Search Count: 46
All
Selected
 |
Organism: Amycolatopsis deserti
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-04-15 Classification: HYDROLASE |
 |
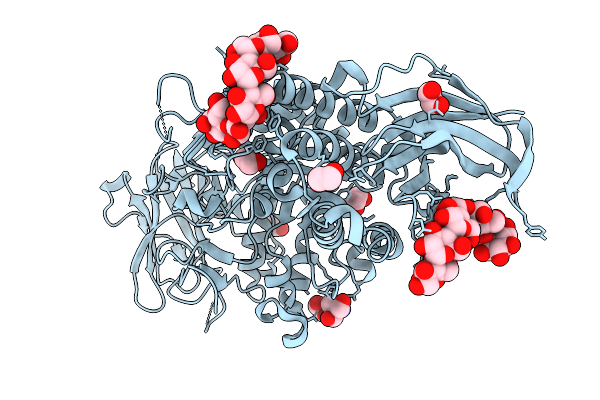
Organism: Faecalibacterium duncaniae
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: TRS, GOL, EDO, PEG |
 |
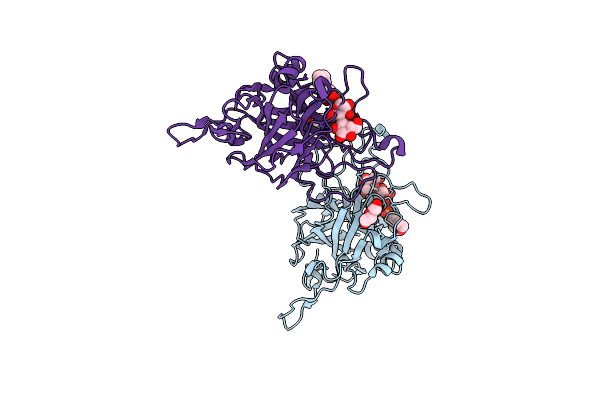
Organism: Faecalibacterium duncaniae
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: TRS, EDO |
 |
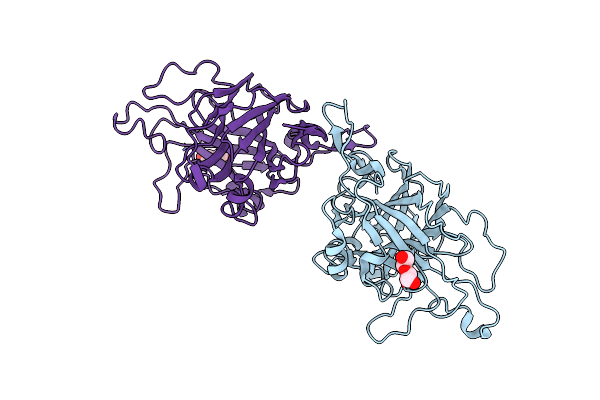
Coprinopsis Cinerea Gh131 Protein Ccgh131B E161A In Complex With Cellobiose
Organism: Coprinopsis cinerea (strain okayama-7 / 130 / atcc mya-4618 / fgsc 9003)
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-11-06 Classification: HYDROLASE Ligands: MES, PEG, PGE |
 |
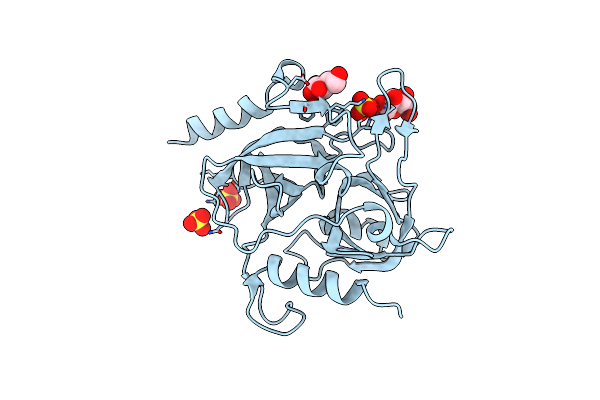
Organism: Coprinopsis cinerea (strain okayama-7 / 130 / atcc mya-4618 / fgsc 9003)
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-11-06 Classification: HYDROLASE Ligands: PEG |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-09-11 Classification: HYDROLASE Ligands: SO4, BGC |
 |
Organism: Human immunodeficiency virus type 1 (new york-5 isolate)
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2024-07-10 Classification: VIRAL PROTEIN Ligands: PG4, A1L1V, SO4 |
 |
Organism: Human immunodeficiency virus type 1 (new york-5 isolate)
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-07-10 Classification: VIRAL PROTEIN Ligands: PG4, A1L1W, SO4 |
 |
Organism: Rhodothermus marinus jcm 9785
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-02-07 Classification: HYDROLASE Ligands: MN, CA, MES, MPD |
 |
Rhodothermus Marinus Alpha-Amylase Rmgh13_47A Cbm48-A-B-C Domains In Complex With Branched Pentasaccharide
Organism: Rhodothermus marinus jcm 9785
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-02-07 Classification: HYDROLASE Ligands: MN, CA, MES, PGE, PEG, GOL |
