Search Count: 551
All
Selected
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: XDC, ZN, MG, CA, GOL, NA, NAG, MAN, PO4, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN, MG, CA, NA, NAG, GOL, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Tissue Non-Specific Alkaline Phosphatase -S110A Bound To Ppi With Ethylene Glycol
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: POP, ZN, MG, CA, NA, NAG, MAN, PO4, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ATP, ZN, MG, CA, GOL, NAG, FLC, PO4 |
 |
Organism: Synthetic construct, Aequorea victoria
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: FLUORESCENT PROTEIN |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2026-02-25 Classification: SUGAR BINDING PROTEIN Ligands: EOH |
 |
Organism: Synthetic construct, Methanosarcina mazei
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: SUGAR BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.46 Å Release Date: 2026-01-14 Classification: ONCOPROTEIN/INHIBITOR Ligands: ANP, MG, SO4, A1BIS |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-14 Classification: TRANSFERASE Ligands: A1IR8 |
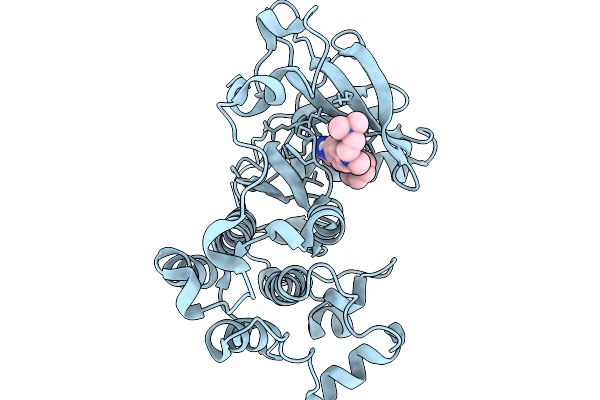 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.99 Å Release Date: 2026-01-14 Classification: TRANSFERASE Ligands: A1IR9 |
