Search Count: 198
All
Selected
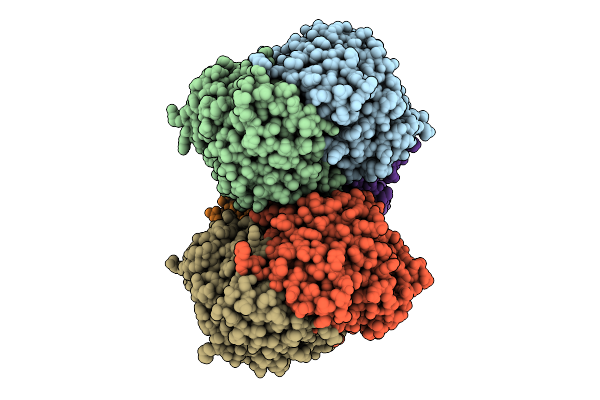 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
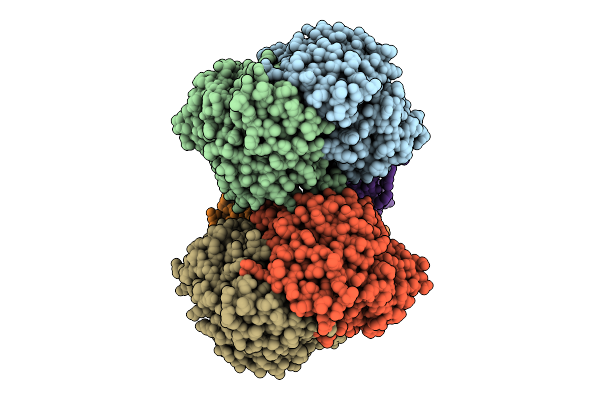 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM |
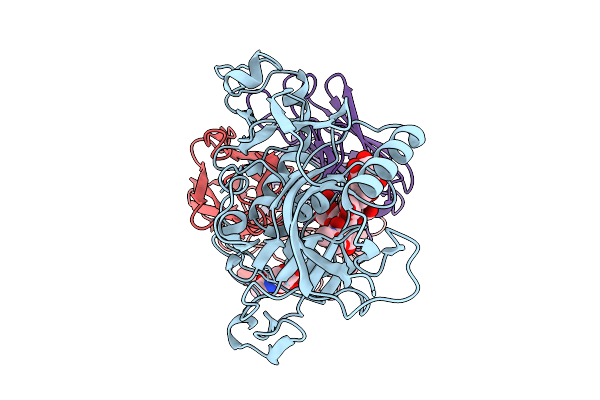 |
Organism: Severe fever with thrombocytopenia syndrome virus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
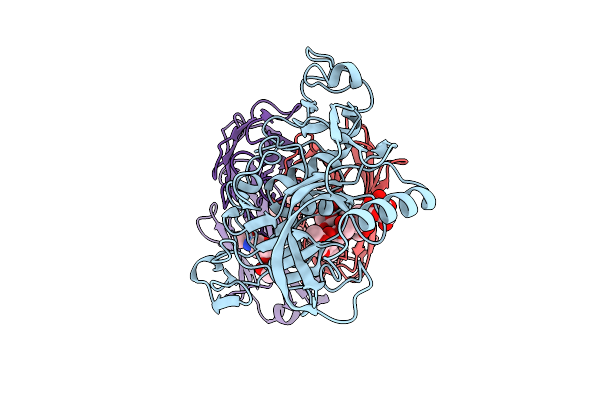 |
Organism: Severe fever with thrombocytopenia syndrome virus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN |
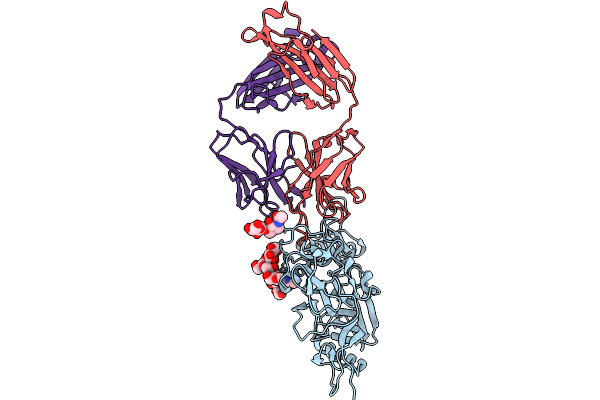 |
Organism: Severe fever with thrombocytopenia syndrome virus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
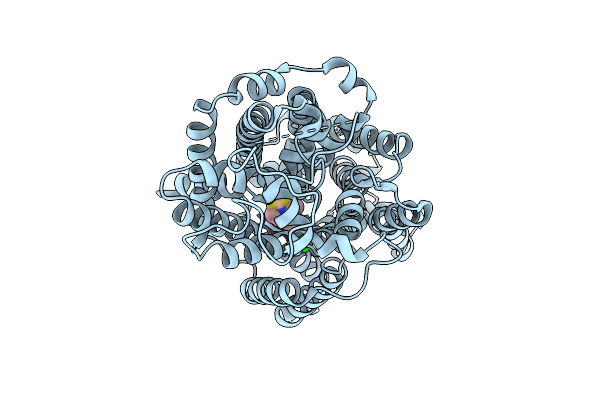 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1EBN, CL |
