Search Count: 455
All
Selected
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: VIRUS Ligands: ZN, PO4 |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PEG, DSN, CA |
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-04-22 Classification: HYDROLASE |
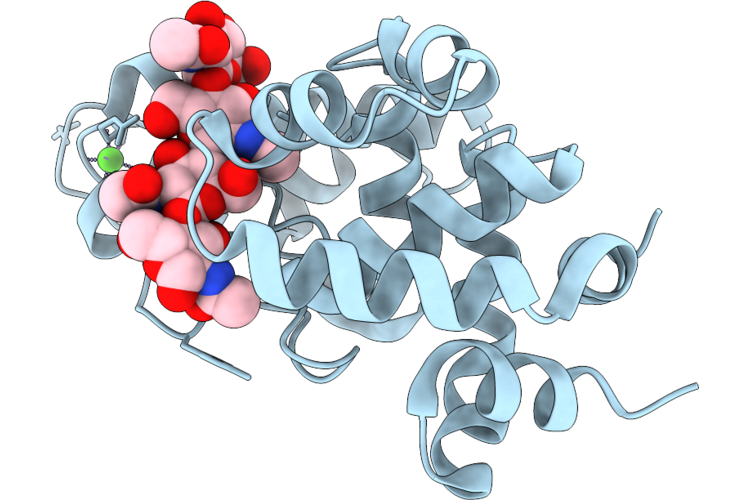 |
Crystal Structure Of Gh19 E228Q Domain Of D29-Lysa In Complex With (Glcnac)4
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, MPD |
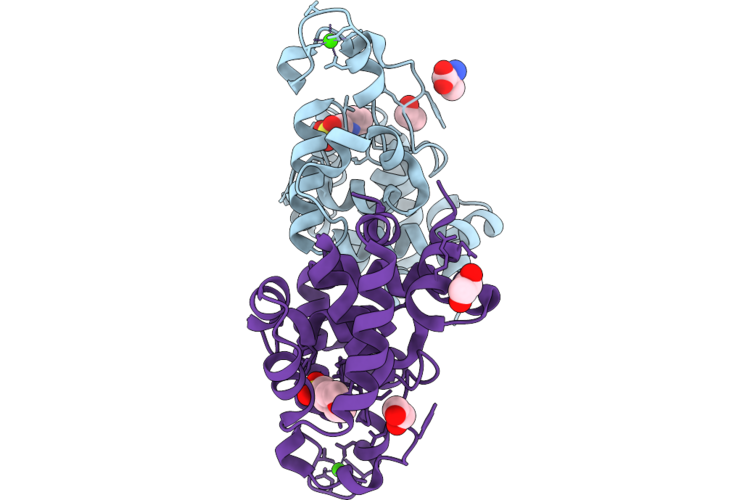 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, EDO, ALA |
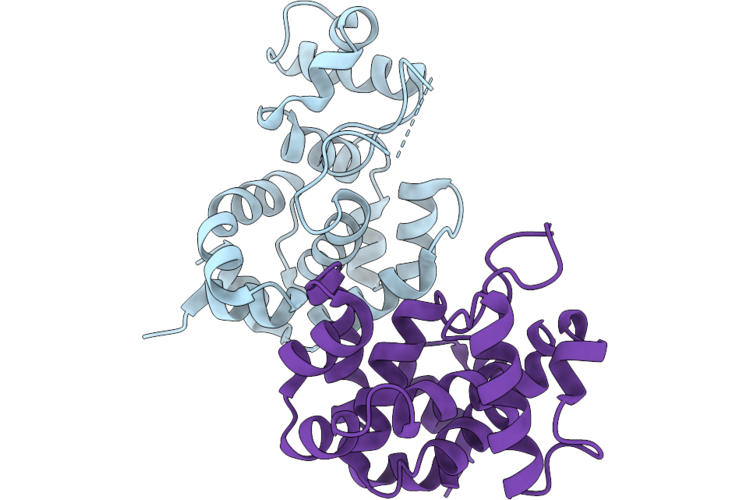 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: HYDROLASE |
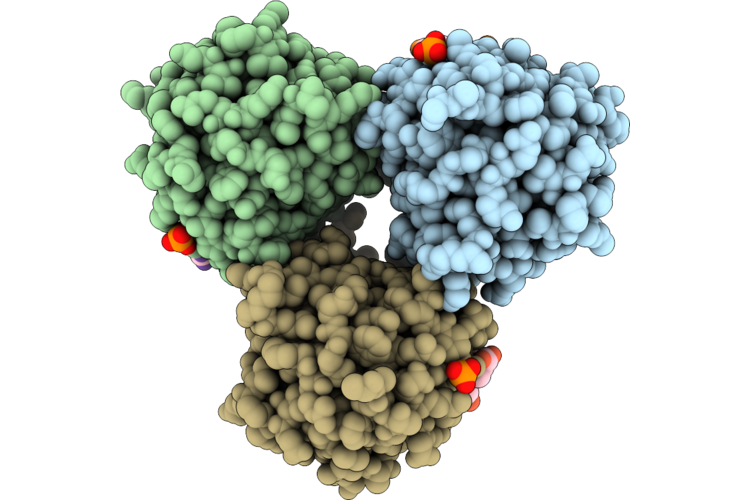 |
