Search Count: 1,436
All
Selected
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And Imidazole
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: IMD, MN |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 2,5,6-Triaminopyrimidin-4(3H)-One.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: A1L8V, IMD, MN |
 |
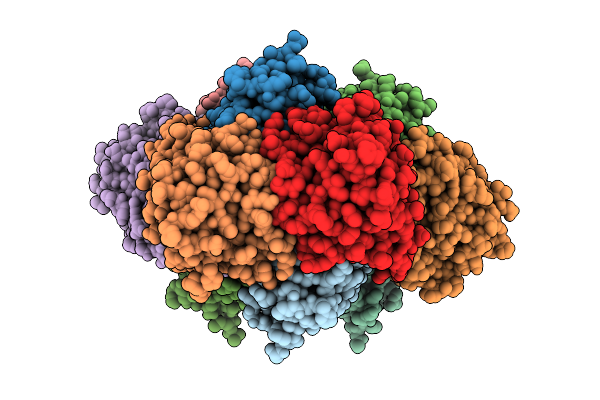
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GOL, A1L8W, MN, IMD |
 |
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Tpp.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-01 Classification: LYASE Ligands: ZN, TPP, MG, GOL |
 |
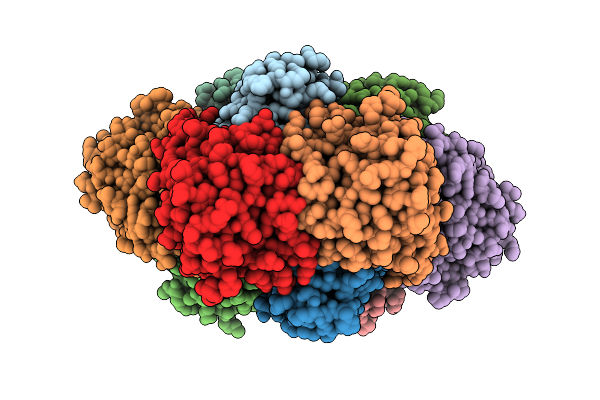
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Ttpp And 6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-04-01 Classification: LYASE Ligands: A1L8W, ZN, SO4, TZD, MG |
 |
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Tpp And 5-Azacytosine.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2026-04-01 Classification: LYASE Ligands: 5AZ, ZN, TPP, MG, GOL |
 |
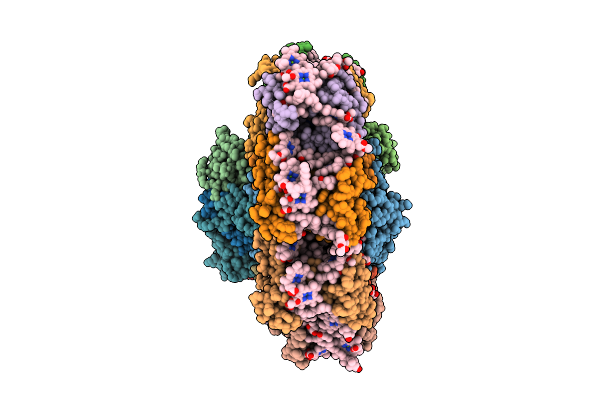
2,2'-Bipyridine Linked Dna Oligomer (Cgcgaat[Bru]Cgcg) In The Presence Of Ni2+ Ion
