Search Count: 2,121
 |
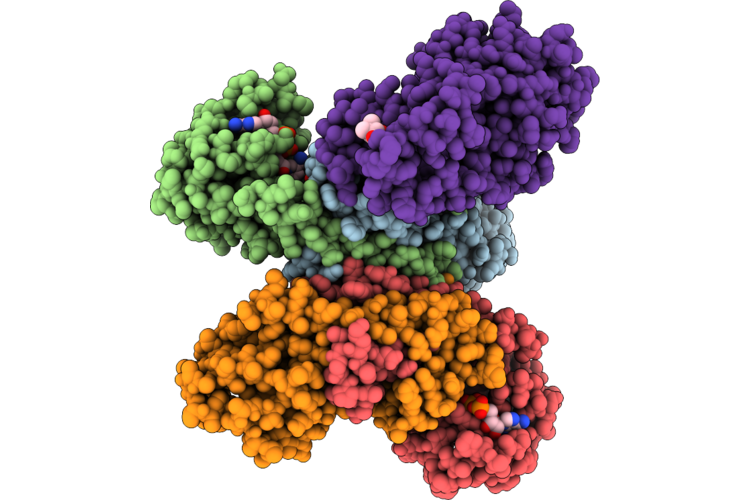
Assembly Intermediate Of Human Mitochondrial Ribosome Small Subunit In Complex With Pus1, Mettl15 And Rbfa(In Conformation) (State M1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-05-27 Classification: RIBOSOME Ligands: NAD, SPM, MG, K, ZN, FES, SAH, ATP, GDP |
 |
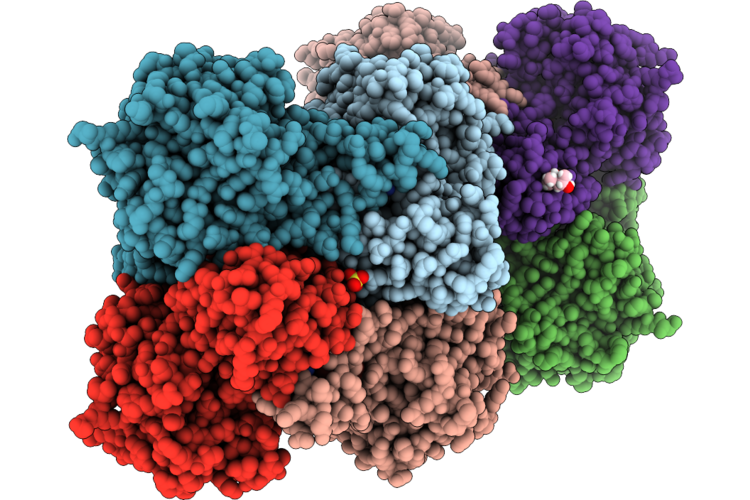
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2026-05-20 Classification: OXIDOREDUCTASE Ligands: SO4, NAD, G8I |
 |
Crystal Structure Of Human S-Adenosyl-L-Homocysteine Hydrolase Complex With Adenosine
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: NAD, ADN, K, SO4, MPD |
 |
Crystal Structure Of Human S-Adenosyl-L-Homocysteine Hydrolase In Complex With Adenosine And Cadmium Ions.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: NAD, ADN, CD, K |
 |
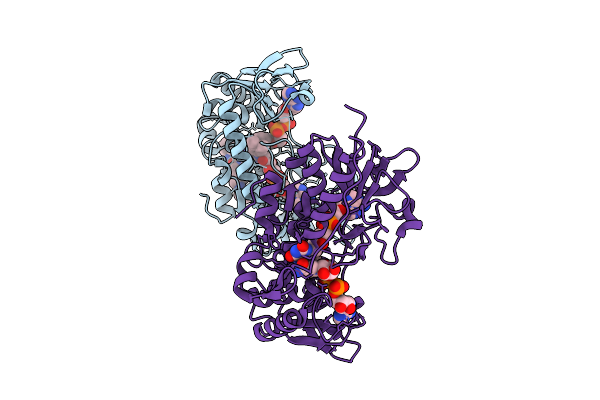
Crystal Structure Of Klebsiella Oxytoca Ribitol Dehydrogenase In Complex With Nad+
Organism: Klebsiella oxytoca
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
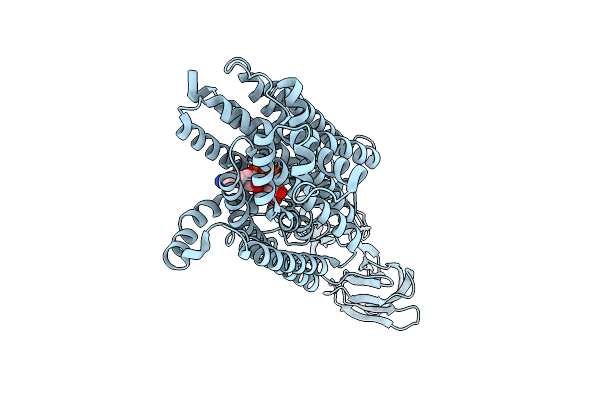
Fsp1 (Tetrapod Ancestor; L323A Mutant) Bound To Fad And Nad+ And Compound 2
Organism: Mammalia
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, NAD, A1I9W, CA |
 |
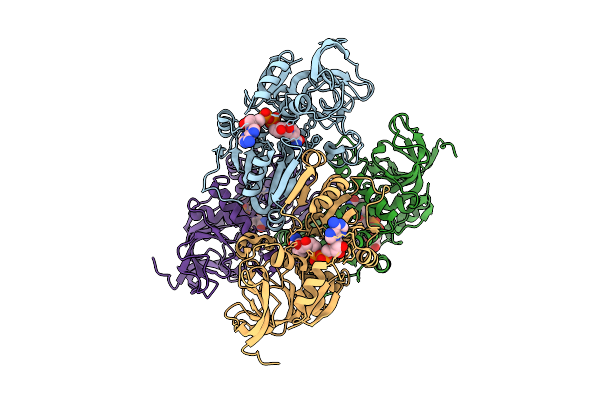
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.38 Å Release Date: 2026-04-15 Classification: MEMBRANE PROTEIN Ligands: NAD |
