Search Count: 2,205
All
Selected
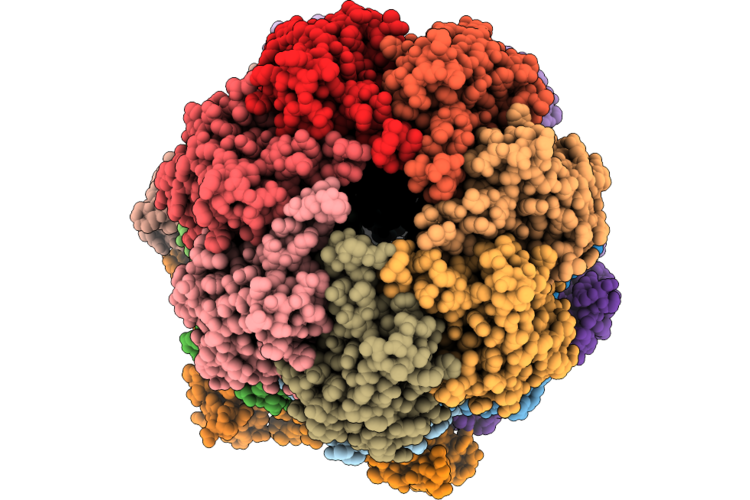 |
Organism: Enterobacter
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-22 Classification: CHAPERONE Ligands: ATP |
 |
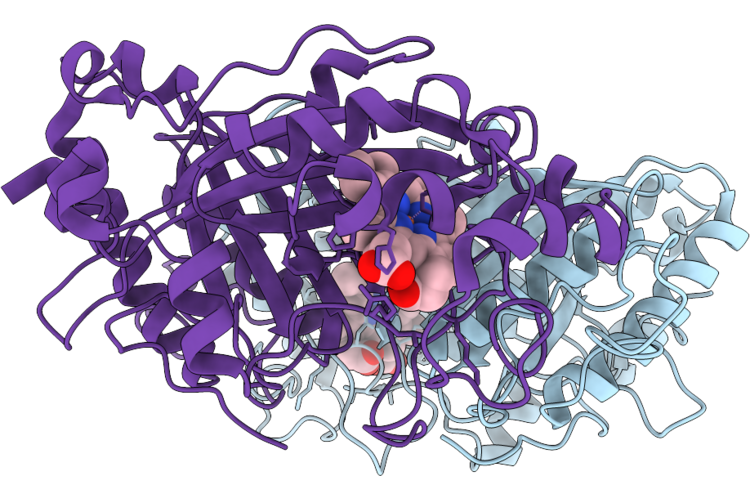
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
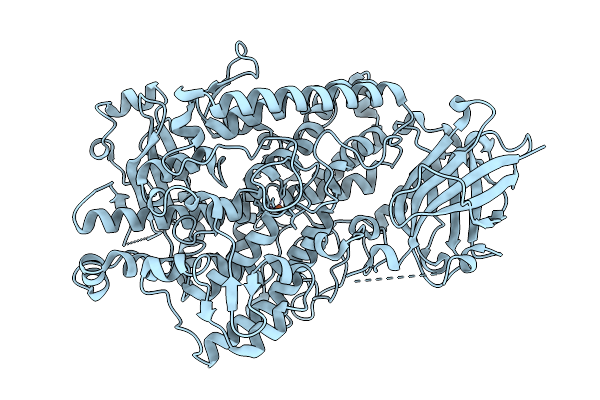
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
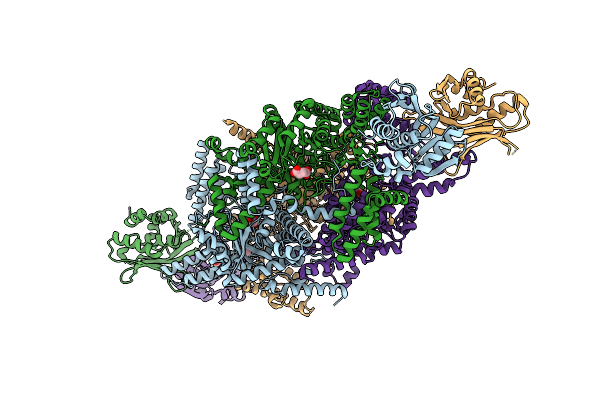
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE |
 |
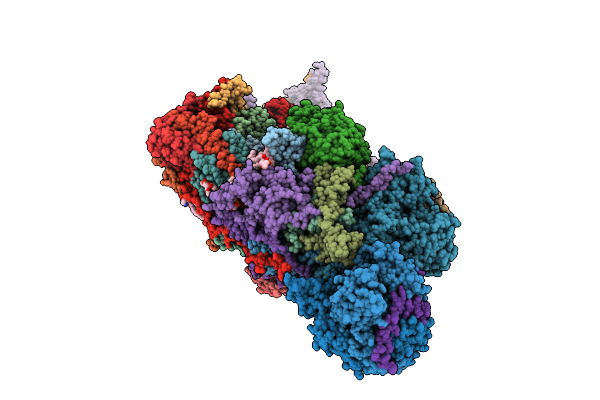
Crystal Structure Of Mycobacterium Tuberculosis Isocitrate Lyase 2 Fixed In The Apo Form With Disulfide Bonds
Organism: Mycobacterium tuberculosis cdc1551
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-03-18 Classification: LYASE Ligands: SIN, EDO |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: 3PE, PC1, SF4, U10, FES, FMN, NAI, K, CDL, DGT, MG, NDP, ZN, EHZ, CHD, MYR |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: PC1, SF4, FES, FMN, NAI, K, U10, 3PE, CDL, DGT, MG, NDP, ZN, EHZ, CHD, MYR |
 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: 3PE, PC1, SF4, FES, NAI, FMN, K, LMT, CDL, DGT, MG, NDP, ZN, EHZ, CHD, MYR |
 |
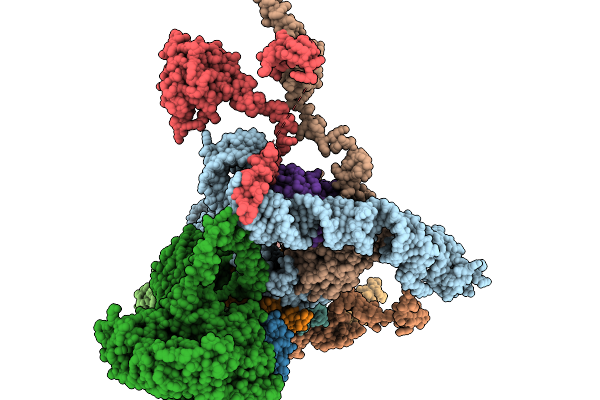
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: ELECTRON TRANSPORT Ligands: 3PE, PC1, SF4, LMT, FES, FMN, NAI, K, CDL, CHD, DGT, MG, NDP, ZN, EHZ, MYR |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSLATION Ligands: KGN |
