Search Count: 3,503
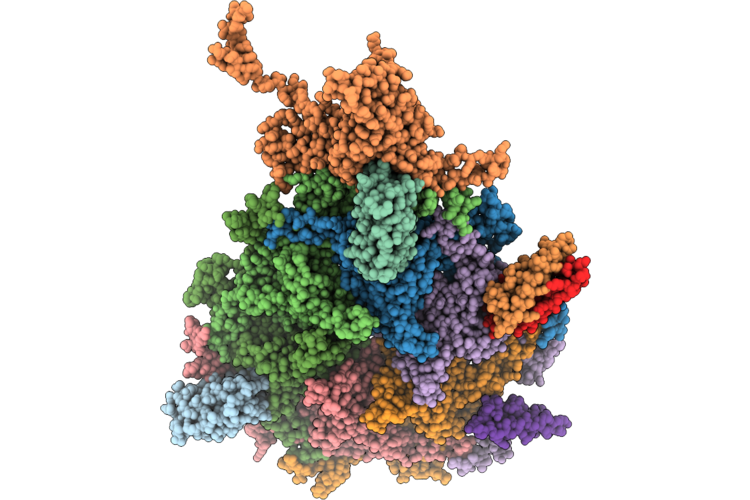 |
Organism: Klebsiella phage ran69
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: VIRUS |
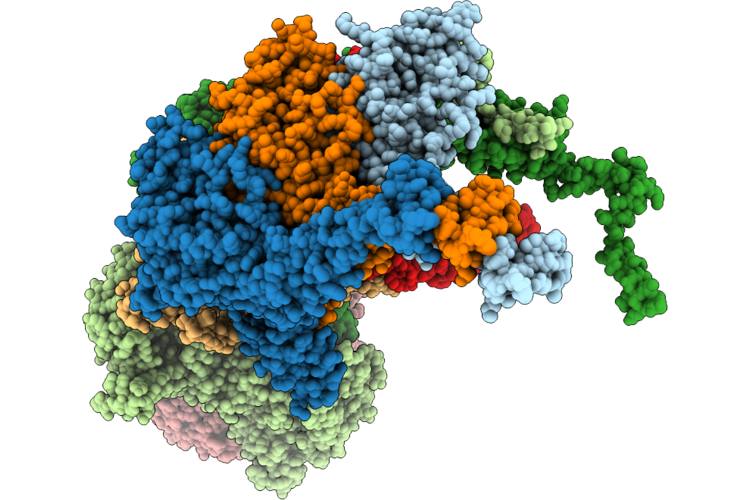 |
The Composite Cryo-Em Structure Of Bacteriophage Ran69 Pre-Ejectosome-Portal Complex
Organism: Klebsiella phage ran69
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: VIRUS |
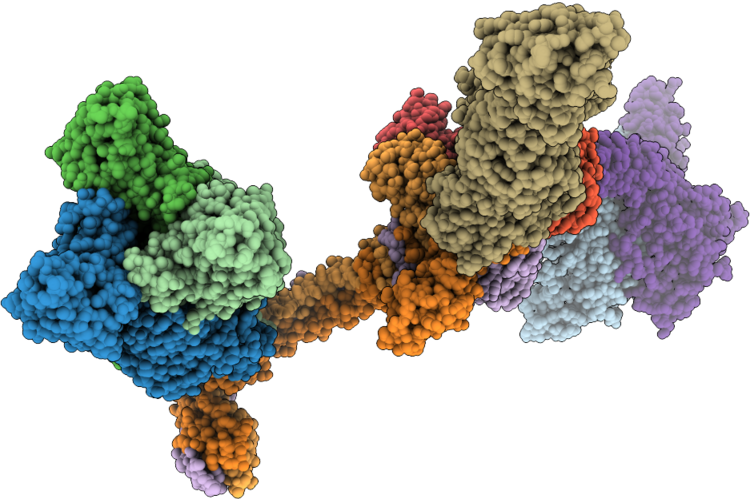 |
Organism: Klebsiella phage ran69
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-05-27 Classification: VIRUS |
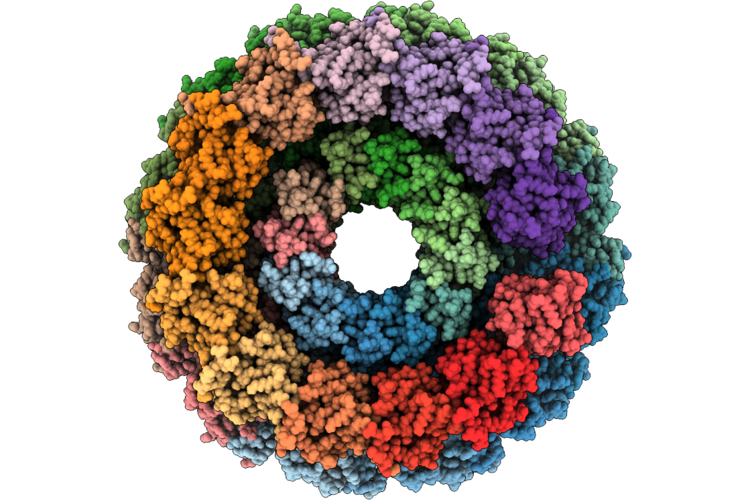 |
Organism: Escherichia phage theodorherzl, Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: HYDROLASE Ligands: MG |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
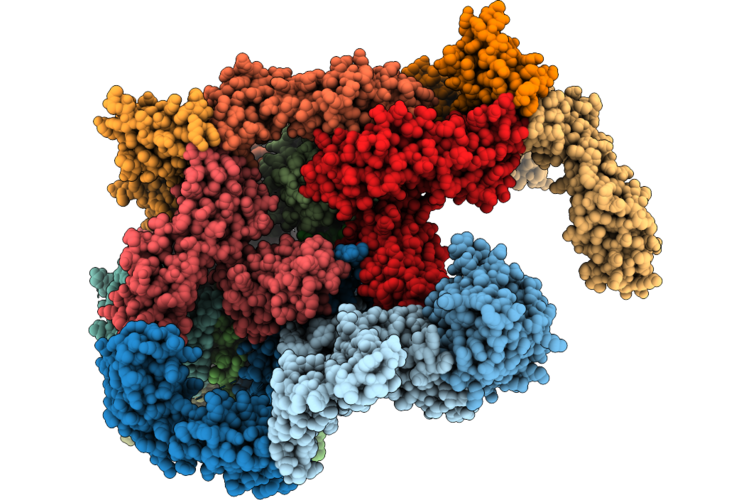 |