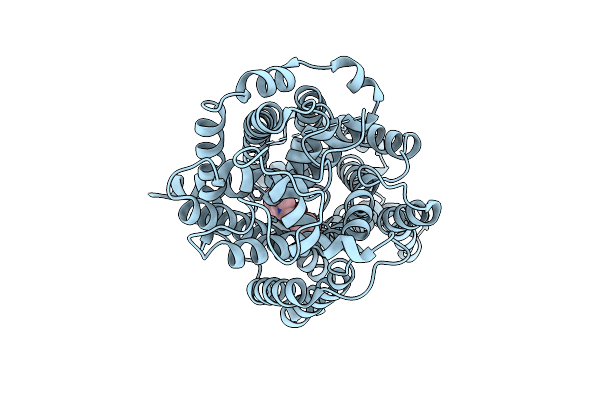 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1EBO, CL |
 |
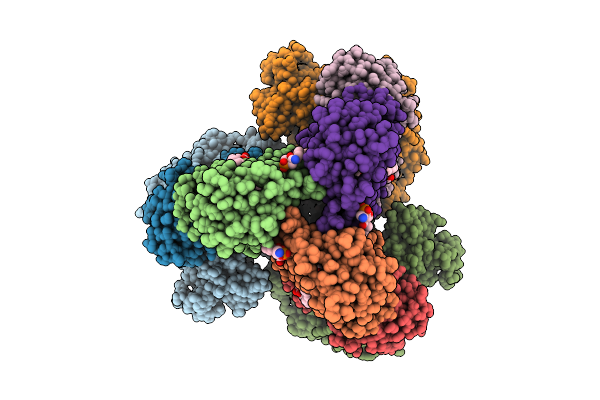
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: MOTOR PROTEIN Ligands: ADP, ATP, ZN |
 |
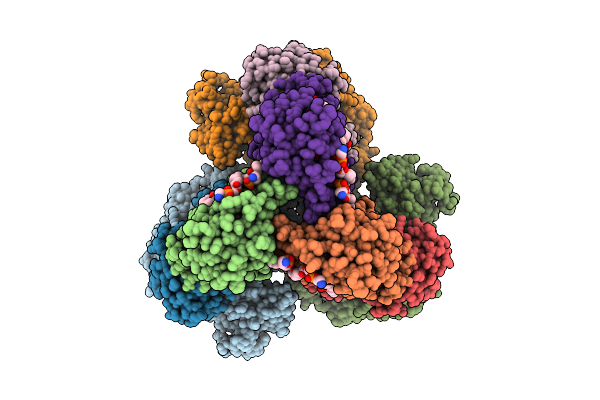
Cryo-Em Structure Of The Apo-Form Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
 |
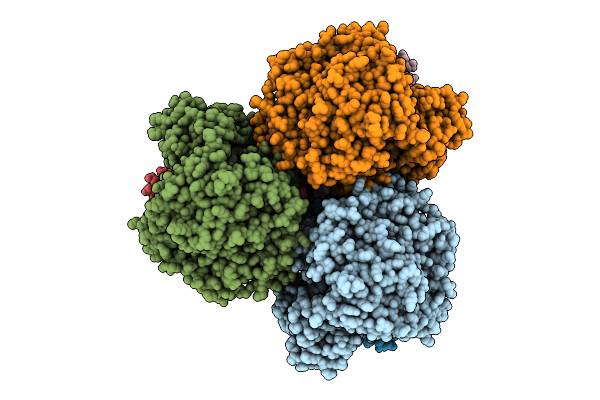
Cryo-Em Structure Of The Lipid-Bound Succiante Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
 |
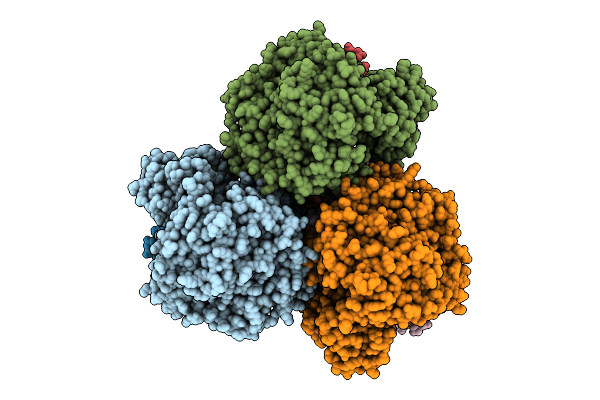
Cryo-Em Structure Of The Mk7-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, MQ7 |
 |
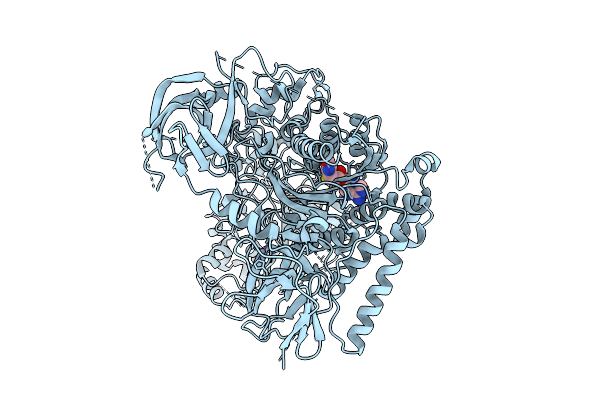
Cryo-Em Structure Of The Mk4-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, 1L3 |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: TRANSFERASE Ligands: SAH, ZN |