Crystal Structure Of Ami2B Domain Of Ds6A-Lysa In Complex With L-Ala-D-Iso-Gln-L-Lys- D-Ala-D-Ala
Organism: Mycobacterium phage ds6a, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN |
 |
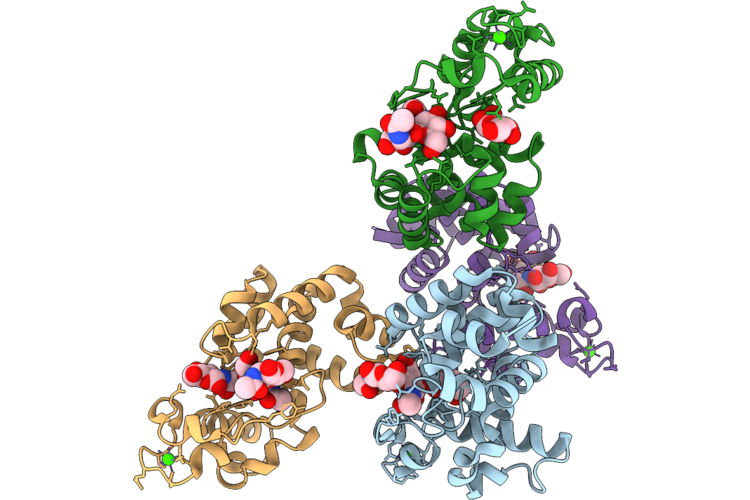
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Product (Glcnac)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG |
 |
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Products (Glcnac Y Glcnac2)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG, CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-04-22 Classification: HYDROLASE |
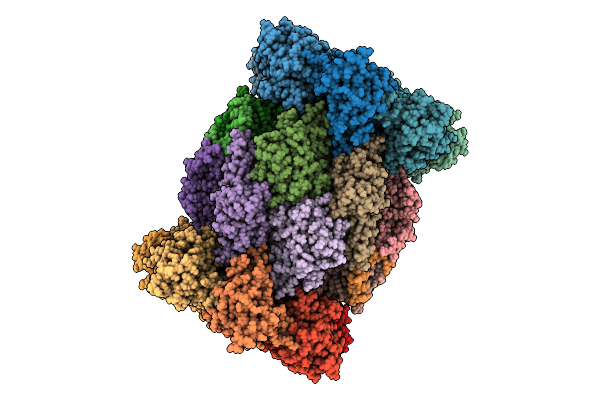 |
Structure Of The Plasmodium Falciparum 20S Proteasome In Complex With A Beta5-Selective Covalent Syringolin Analogue Inhibitor.
