Search Count: 63
All
Selected
 |
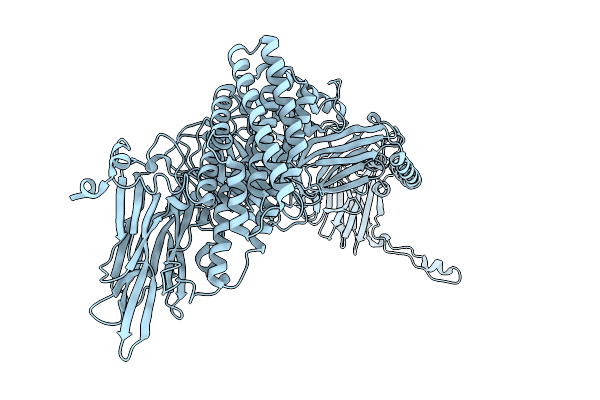
Native Structure Of Full-Length Pesticidal Protein Cry1Ca18 At Ph9, From Crystals Formed In Vivo
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-02-11 Classification: TOXIN Ligands: CA, ZN |
 |
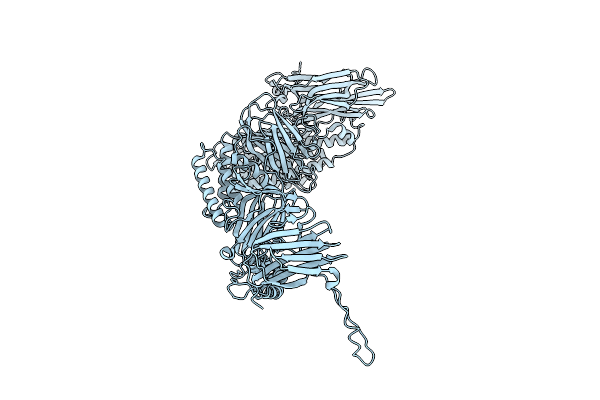
Native Structure Of The Full-Length Pesticidal Protein Cry8Ba2, From Crystals Formed In Vivo (Form 1)
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2025-12-03 Classification: TOXIN |
 |
Native Structure Of The Full-Length Pesticidal Protein Cry8Ba2, From Crystals Formed In Vivo (Form 2)
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2025-12-03 Classification: TOXIN |
 |
Native Structure Of The Full-Length Pesticidal Protein Cry1Ca18 At Ph7, From Crystals Formed In Vivo
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-11-12 Classification: TOXIN Ligands: ZN, CA |
 |
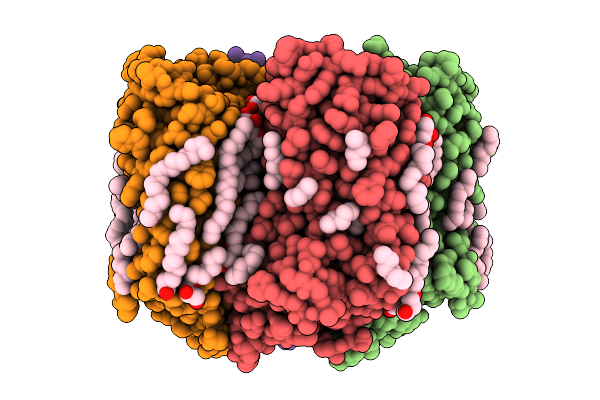
Room Temperature Structure Of Kr2 Rhodopsin In Pentameric Form At 95% Relative Humidity
Organism: Dokdonia eikasta
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: RET, NA, OLC, LFA |
 |
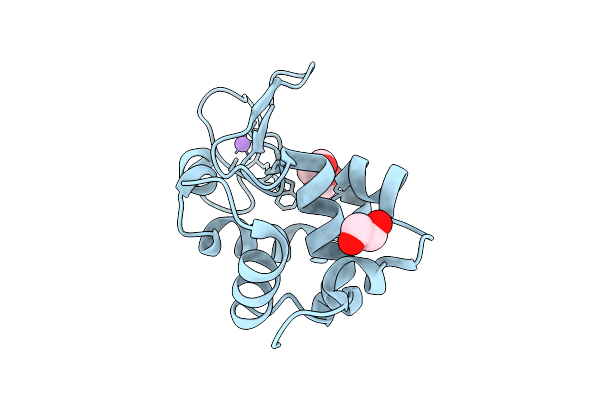
Room-Temperature Structure Of Kr2 Rhodopsin In Pentameric Form At 85% Relative Humidity
Organism: Dokdonia eikasta
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: RET, NA, OLC, LFA |
 |
Hen Egg-White Lysozyme Structure Embedded In Lcp Medium At 95% Relative Humidity
