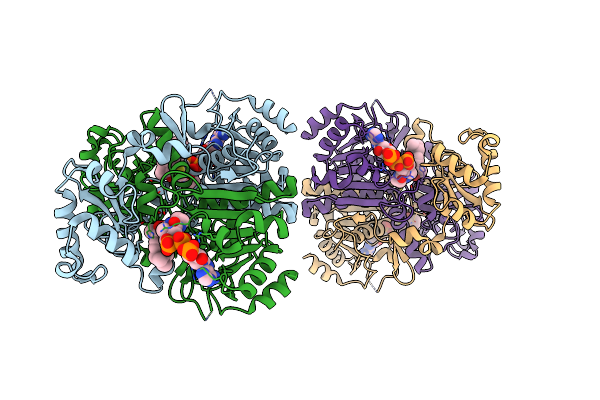
The Enolisation Chemistry Of A Thioester-Dependent Racemase: The 1.4 A Crystal Structure Of A Complex With A Planar Reaction Intermediate Analogue
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2012-03-28 Classification: ISOMERASE Ligands: GOL, MC4, PO4 |
 |
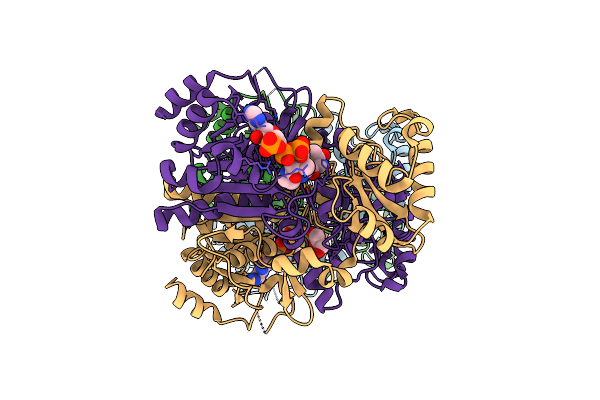
The 1,1-Proton Transfer Reaction Mechanism By Alpha-Methylacyl-Coa Racemase Is Catalyzed By An Aspartate/Histidine Pair And Involves A Smooth, Methionine-Rich Surface For Binding The Fatty Acyl Moiety
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2007-02-20 Classification: ISOMERASE Ligands: SFC, RFC |
 |
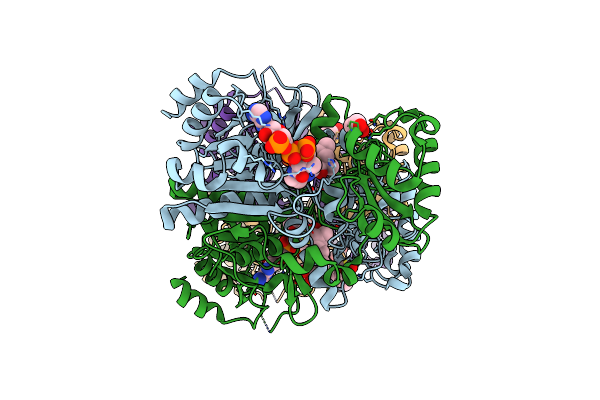
The 1,1-Proton Transfer Reaction Mechanism By Alpha-Methylacyl-Coa Racemase Is Catalyzed By An Aspartate/Histidine Pair And Involves A Smooth, Methionine-Rich Surface For Binding The Fatty Acyl Moiety
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2007-02-20 Classification: ISOMERASE Ligands: CAA, GOL |
 |
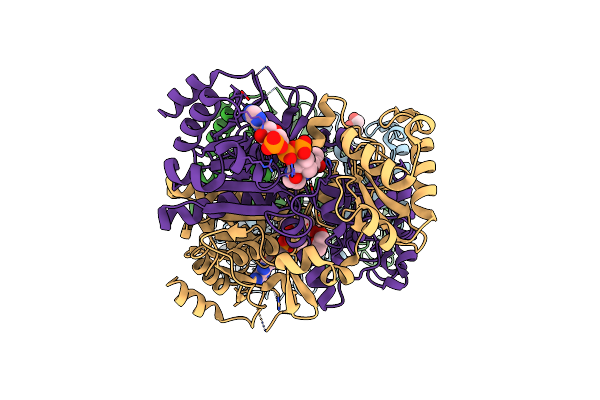
The 1,1-Proton Transfer Reaction Mechanism By Alpha-Methylacyl-Coa Racemase Is Catalyzed By An Asparte/Histidine Pair And Involves A Smooth, Methionine-Rich Surface For Binding The Fatty Acyl Moiety
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2007-02-20 Classification: ISOMERASE Ligands: MRR, GOL |
 |
The 1,1-Proton Transfer Reaction Mechanism By Alpha-Methylacyl-Coa Racemase Is Catalyzed By An Aspartate/Histidine Pair And Involves A Smooth, Methionine-Rich Surface For Binding The Fatty Acyl Moiety
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2007-02-20 Classification: ISOMERASE Ligands: ACO, GOL |
 |
The 1,1-Proton Transfer Reaction Mechanism By Alpha-Methylacyl-Coa Racemase Is Catalyzed By An Aspartate/Histidine Pair And Involves A Smooth, Methionine-Rich Surface For Binding The Fatty Acyl Moiety
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2007-02-20 Classification: ISOMERASE Ligands: MRS, GOL |
 |
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-03-25 Classification: ISOMERASE Ligands: PP3 |
 |
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2026-03-25 Classification: ISOMERASE Ligands: PDD |