 |
Crystal Structure Of (R)-Carbonyl Reductase Mutant I51L/Y61F/D147E From Candida Parapsilosis In Complex With Nad
Organism: Candida parapsilosis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-15 Classification: BIOSYNTHETIC PROTEIN Ligands: NAD, ZN |
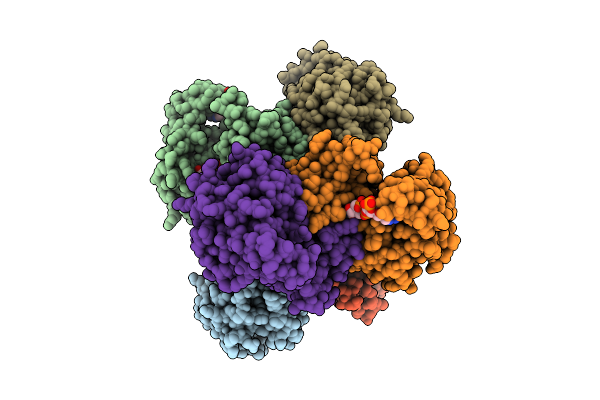 |
Organism: Faecalibacterium prausnitzii l2-6
Method: X-RAY DIFFRACTION Resolution:3.29 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: NAD, CAA |
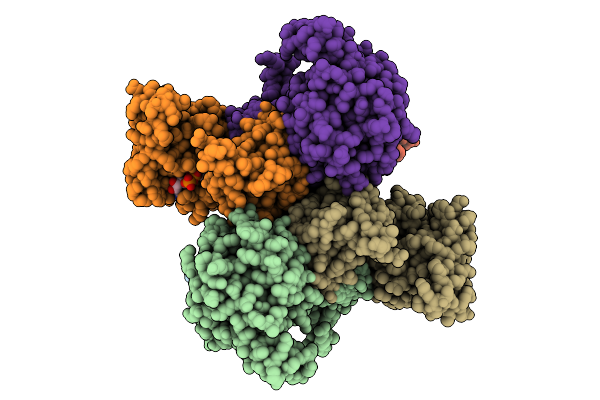 |
Organism: Faecalibacterium prausnitzii l2-6
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
Crystal Structure Of The S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa, Q65N Mutant Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, GOL, PO4 |
 |
Crystal Structure Of S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa: Q65A Mutant Soaked With Adenosine And Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, ADN, PO4 |
 |
Crystal Structure Of S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa, Q65N Mutant Soaked With Adenosine And Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, ADN, PO4 |
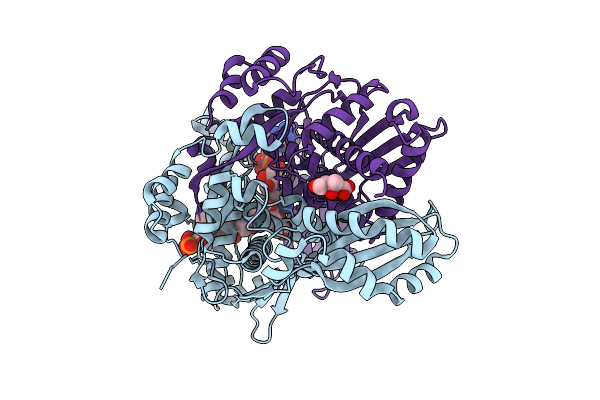 |
Structure Of Pmhmgr Bound To Mevalonate, Coa And Nad 1 Minute After Reaction Initiation At Ph 9
Organism: Pseudomonas sp. 'mevalonii'