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
 |
Organism: Gallus gallus
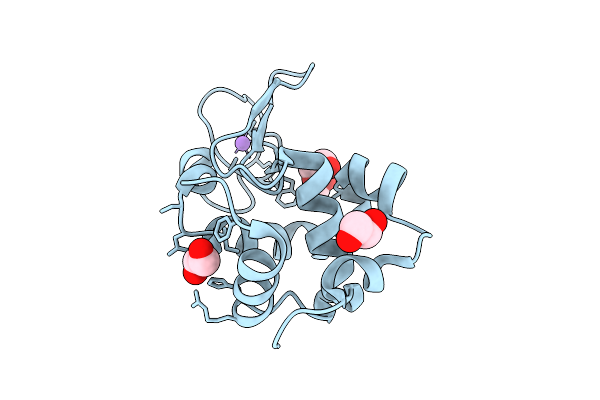
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
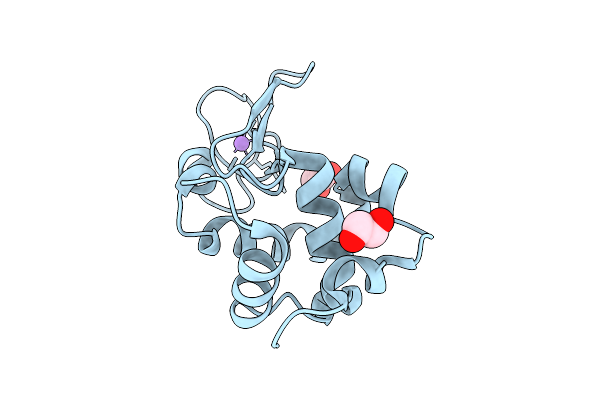 |
Hen Egg-White Lysozyme Structure Collected At Euxfel Spb/Sfx With Hve Injection Method
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
 |
Hen Egg-White Lysozyme Structure Embedded In Lcp Medium At 95% Relative Humidity
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
 |
Hen Egg-White Lysozyme Structure Embedded In Lcp Medium At 85% Relative Humidity
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
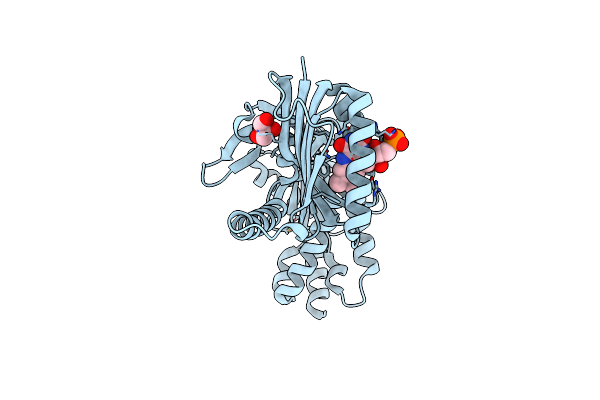 |
Organism: Oscillatoria acuminata
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-02-26 Classification: LYASE Ligands: FMN, GOL, CA |
 |
Room Temperature Structure Of Fad-Containing Ferrodoxin-Nadp Reductase From Brucella Ovis At Euxfel
Organism: Brucella ovis atcc 25840
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Room Temperature Structure Of Fad-Containing Ferrodoxin-Nadp Reductase From Brucella Ovis At Lcls
Organism: Brucella ovis atcc 25840
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (3 Picosecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (300 Ps Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (1 Nanosecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |

