Search Count: 301
All
Selected
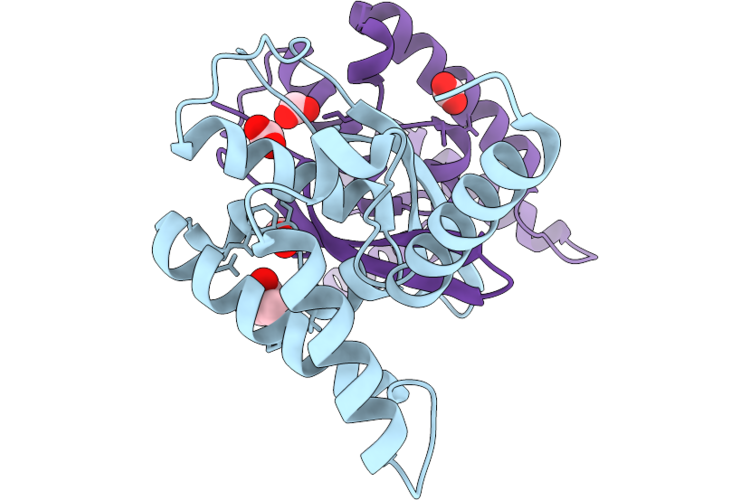 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: FMT, GOL |
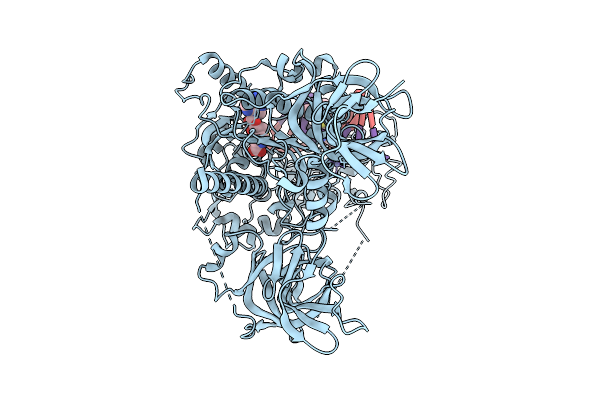 |
Human Dnmt1 (Aa 698-1616) In Complex With Hemimethylated Dsdna And Inhibitor Dmt207
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: ZN, SAH, A1EQ2 |
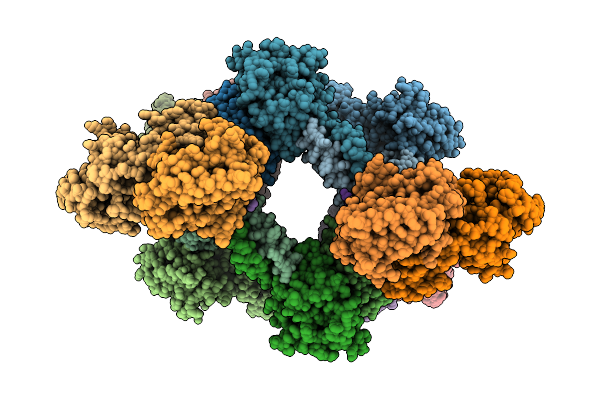 |
Organism: Enterococcus faecalis
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN Ligands: CA |
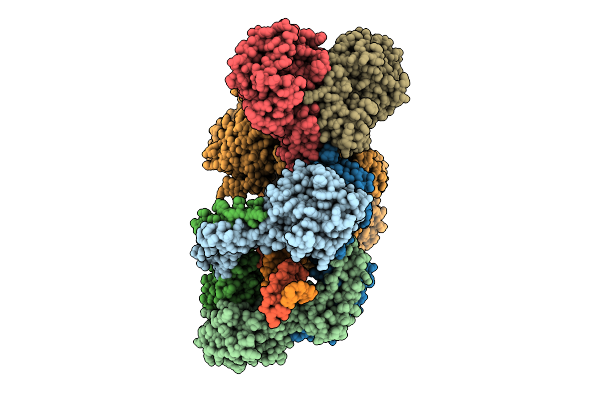 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
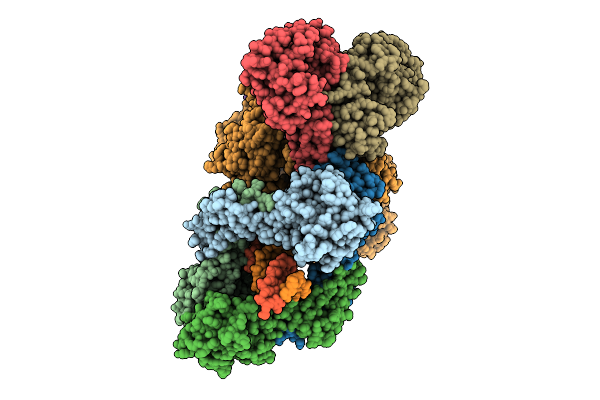 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
