Search Count: 120
All
Selected
 |
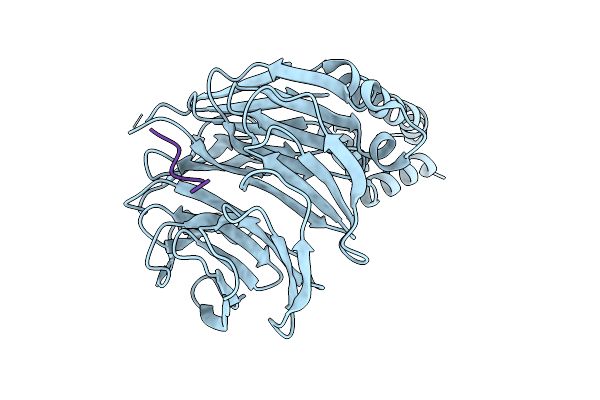
Crystal Structure Of Beta-Trcp Bound By Monophosphorylated Nrf2 Degron Peptide.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-08 Classification: LIGASE |
 |
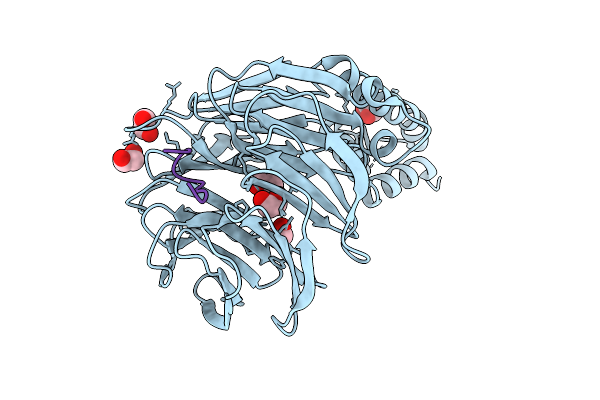
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated Nrf2 Degron Peptide.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-08 Classification: LIGASE |
 |
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated I-Kappa-B-Alpha Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.16 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: EDO |
 |
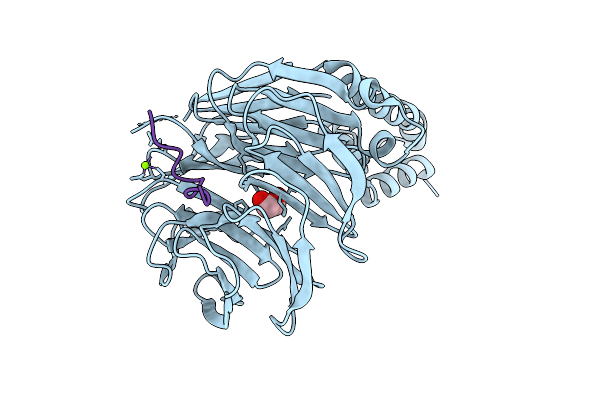
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated Claspin Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: EDO, MG |
 |
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated Pdcd4 Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: MG, EDO |
 |
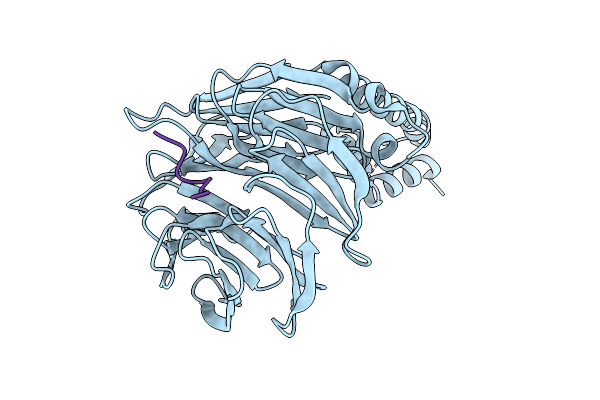
Crystal Structure Of Beta-Trcp Bound By Monophosphorylated Atf4 Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Crystal Structure Of Beta-Trcp Bound By Monophosphorylated Wee1 Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2024-08-14 Classification: TRANSFERASE Ligands: A1H8A, MTA, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-08-14 Classification: TRANSFERASE Ligands: A1H76, MTA, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-08-14 Classification: TRANSFERASE Ligands: A1H8B, MTA, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-08-14 Classification: TRANSFERASE Ligands: A1H73, MTA, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-07-03 Classification: HYDROLASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.73 Å Release Date: 2024-07-03 Classification: HYDROLASE Ligands: MG, A1IA7 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2024-07-03 Classification: HYDROLASE Ligands: MG, EOH, A1IA8 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2024-06-19 Classification: HYDROLASE Ligands: A1IA5, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.91 Å Release Date: 2024-06-19 Classification: HYDROLASE Ligands: A1IA6, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.47 Å Release Date: 2024-06-19 Classification: HYDROLASE Ligands: MG, A1IA4 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-08-23 Classification: MEMBRANE PROTEIN |