Phage 812 Baseplate In The Pre-Contraction State - Core And Wedge Module Proteins
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
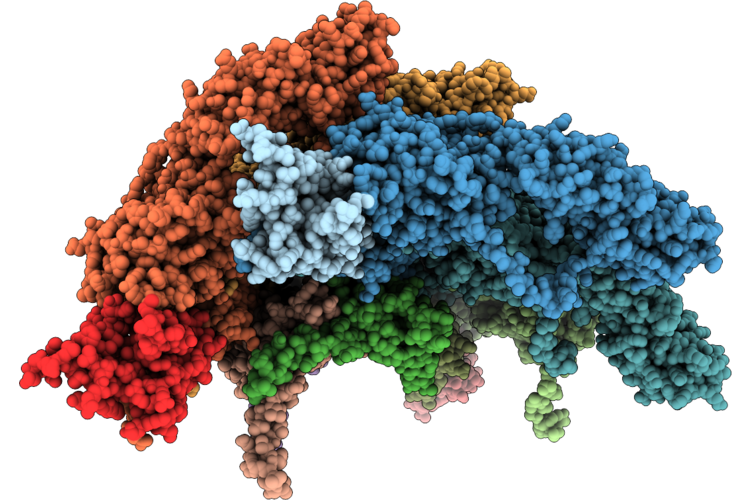 |
Phage 812 Baseplate In The Pre-Contraction State - Tail Sheath Initiator And Baseplate-Proximal Tail Proteins
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
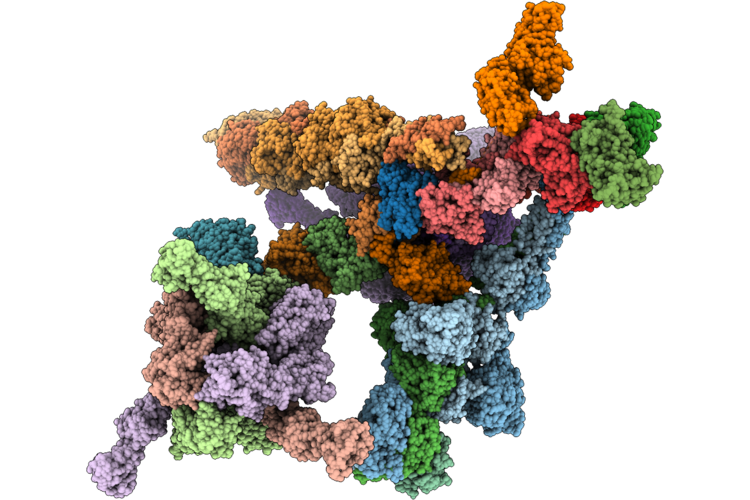 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
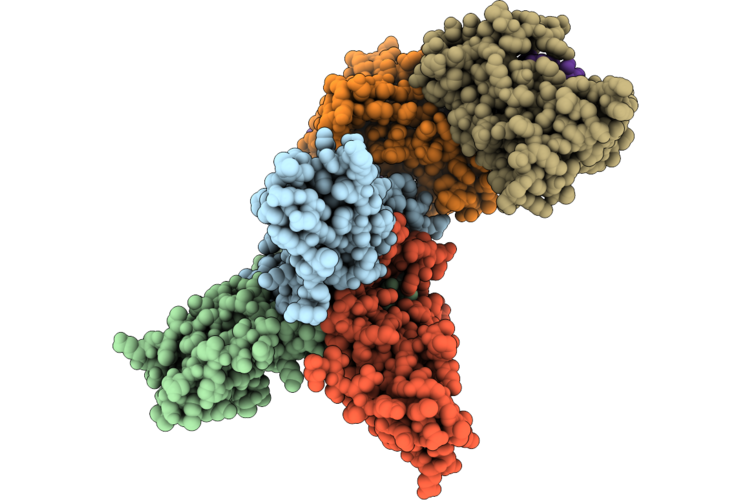 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Phage 812 Baseplate In The Pre-Contraction State - Upper Arm (Segment Cdef)
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |
 |
Organism: Staphylococcus phage 812k1/420
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: VIRUS |