 |
Organism: Beijerinckia indica subsp. indica
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2023-06-14 Classification: TRANSFERASE Ligands: GOL |
 |
Beijerinckia Indica Beta-Fructosyltransferase Variant H395R/F473Y In Complex With Fructose
Organism: Beijerinckia indica subsp. indica nbrc 3744
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2023-06-14 Classification: TRANSFERASE Ligands: MG, FRU, BDF |
 |
Organism: Aspergillus sojae nbrc 4239
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2022-06-15 Classification: HYDROLASE Ligands: NAG, NA, GLC |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2022-03-30 Classification: TRANSFERASE |
 |
Organism: Frischella perrara
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2022-03-09 Classification: HYDROLASE Ligands: PEG, PGE |
 |
Organism: Frischella perrara
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2022-03-09 Classification: HYDROLASE Ligands: FRU, PEG, 1PE |
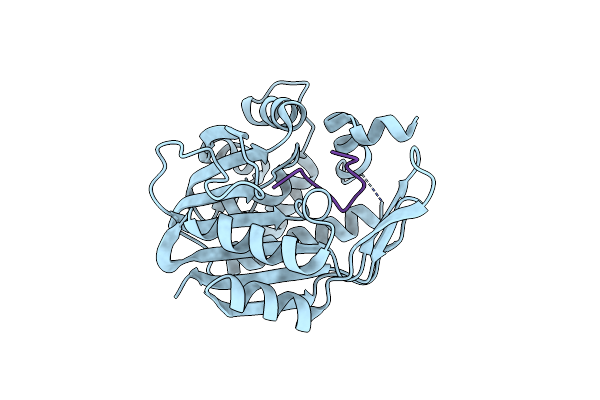 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2021-12-15 Classification: TRANSFERASE |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2021-12-15 Classification: TRANSFERASE |
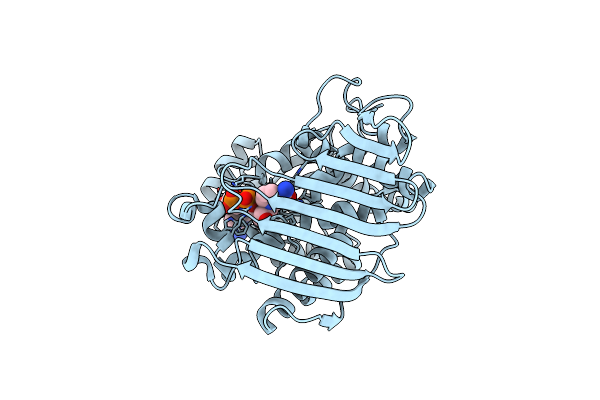 |
Crystal Structure Of Inositol Dehydrogenase Homolog Complexed With Nad+ From Azotobacter Vinelandii
Organism: Azotobacter vinelandii (strain dj / atcc baa-1303)
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2021-09-29 Classification: OXIDOREDUCTASE Ligands: NAD |
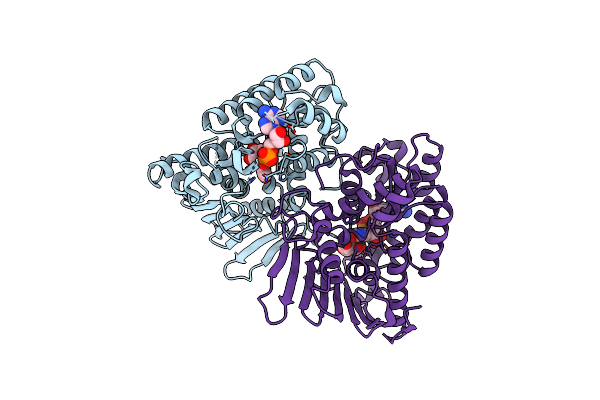 |
Crystal Structure Of Inositol Dehydrogenase Homolog Complexed With Nadh And Myo-Inositol From Azotobacter Vinelandii
Organism: Azotobacter vinelandii (strain dj / atcc baa-1303)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2021-09-29 Classification: OXIDOREDUCTASE Ligands: NAI, INS |

