Search Count: 79
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: PROTEIN FIBRIL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: PROTEIN FIBRIL |
 |
Pentameric Structure Of Mers-Cov Envelope Protein Transmembrane Domain Determined By Solid-State Nmr
Organism: Betacoronavirus england 1
Method: SOLID-STATE NMR Release Date: 2025-09-24 Classification: VIRAL PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-04-16 Classification: CELL CYCLE Ligands: A1A1H |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2025-04-16 Classification: CELL CYCLE Ligands: A1A1H, PEG, GOL, EDO |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: CELL CYCLE Ligands: ZN, A1A1I |
 |
Cryo-Em Structure Of Cdk2/Cycline1 In Complex With Crbn/Ddb1 And Cpd 4 (Local Mask)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: CELL CYCLE Ligands: ZN, A1A1I |
 |
Activation Mechanism And Novel Binding Site Of The Bkca Channel Activator Ctibd
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-31 Classification: MEMBRANE PROTEIN Ligands: A1L04, Y01, MG, CA |
 |
Organism: Homo sapiens, Human adenovirus c serotype 2
Method: ELECTRON MICROSCOPY Release Date: 2024-07-10 Classification: MEMBRANE PROTEIN Ligands: NAG, 65I |
 |
De Novo Designed Protein Binds Poly Adp Ribose Polymerase Inhibitors (Parpi) - Apo
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2024-04-24 Classification: DE NOVO PROTEIN Ligands: SO4 |
 |
De Novo Designed Protein Binds Poly Adp Ribose Polymerase Inhibitors (Parpi) - Holo Rucaparib
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2024-04-24 Classification: DE NOVO PROTEIN Ligands: RPB |
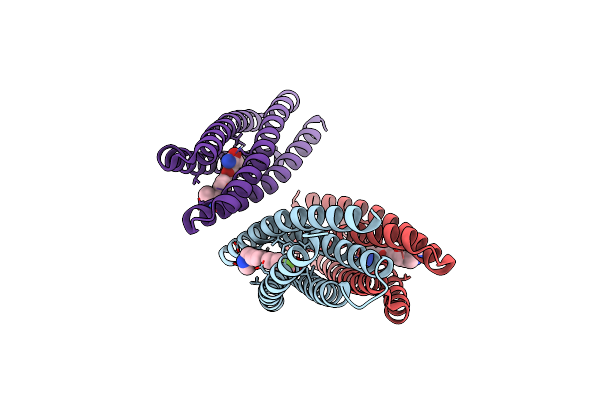 |
De Novo Designed Protein Binds Poly Adp Ribose Polymerase Inhibitors (Parpi) - Holo Mefuparib
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2024-04-24 Classification: DE NOVO PROTEIN Ligands: IQM |
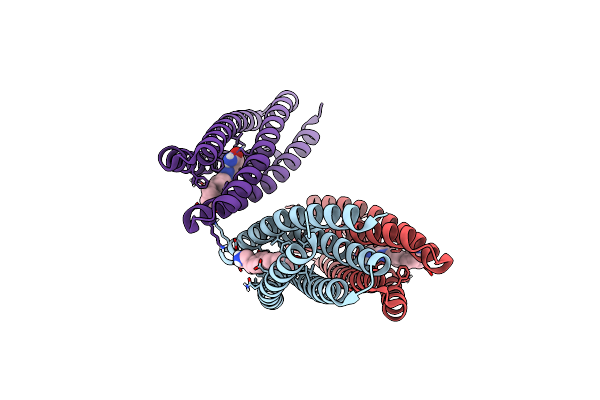 |
De Novo Designed Protein Binds Poly Adp Ribose Polymerase Inhibitors (Parpi) - Holo Niraparib
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2024-04-24 Classification: DE NOVO PROTEIN Ligands: 3JD |
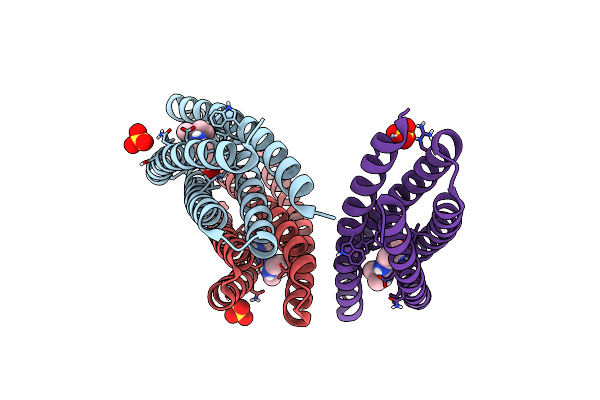 |
De Novo Designed Protein Binds Poly Adp Ribose Polymerase Inhibitors (Parpi) - Holo Veliparib
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.29 Å Release Date: 2024-04-24 Classification: DE NOVO PROTEIN Ligands: 78P, SO4 |
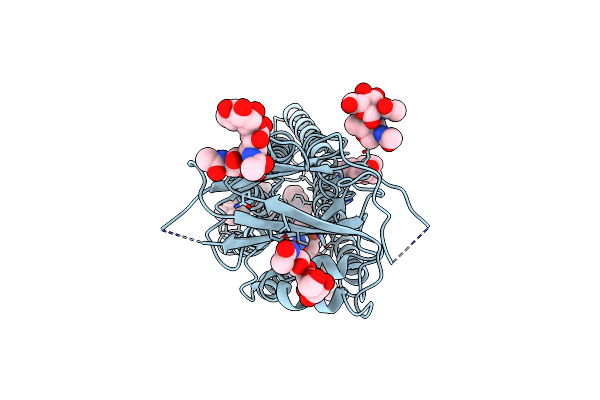 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-12-27 Classification: MEMBRANE PROTEIN Ligands: L9Q, CLR |
 |
Organism: Homo sapiens, Human adenovirus 2
Method: ELECTRON MICROSCOPY Release Date: 2023-12-27 Classification: MEMBRANE PROTEIN Ligands: GCO |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2023-11-22 Classification: LIGASE Ligands: FAD, TPP, MG, UQ0 |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2023-11-22 Classification: LIGASE Ligands: FAD, TPP, MG, UQ0 |
 |
Crystal Structure Of Escherichia Coli Glyoxylate Carboligase Double Mutant In Complex With Glycolaldehyde
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2023-11-22 Classification: LIGASE Ligands: MG, FAD, TPP, DW3, UQ1 |
 |
Crystal Structure Of Escherichia Coli Glyoxylate Carboligase Quadruple Mutant
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2023-11-22 Classification: LIGASE Ligands: FAD, TPP, MG, UQ0 |