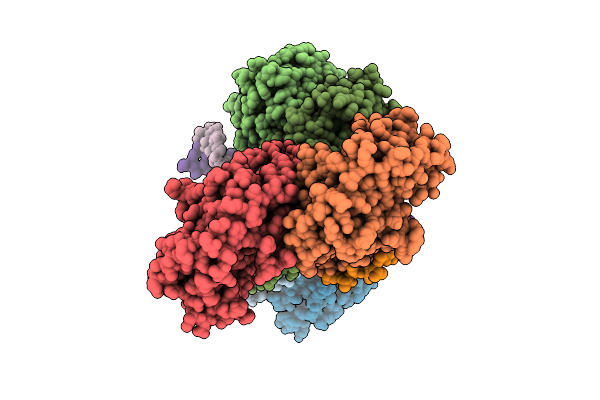 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.76 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA Ligands: CA |
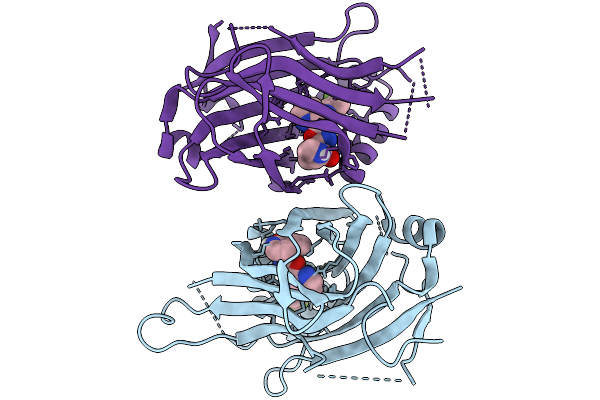 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.13 Å Release Date: 2026-03-04 Classification: TRANSCRIPTION Ligands: A1BXX |
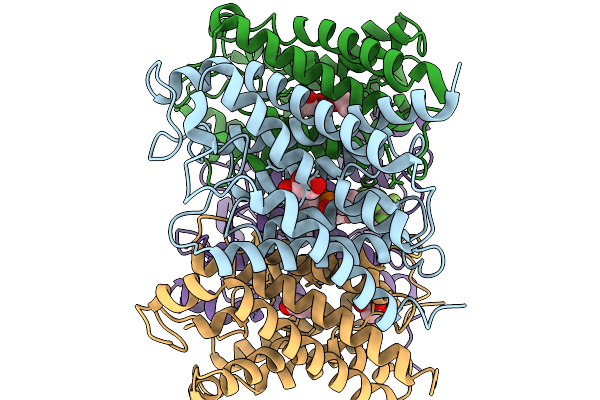 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: GOL, A1H8K |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN |
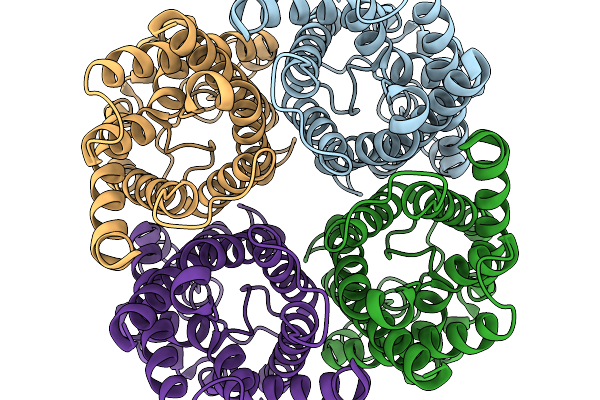 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: PEO |
 |
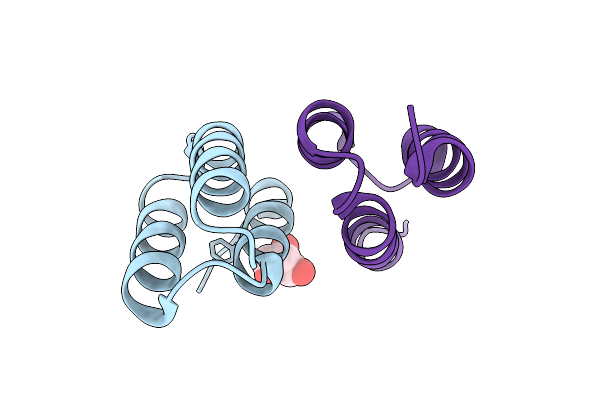
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
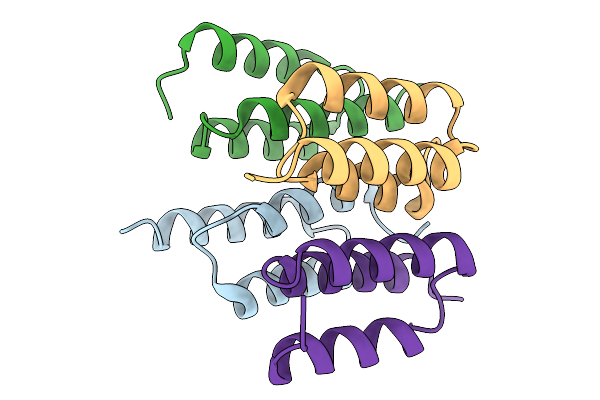
