Search Count: 422
All
Selected
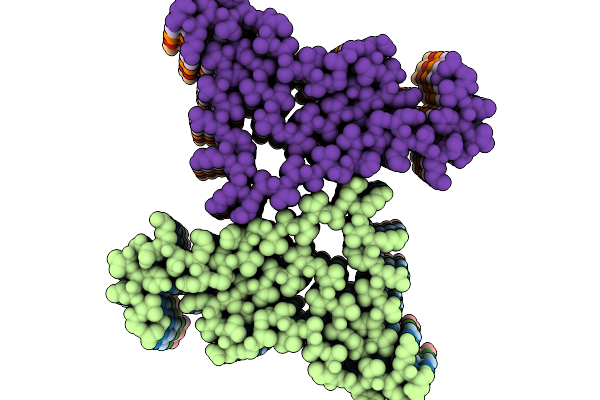 |
Structure Of Amplified Asyn Filament By Using Seed Amplification Assay (Saa) From Msa Patient Csf.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.18 Å Release Date: 2026-02-25 Classification: PROTEIN FIBRIL |
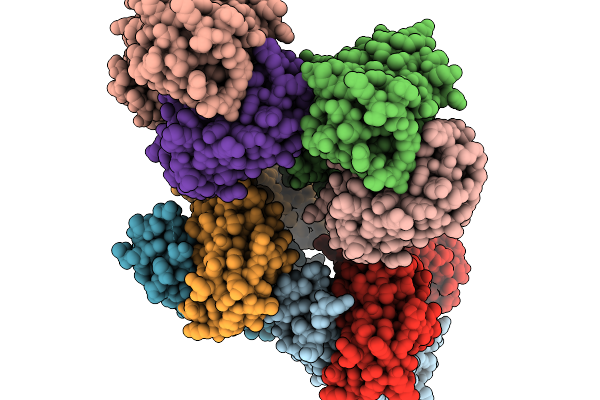 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM |
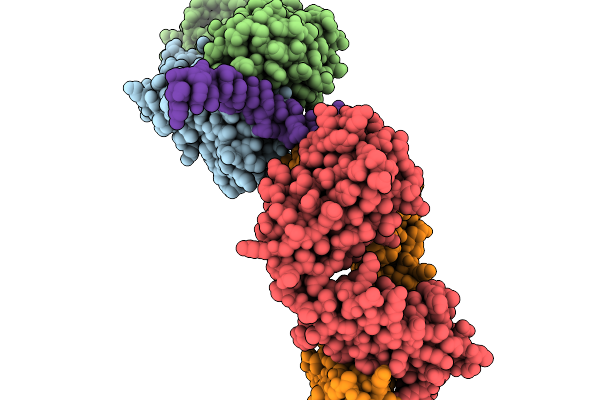 |
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.27 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM |
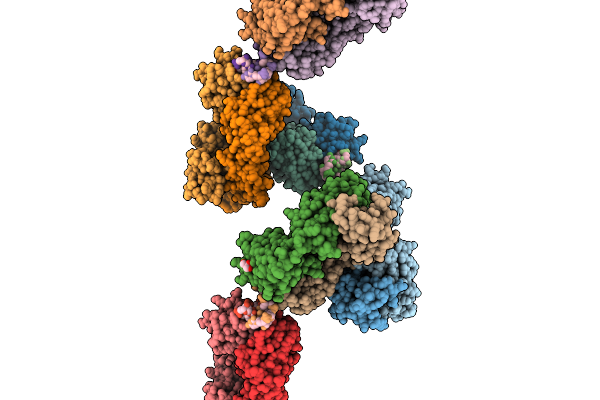 |
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM Ligands: GOL |
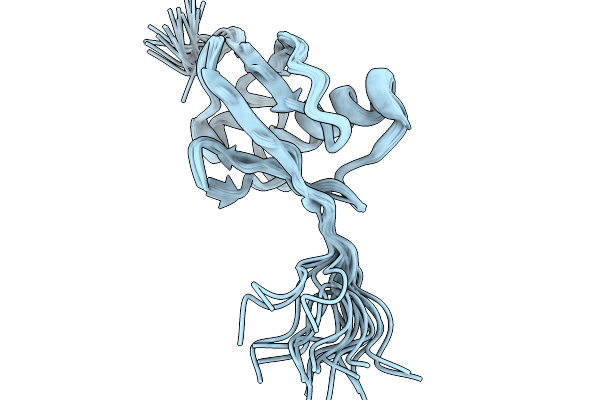 |
Nmr Structure Of Proteinmpnn-Desighed Ubiquitin Variant R4 At Ph 3 With 8 M Urea
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
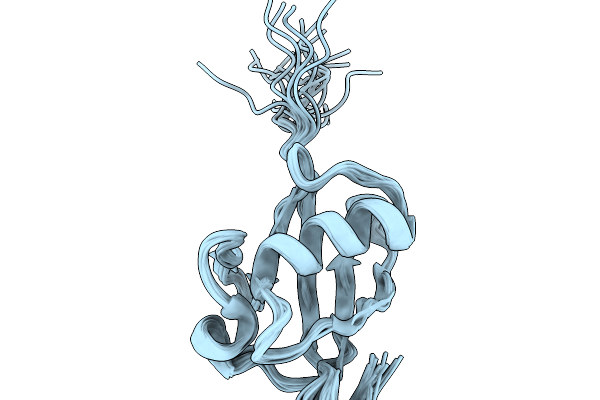 |
Nmr Structure Of Proteinmpnn-Desighed Ubiquitin Variant R4 At Ph 6.3 With 8 M Urea
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