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-04-01 Classification: DNA |
 |
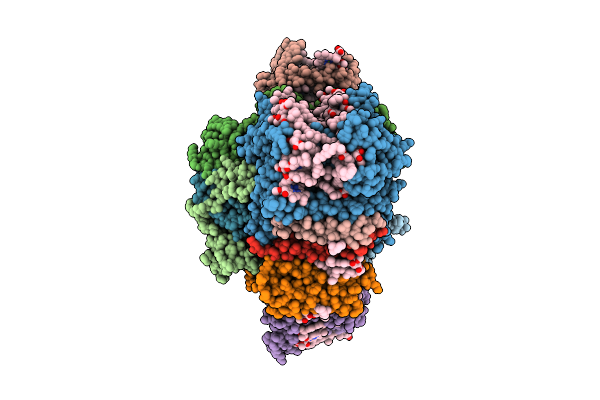
Organism: Chrysotila roscoffensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, LMU, CL0, SF4, DGD, SQD, DD6, LMG, KC1, A86 |
 |
Organism: Chrysotila roscoffensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, DD6, LMU, CL0, SF4, DGD, LMG, A86, KC1 |
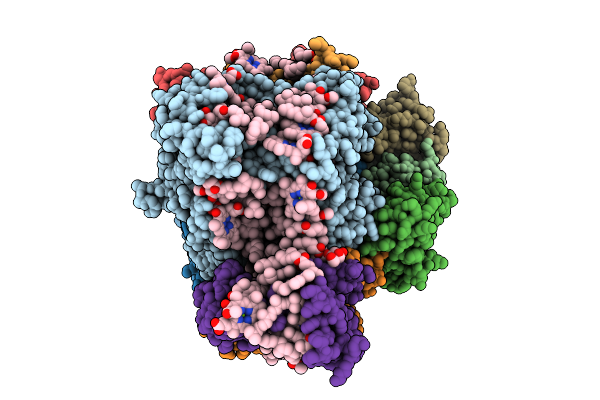 |
Organism: Chrysotila roscoffensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, CL0, SF4, DGD, DD6, LMG, A86, KC1 |
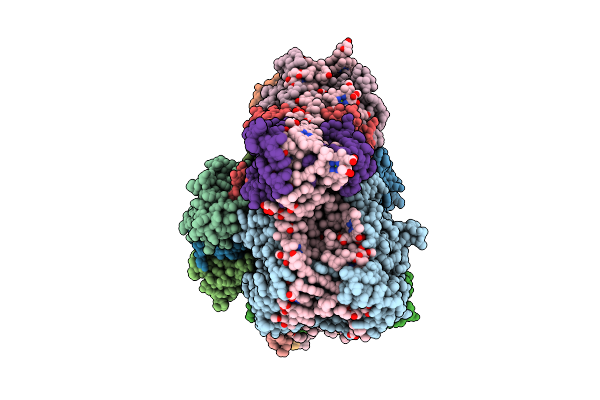 |
Organism: Chrysotila roscoffensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, CL0, SF4, DD6, DGD, SQD, LMG, KC1, A86 |
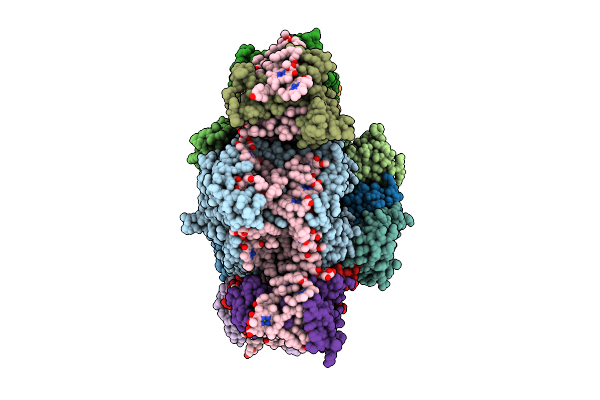 |
Organism: Chrysotila roscoffensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, CL0, SF4, DD6, DGD, SQD, LMG, KC1, A86 |
 |
Organism: Streptomyces prunicolor
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GOL, PEG |
 |
Aminoacyl-Trna-Dependent Peptide Synthase, Sbb17, Complexed With Streptothrisamine
Organism: Streptomyces prunicolor