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING Ligands: GOL |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
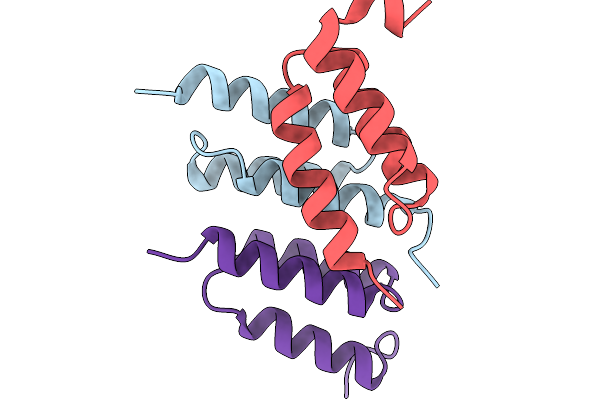
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING Ligands: SO4 |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING |
 |
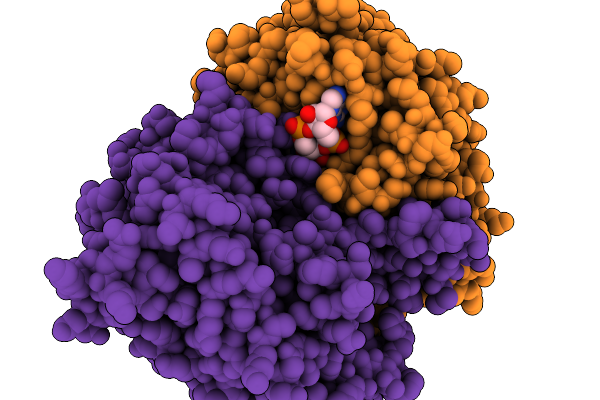
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-04 Classification: PROTEIN BINDING Ligands: EPE |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: HYDROLASE Ligands: 4BW |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2025-11-19 Classification: HYDROLASE |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-11-19 Classification: HYDROLASE |