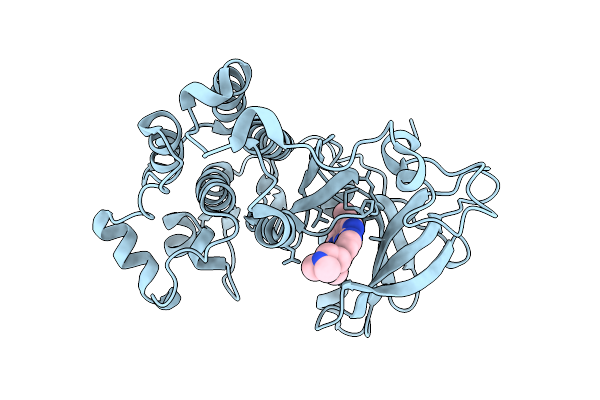 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-01-14 Classification: TRANSFERASE Ligands: A1JLU |
 |
Organism: Severe acute respiratory syndrome coronavirus
Method: SOLUTION NMR Release Date: 2025-10-01 Classification: RNA |
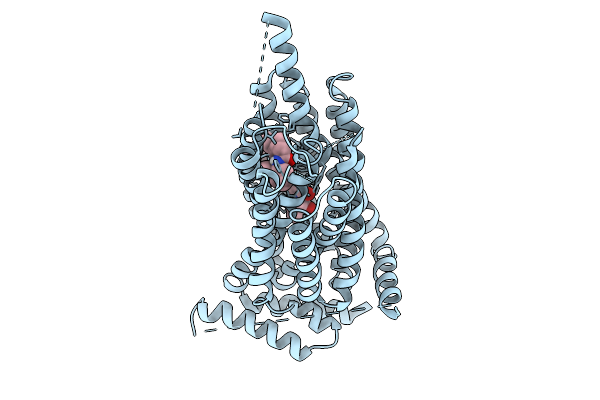 |
Structure Of Synaptic Vesicle Protein 2A Bound To Brivaracetam And Ucb1244283
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN Ligands: A1B1J, VLX, CLR, A1B1I |
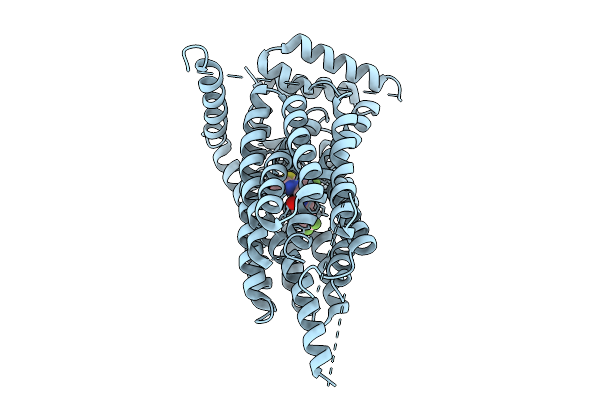 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN Ligands: A1CHN |
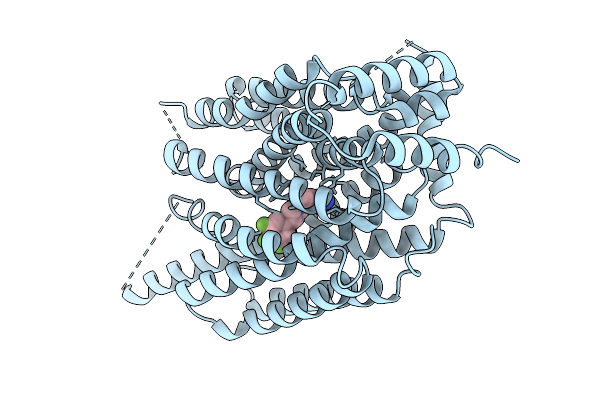 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN Ligands: A1CCA |
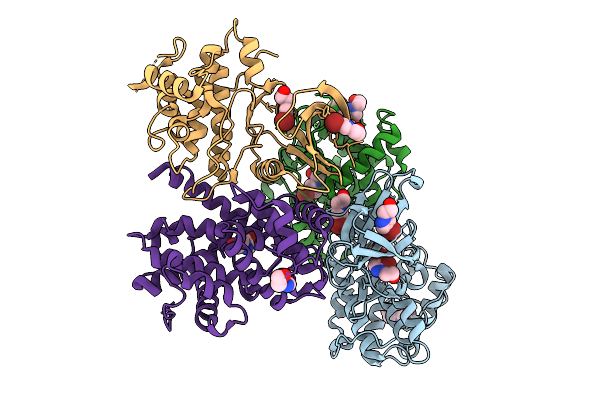 |
Organism: Homo sapiens, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2025-09-03 Classification: CELL CYCLE Ligands: UU1 |
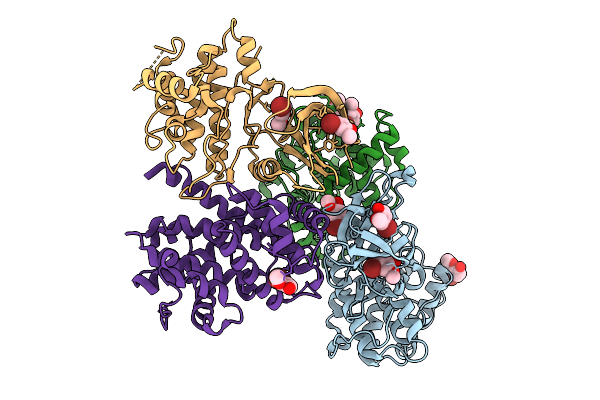 |
Organism: Homo sapiens, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-09-03 Classification: CELL CYCLE Ligands: 3Z7 |
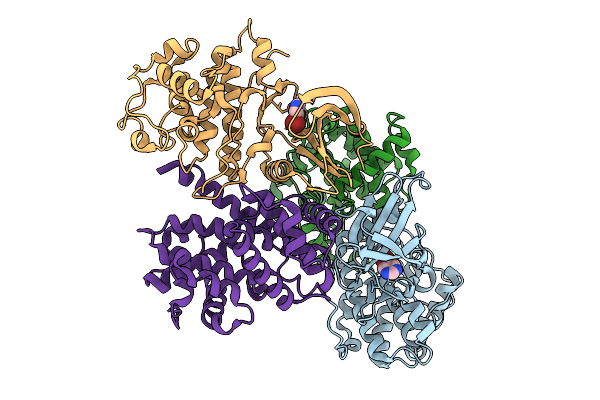 |
Organism: Homo sapiens, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2025-09-03 Classification: CELL CYCLE Ligands: HHN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2025-07-16 Classification: TRANSFERASE Ligands: A1IHC |

