Search Count: 8
All
Selected
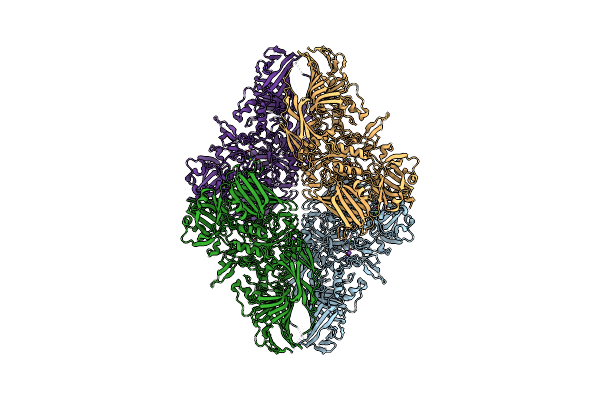 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: HYDROLASE Ligands: MG, NA |
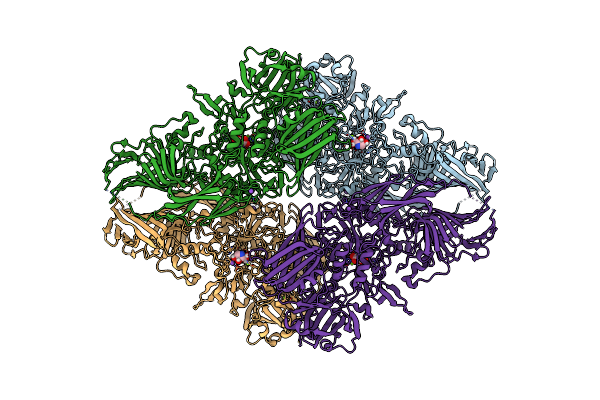 |
Beta-Galactosidase (Lacz) In Complex With Glycosyrin From Pseudomonas Syringae.
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: HYDROLASE Ligands: A1H05, MG, NA |
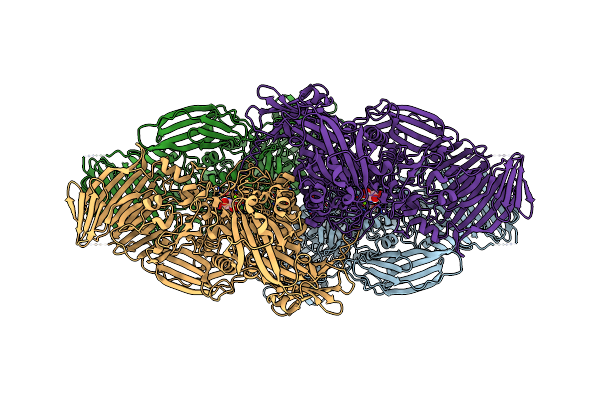 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: HYDROLASE Ligands: A1H05, MG, NA |
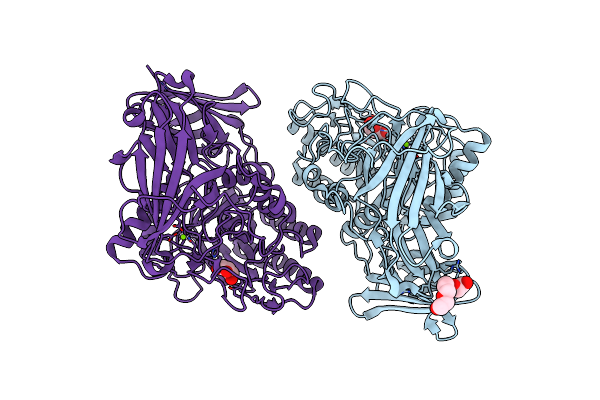 |
Beta-D-Galactofuranosidase Orf1110 From Streptomyces Sp. Jha19, Ligand-Free Form
Organism: Streptomyces sp. jha19
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-11-27 Classification: HYDROLASE Ligands: GOL, PG4, MG |
 |
Beta-D-Galactofuranosidase Orf1110 From Streptomyces Sp. Jha19, D-Iminogalactitol Complex
Organism: Streptomyces sp. jha19
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-11-27 Classification: HYDROLASE Ligands: P6G, MPD, A1L3Z, MG, PG4 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2010-05-19 Classification: HYDROLASE Ligands: MN3, EDO, PO4, CIT |
 |
Crystal Structure Of An N-Terminal Mutant Of The Plasmid Pcu1 Trai Relaxase Domain
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2010-05-19 Classification: HYDROLASE Ligands: CIT, EDO, NI, CL, PO4, MG, NA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2000-02-28 Classification: DNA BINDING PROTEIN |