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: A1L3W |
 |
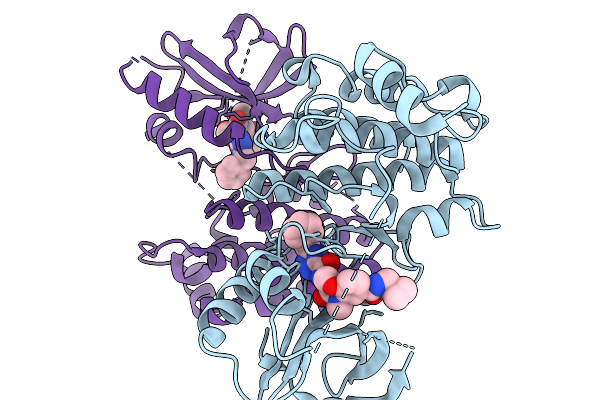
Aminoacyl-Trna-Dependent Peptide Synthase, Sba18, Complexed With Streptothrisamine
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: A1L3W, TRS |
 |
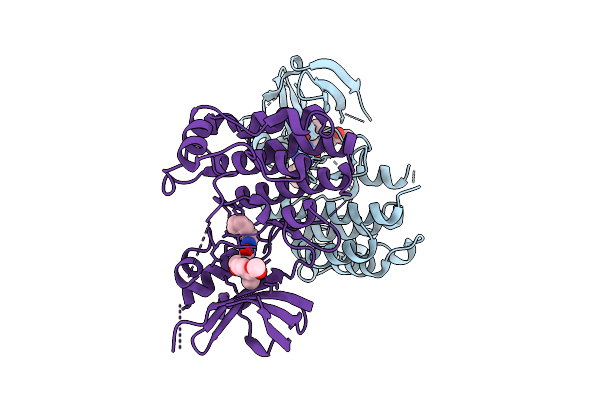
Crystal Structure Of Human Ripk1 Kinase Domain In Complex With Compound Hr97
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: A1E6R |
 |
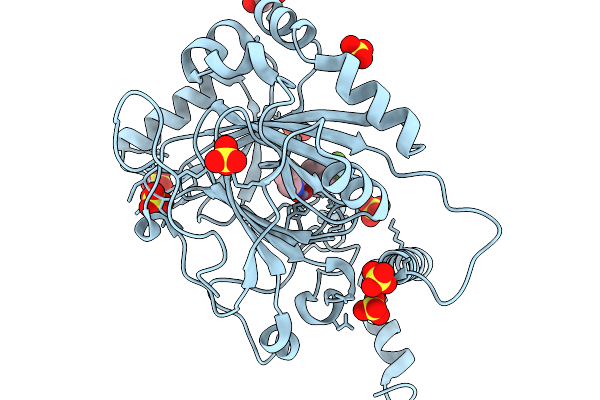
Crystal Structure Of Human Ripk1 Kinase Domain In Complex With Compound Hr10
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: A1E6Q |
 |
Fih [Factor Inhibiting Hif (Hypoxia Inducible Factor)] In Complex With Zn(Ii) And N-((4-(Trifluoromethyl)Phenyl)Sulfonyl)Picolinamide (Sb133)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: GOL, ZN, A1L8D, SO4 |
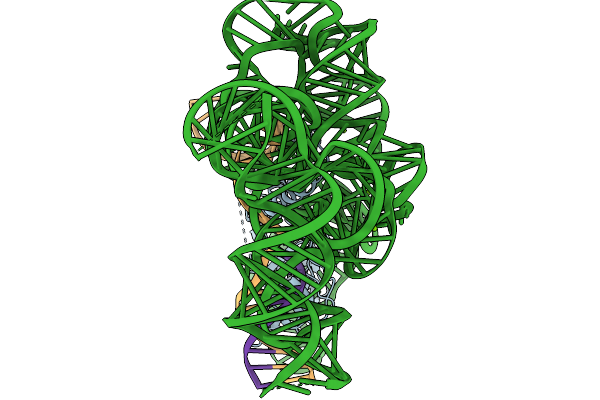 |
Organism: Metagenome, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN Ligands: MG |
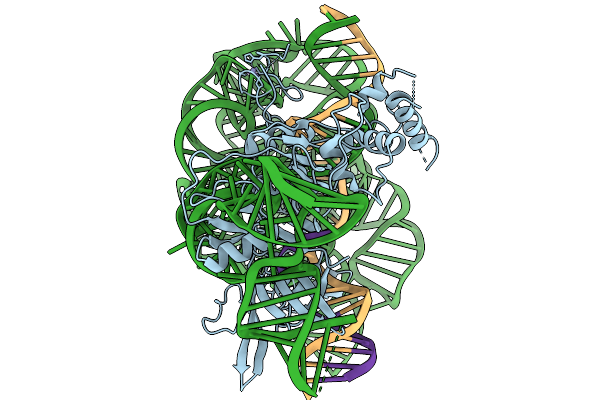 |
Organism: Tissierellia bacterium, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |