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE/SUBSTRATE/INTERMEDIATE Ligands: COA, A1AFY, NAI, NAD, MEV |
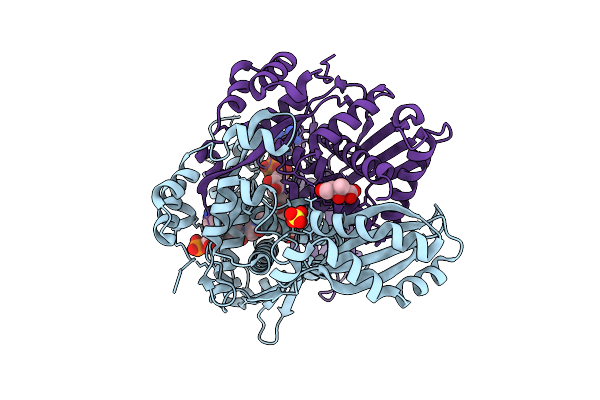 |
Pmhmgr Bound To Mevalonate, Coa, And Nad, Buffer-Exchanged To Ammonium Acetate Environment At Ph 6.7
Organism: Pseudomonas sp. 'mevalonii'
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: COA, MEV, SO4, NAD |
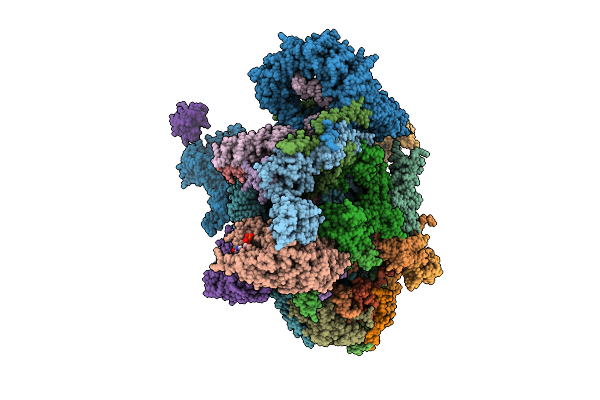 |
Assembly Intermediate Of Human Mitochondrial Ribosome Small Subunit Bound To Mettl15, Rbfa, And Mtif2 (State M2.1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: RIBOSOME Ligands: MG, ZN, FES, ATP, GDP, NAD, SPM |
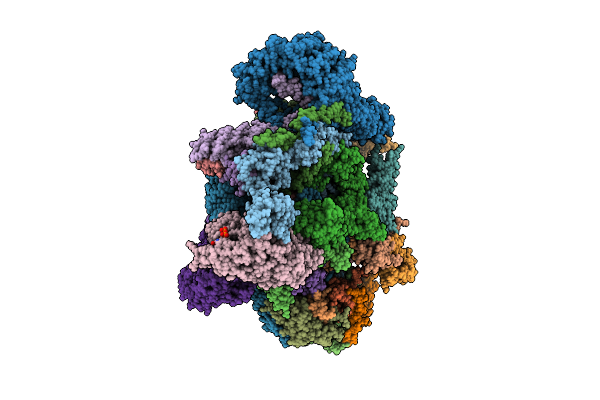 |
Assembly Intermediate Of Human Mitochondrial Ribosome Small Subunit Bound To Mettl15 And Rbfa (Inward Conformation) (State M2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: RIBOSOME Ligands: MG, FES, ZN, ATP, GDP, NAD, SPM |
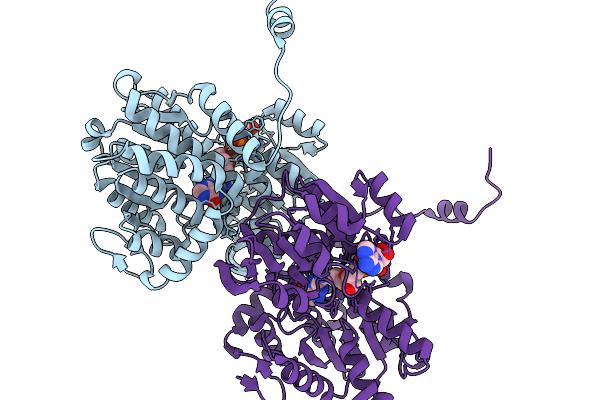 |
Crystal Structure Of S-Adenosyl-L-Homocysteine Hydrolase From Pyrococcus Furiosus In Complex With Hypoxanthine
Organism: Pyrococcus furiosus
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: NAD, HPA |
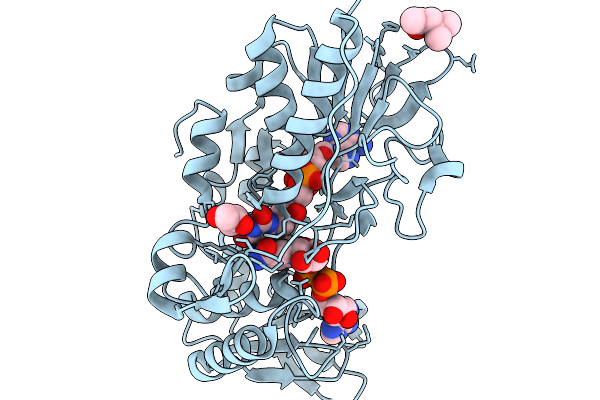 |
Organism: Mammalia
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: FAD, NAD, MPD, GOL |
 |
Organism: Mammalia
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: FAD, NAD, A1I3Y, CL, MG |