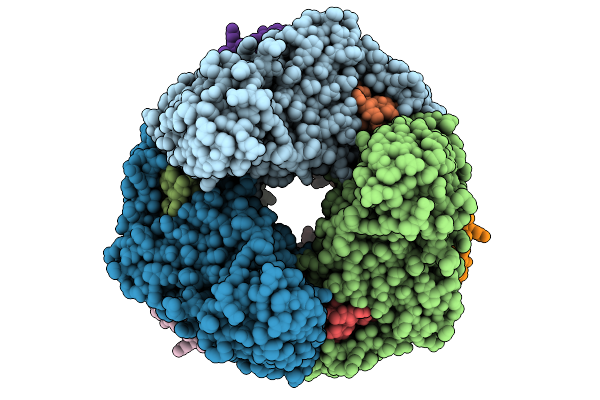 |
Organism: Nitrosospira briensis
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: OXIDOREDUCTASE Ligands: CU, 6PL |
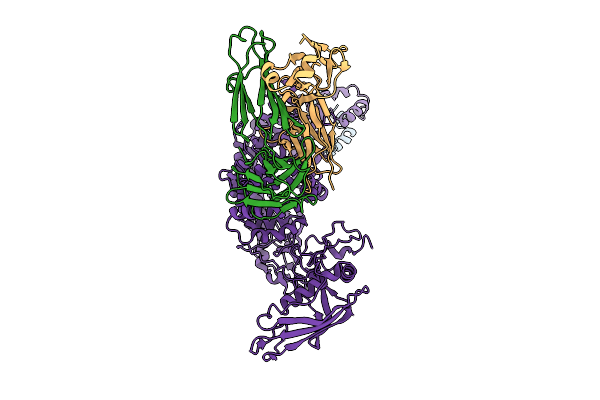 |
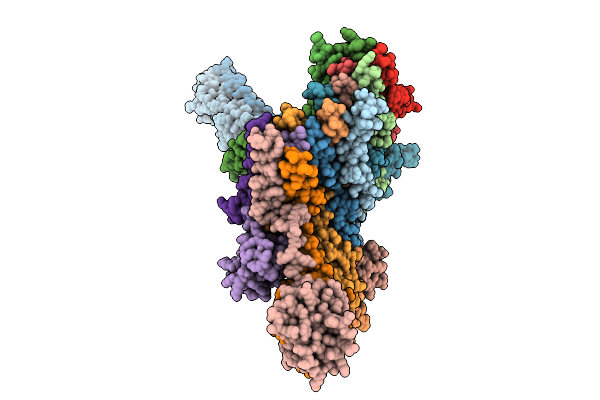
P116 From Mycoplasma Pneumoniae In Complex With Mild Growth Suppressor Monoclonal Antibody
Organism: Mus musculus, Mycoplasmoides pneumoniae m129
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: PROTEIN TRANSPORT |
 |
Cryo-Em Structure Of The Vaccinia Virus Entry/Fusion Complex (Efc) Lacking The F9 Subunit
Organism: Orthopoxvirus vaccinia
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN |
 |
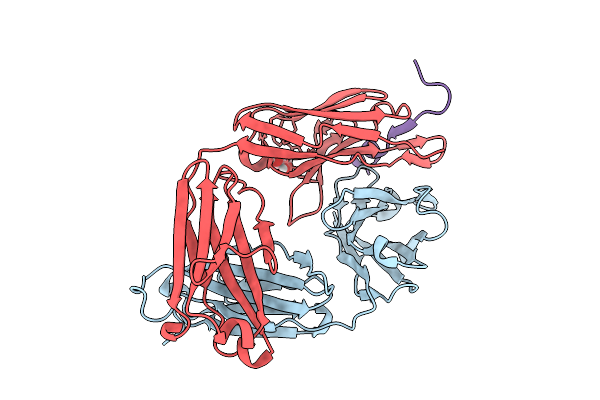
Cryo-Em Structure Of The Vaccinia Virus Entry/Fusion Complex (Efc) Including The F9 Subunit
Organism: Orthopoxvirus vaccinia
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN |
 |
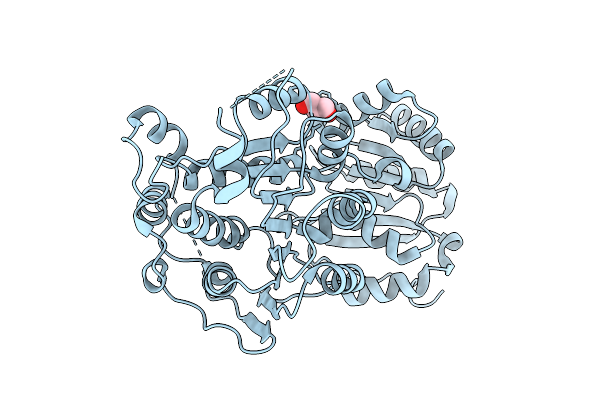