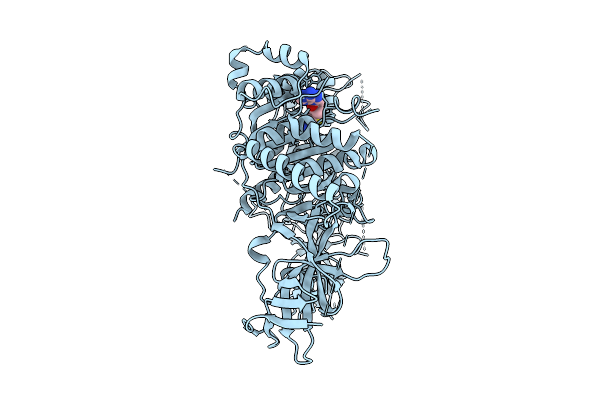 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: TRANSFERASE Ligands: ZN, SAH |
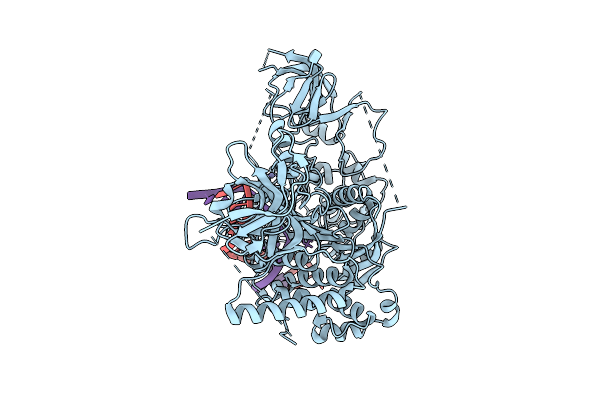 |
Organism: Arabidopsis thaliana, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: TRANSFERASE/DNA Ligands: ZN |
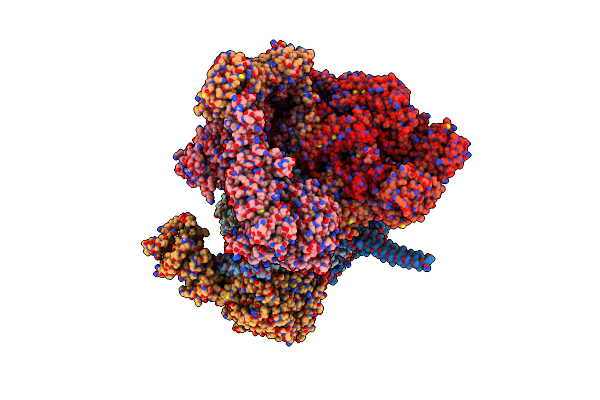 |
Structure Of Human Substrate-Free 26S Proteasome In The Presence Of Atpgs And Mg-132,Sa-Like State (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: ATP, MG, ADP, LDZ, ZN |
 |
Structure Of Substrate Engaged Midn-Bound Human 26S Proteasome, Eb-Midn (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: LDZ, ATP, MG, ADP, ZN |
 |
Structure Of Substrate Engaged Midn-Bound Human 26S Proteasome, Eb Midn_Ubl State (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: ATP, MG, ADP, ZN |