 |
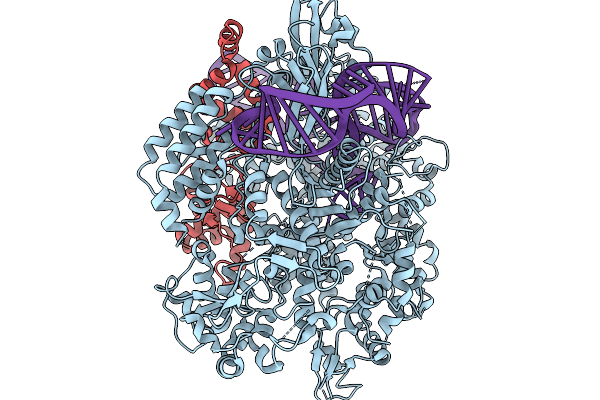
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSLATION Ligands: IHP |
 |
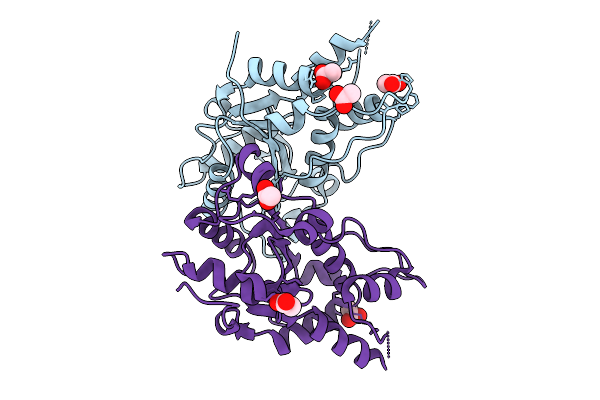
Organism: Streptococcus pyogenes m1 gas, Streptococcus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: IMMUNE SYSTEM |
 |
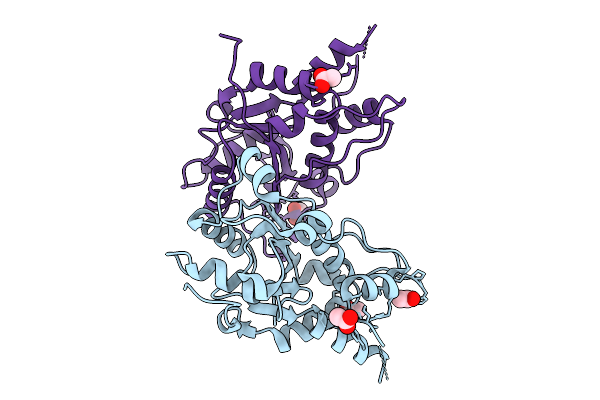
The Apo-Mtrex1 Crystal Structure For Soaking Experiments (Soaking Condition 1)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: ACT |
 |
The Apo-Mtrex1 Crystal Structure For Soaking Experiments (Soaking Condition 2)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: ACT |
 |
The Apo-Mtrex1 Crystal Structure For Soaking Experiments (Soaking Condition 3)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: GOL |
 |
The Mtrex1-Nsc 37203 Complex Structure By Co-Crystallization (Nsc 37203 Complex 1)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: DMS, LI, EPE, SO4, A1L5X |
 |
The Mtrex1-Nsc 37203 Complex Structure By Soaking In Soaking Condition 1 (Nsc 37203 Complex 2)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: DMS, MG, ACT, A1L5X, CAC, NA |
 |
The Mtrex1-Nsc 37203 Complex Structure By Co-Crystallization (Nsc 37203 Complex 3)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: SO4, A1L5X |
 |
The Mtrex1-Nsc 37204 Complex Structure By Soaking In Soaking Condition 2 (Nsc 37204 Complex 1)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: ACT, A1L5Z |

