Search Count: 187
All
Selected
 |
Cryo-Em Structure Of Human Complement C1S Cub Domain In Complex With Ray121
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: IMMUNE SYSTEM Ligands: CA |
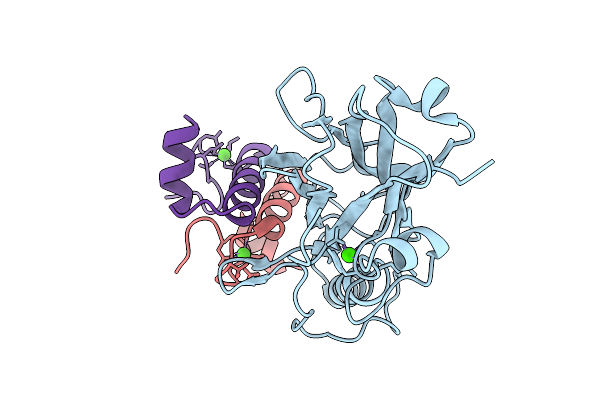 |
Ruminococcus Flavefaciens Coh-Doc Complex Between A Group 2 Dockerin And The Cohesin From Cell Surface Attached Scaffoldin Scae
Organism: Ruminococcus flavefaciens fd-1
Method: X-RAY DIFFRACTION Resolution:2.97 Å Release Date: 2025-10-08 Classification: PROTEIN BINDING Ligands: CA |
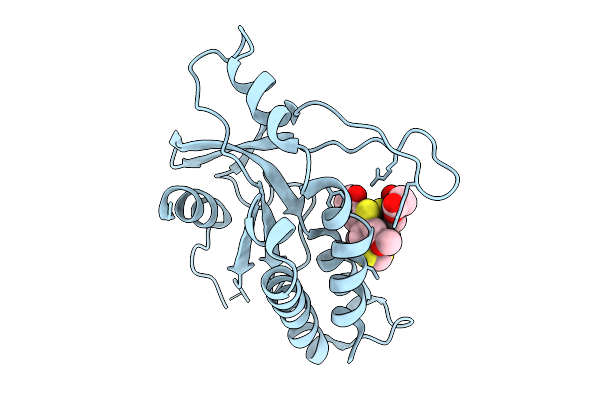 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2025-09-24 Classification: SIGNALING PROTEIN Ligands: A1JA0 |
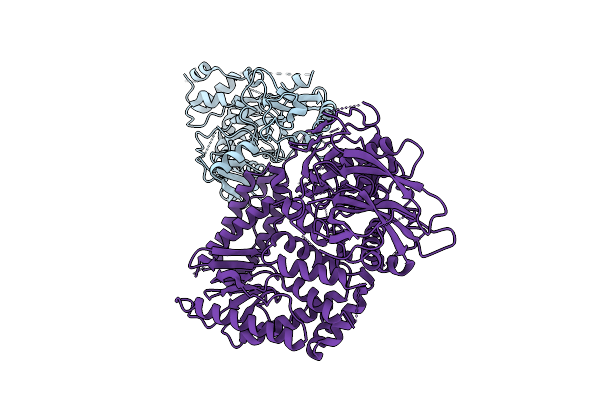 |
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna And Atp-Gamma-S
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
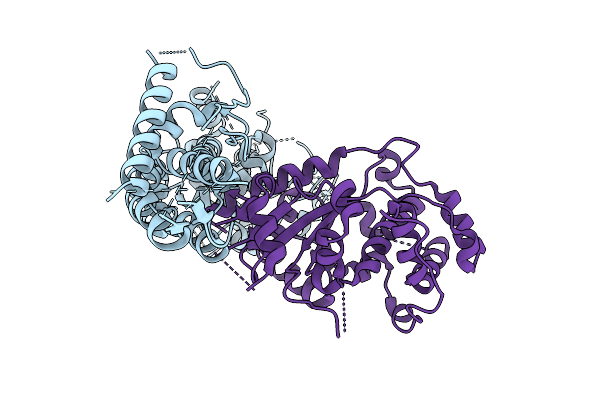 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna And Atp-Gamma-S
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
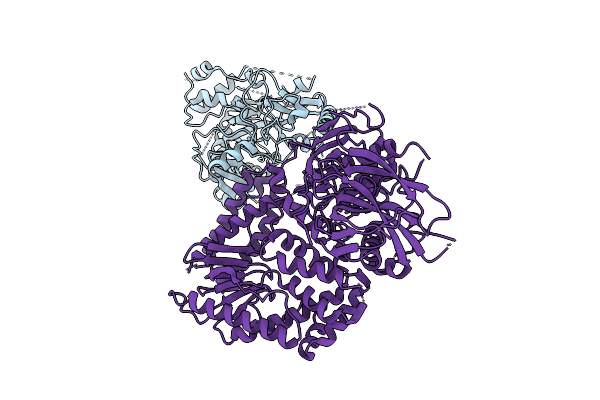 |
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna, Atp-Gamma-S And Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
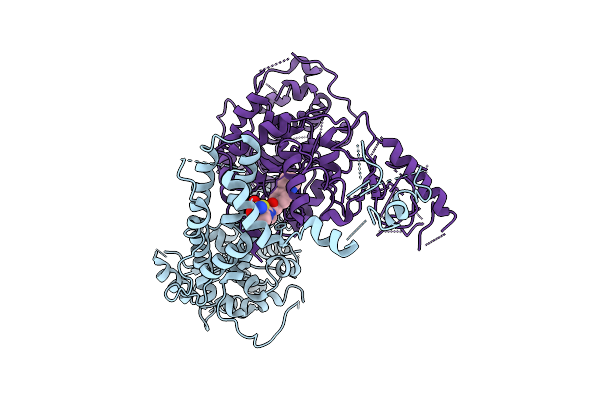 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna, Atp-Gamma-S And Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXB |
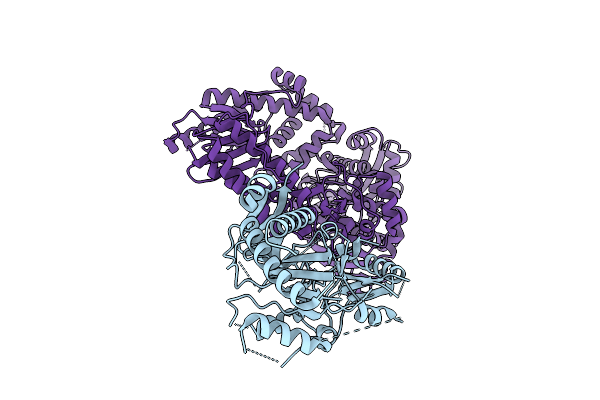 |
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR |
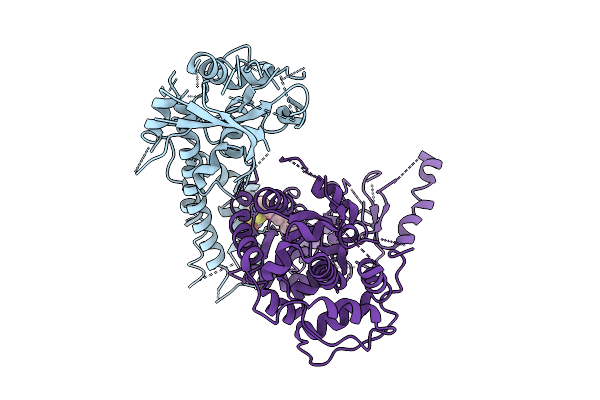 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXB |
 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Amenamevir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXD |
 |
Glycoside Hydrolase Family 157 From Labilibaculum Antarcticum, Wild Type Semet Derivative (Lagh157)
Organism: Labilibaculum antarcticum
Method: X-RAY DIFFRACTION Resolution:2.44 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: ACT, GOL, MLA |
 |
Glycoside Hydrolase Family 157 From Labilibaculum Antarcticum (Lagh157) E224A Mutant In Complex With Laminaritriose And Glucose
Organism: Labilibaculum antarcticum
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2025-05-21 Classification: HYDROLASE Ligands: EDO, BGC, GOL, PGE |
 |
Glycoside Hydrolase Family 157 From Labilibaculum Antarcticum (Lagh157) In Complex With Laminaribiose
Organism: Labilibaculum antarcticum
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2025-05-21 Classification: HYDROLASE Ligands: MLA, GOL, BGC, PG4 |
 |
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-12-11 Classification: OXIDOREDUCTASE Ligands: GOL, CU |
 |
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-12-04 Classification: OXIDOREDUCTASE Ligands: CU |
 |
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2024-12-04 Classification: OXIDOREDUCTASE Ligands: GOL, CU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2024-11-20 Classification: CYTOSOLIC PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-20 Classification: CYTOSOLIC PROTEIN Ligands: ANP |
 |
Complex Of Npr1 Ectodomain With Anp Plus An Allosteric Activating Antibody, Regn5381
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Complex Of Npr1 Ectodomain And Regn5381 Fab In An Active-Like State With No Anp Bound
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: IMMUNE SYSTEM |

