Search Count: 46
All
Selected
 |
Organism: Homo sapiens
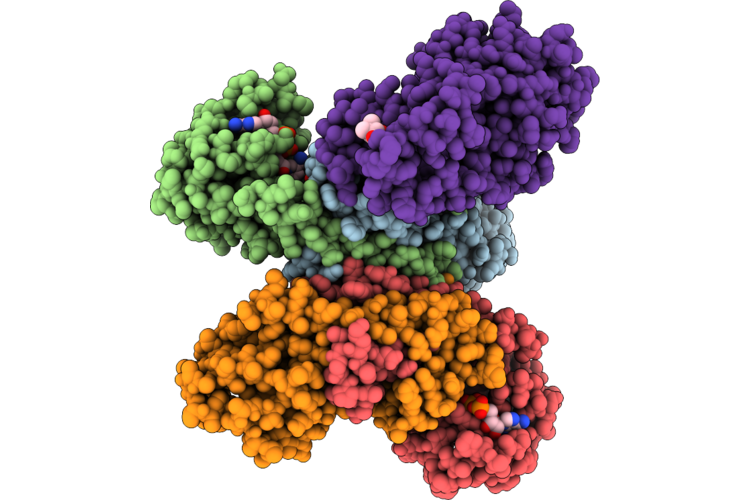
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2026-05-20 Classification: OXIDOREDUCTASE Ligands: SO4, NAD, G8I |
 |
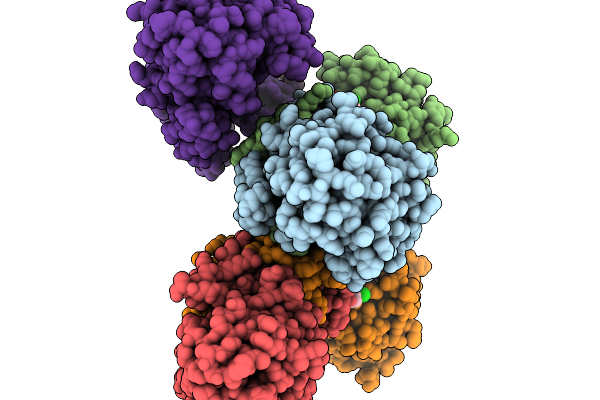
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 1-Benzyl-1H-1,3-Benzodiazole-2-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: A1CFK, SO4 |
 |
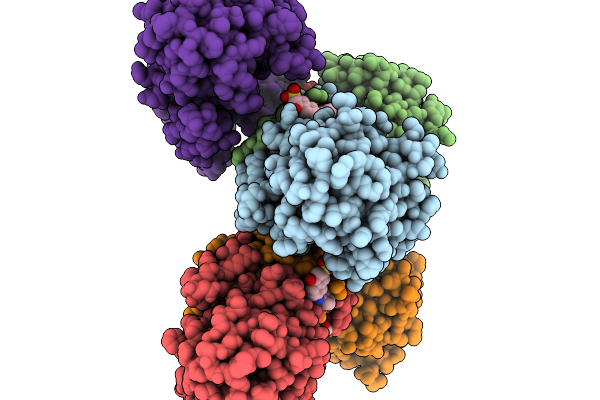
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 2-[(2,6-Dichlorophenyl)Amino]Pyridine-3-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE Ligands: A1CFJ, SO4, PEG, PGE |
 |
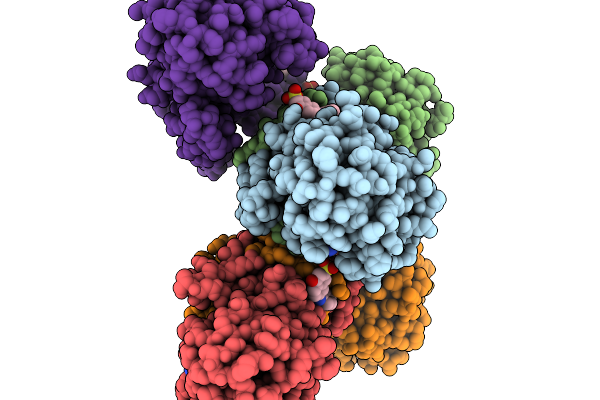
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 4-Hydroxy-7-(Phenylamino)Naphthalene-2-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: A1CFI, SO4, PGE |
 |
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 4-Hydroxy-7-(Phenylamino)Naphthalene-2-Sulfonic Acid In A Remote Site And Nadh And (S)-(-)-2-Hydroxy-3,3-Dimethylbutyric Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: G8I, A1CFI, NAI, PEG |
 |
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 7-Benzamido-4-Hydroxynaphthalene-2-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: SO4, PEG, A1CFH, PGE |
 |
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 7-Benzamido-4-Hydroxynaphthalene-2-Sulfonic Acid In A Remote Site And Nadh And (S)-(-)-2-Hydroxy-3,3-Dimethylbutyric Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: G8I, A1CFH, NAI |
 |
Quaternary Complex Of Pycr1 With The Allosteric Inhibitor 1-Benzyl-1H-1,3-Benzodiazole-2-Sulfonic Acid In A Remote Site And Nadh And (S)-Tetrahydrofuran-2-Carboxylic Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: TFB, NAI, A1CFK, PEG |
 |