Organism: Oryctolagus cuniculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2025-11-12 Classification: ANTITUMOR PROTEIN Ligands: GOL |
 |
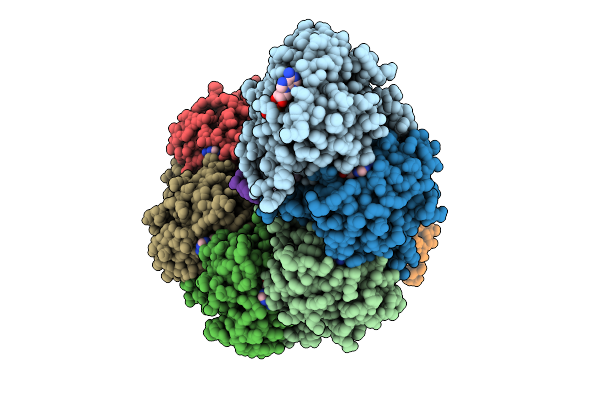
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-10-29 Classification: RNA BINDING PROTEIN Ligands: GOL |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: DNA BINDING PROTEIN/DNA Ligands: ANP, MG |
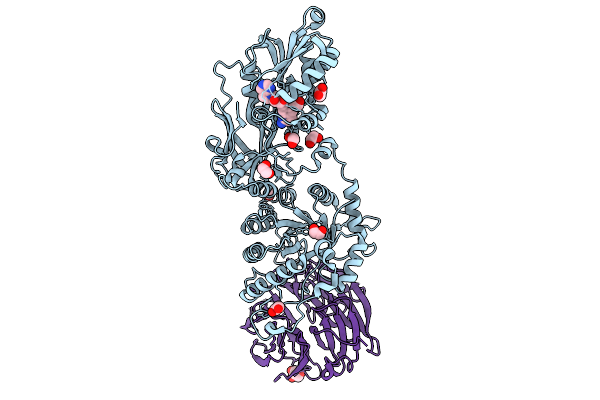 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-09-10 Classification: TRANSLOCASE Ligands: A1CHO, EDO, GOL |
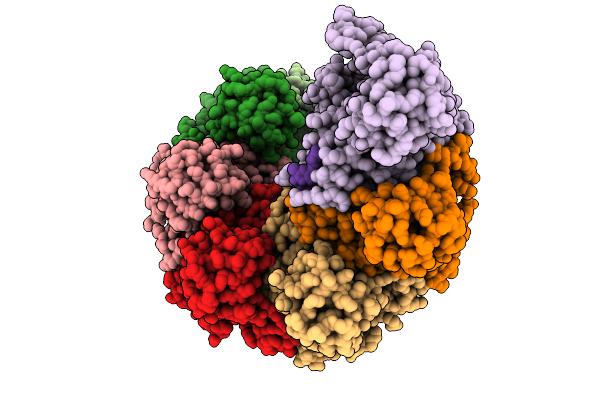 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: DNA BINDING PROTEIN/DNA Ligands: ANP, MG |
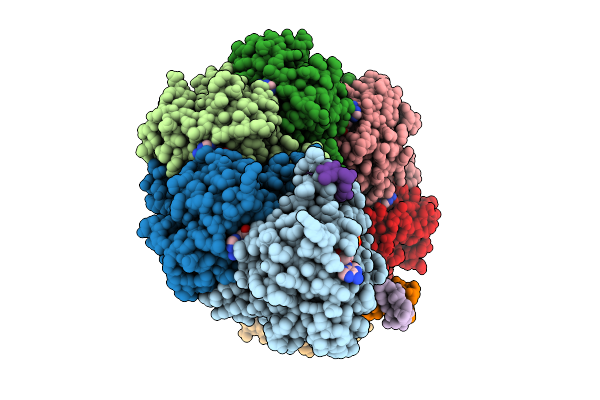 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: DNA BINDING PROTEIN/DNA Ligands: MG, ANP |
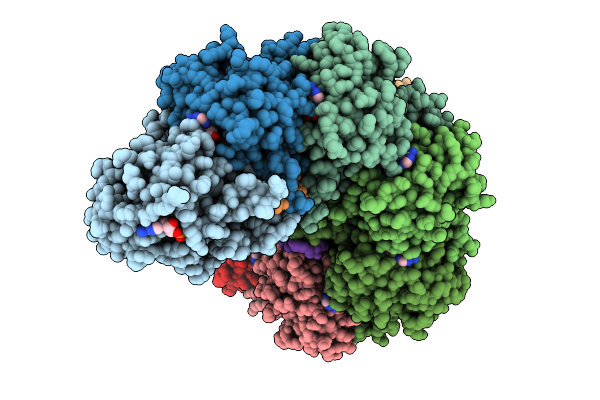 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: DNA BINDING PROTEIN/DNA Ligands: ANP, MG |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: DNA BINDING PROTEIN/DNA Ligands: ANP, MG |