Organism: Plasmodium falciparum 3d7
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: A1CY6 |
 |
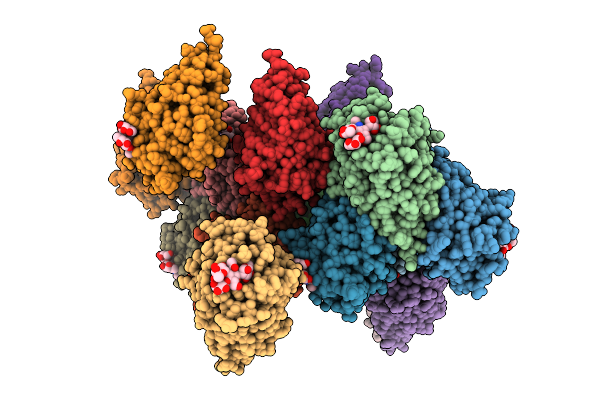
Structure Of The Human 20S Proteasome In Complex With A Beta5-Selective Covalent Syringolin Analogue Inhibitor.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: A1CY5 |
 |
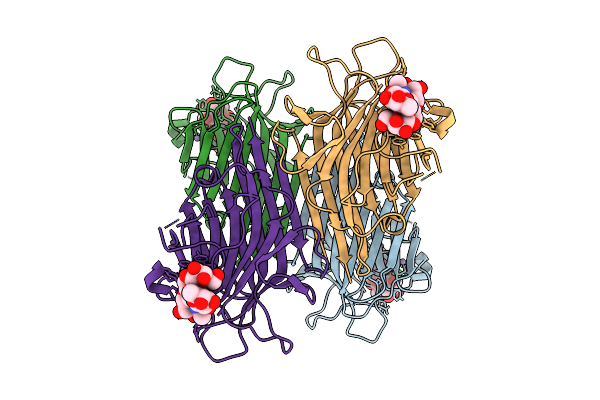
Organism: Lablab purpureus
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: CA, MN |
 |
Organism: Lablab purpureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: CA, MN |
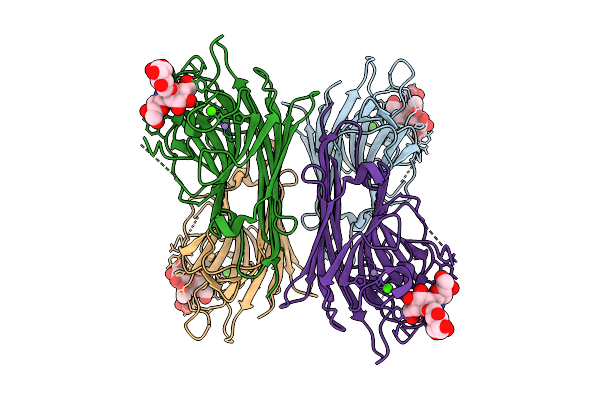 |
Organism: Lablab purpureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: CA, MN |
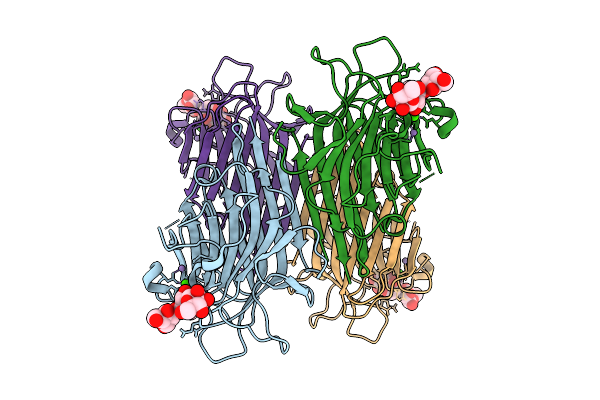 |
Organism: Lablab purpureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: CA, MN |
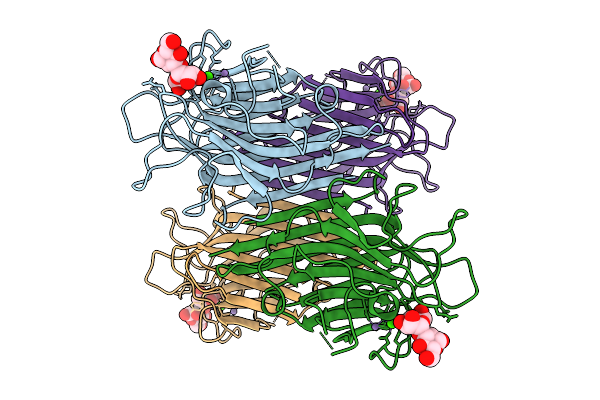 |
Organism: Lablab purpureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: CA, MN |
 |
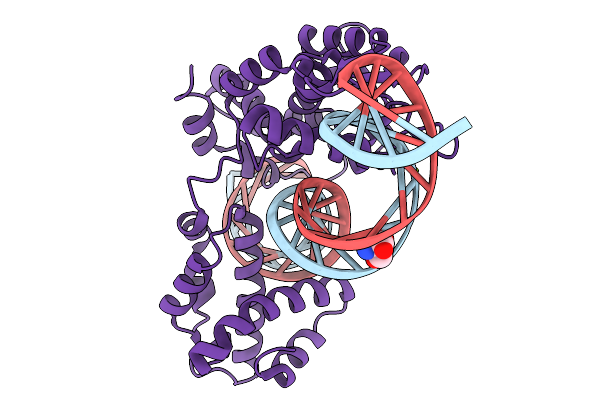
Arbitrium Controls Lysis-Lysogeny Through The Activation Of A Small Antirepressor Protein In The Majority Of Arbitrium-Coding Phages
Organism: Bacillus phage vb_bts_bmbtp14
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2026-01-14 Classification: DNA BINDING PROTEIN Ligands: SO4 |
 |
Organism: Bacillus phage phi3t
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-01-14 Classification: DNA BINDING PROTEIN Ligands: TRS, MG |