Ternary Complex Of Pycr1 With The Allosteric Inhibitor 2-[(2,6-Dichlorophenyl)Amino]Pyridine-3-Sulfonic Acid In Two Remote Sites And The Active Site, And (S)-(-)-2-Hydroxy-3,3-Dimethylbutyric Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: G8I, A1CFJ, SO4 |
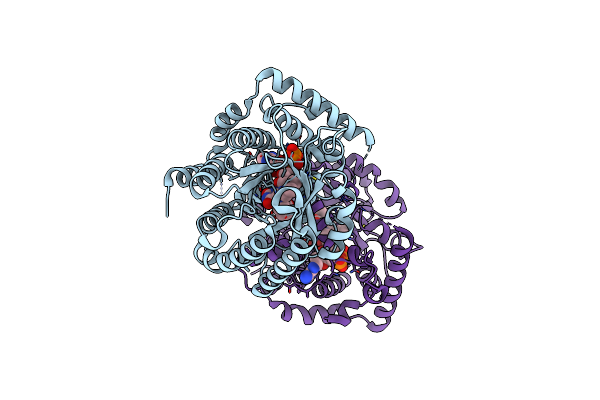 |
Minimal Puta Proline Dehydrogenase Domain (Design #2) Covalently Inactivated By (S)-But-3-Yn-2-Ylglycine
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-02-05 Classification: OXIDOREDUCTASE Ligands: A1BC6, FMT |
 |
Proline Utilization A (Puta) From Sinorhizobium Meliloti Inactivated By N-Propargylglycine
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2025-02-05 Classification: OXIDOREDUCTASE/INHIBITOR Ligands: P5F, NAD, SO4, MG, FMT, PEG, PG4, PGE |
 |
Structure Of Proline Utilization A Complexed With 2,3-Dihydro-1,4-Benzodioxine-5-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, 53E, FMT, NAD, SO4, MG, PGE, PG4 |
 |
Structure Of Proline Utilization A Complexed With Quinoline-2-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, QNC, PEG, FMT, SO4, MG, PG4 |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: PGE, PEG, PYS, FMT, SO4, FAD |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, FMT, PEG, PG4, A1BDA, SO4 |
 |
Structure Of Proline Utilization A Complexed With Piperidine-3-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, PGE, FMT, A1BDB, ID7, SO4, NAD, MG, PG4 |
 |
Structure Of Proline Utilization A Complexed With 1-(4-Fluorophenyl)Thiourea
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, PGE, 8P4, FMT, SO4 |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, A1BDD, PGE, PG4, FMT, NAD, SO4, MG |
 |
Structure Of Proline Utilization A Co-Crystallized With 4-Methoxybenzyl Alcohol
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, A1BDD, PGE, FMT, PEG, NAD, SO4, MG, 1PE |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, FMT, PEG, NAD, SO4, MG, PGE, A1BDW |

