Search Count: 118
All
Selected
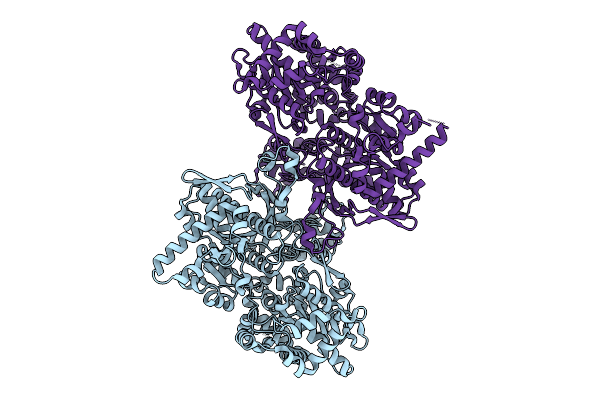 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
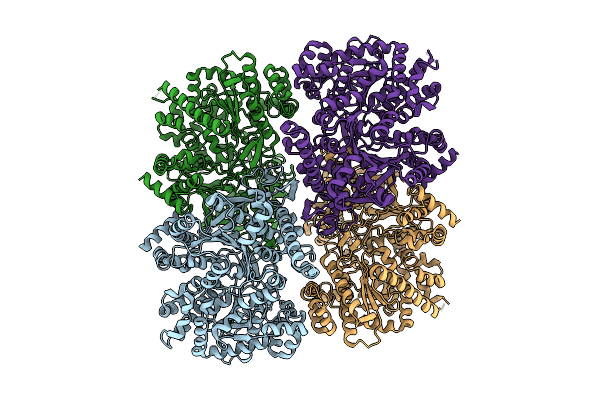 |
Structure Of Glycogen Phosphorylase - Tetrameric Form - From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
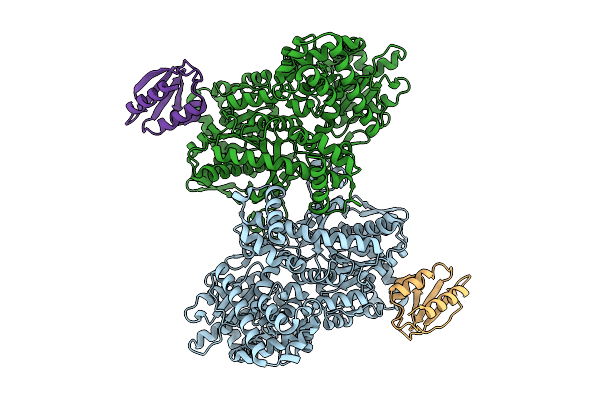 |
Structure Of Glycogen Phosphorylase - Dimeric Form - In Complex With Hpr From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
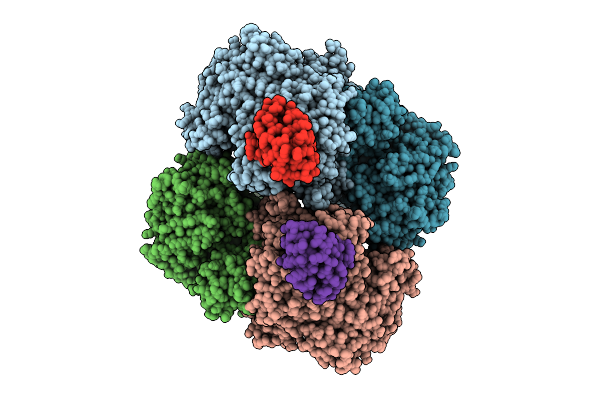 |
Structure Of Glycogen Phosphorylase - Tetrameric Form - In Complex With Hpr From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Organism: Bacteroides thetaiotaomicron vpi-5482
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-07-16 Classification: HYDROLASE |
 |
Crystal Structure Of Gh139 Glycoside Hydrolase From Verrucomicrobium Sp. In The Hexagonal Space Group P6522
Organism: Verrucomicrobium sp.
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: GOL |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2025-02-12 Classification: CYTOSOLIC PROTEIN Ligands: NAD, SER, 2HG |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2025-01-15 Classification: CYTOSOLIC PROTEIN Ligands: NAD, PI |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-01-15 Classification: CYTOSOLIC PROTEIN Ligands: NAD, HPV, GOL |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-01-15 Classification: CYTOSOLIC PROTEIN Ligands: NAD, 3PG, BU1 |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2025-01-15 Classification: CYTOSOLIC PROTEIN Ligands: NAD, SER |
 |
Organism: Rhodopirellula
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2025-01-08 Classification: HYDROLASE Ligands: MPO, SO4, CA |
 |
Crystal Structure Of Pbfuca From Planctomycetes Bacterium K23_9 In P 1 21 1
Organism: Planctomycetes bacterium k23_9
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-01-01 Classification: HYDROLASE Ligands: EDO |
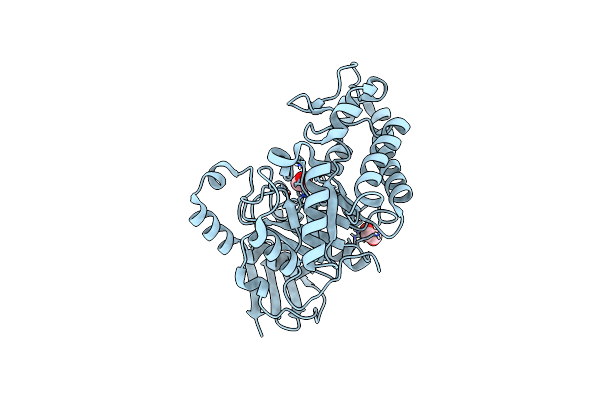 |
Crystal Structure Of Pbfuca From Planctomycetes Bacterium K23_9 In P 21 21 21
Organism: Planctomycetes bacterium k23_9
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-01-01 Classification: HYDROLASE Ligands: EDO |
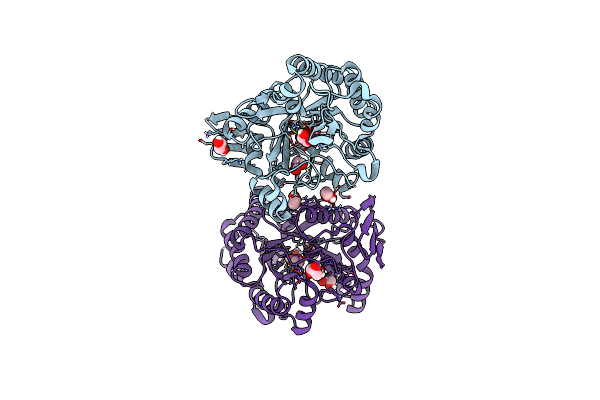 |
Crystal Structure Of Pbfuca From Planctomycetes Bacterium K23_9 In P 4 21 2
Organism: Planctomycetes bacterium k23_9
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2025-01-01 Classification: HYDROLASE Ligands: EDO, GOL |
 |
Crystal Structure Of Igg1-Fc Fragment (E382S) In Complex With Corynebacterial Engase Cu43 (D187A-E189A)
Organism: Corynebacterium ulcerans, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.62 Å Release Date: 2024-10-30 Classification: IMMUNE SYSTEM |
 |
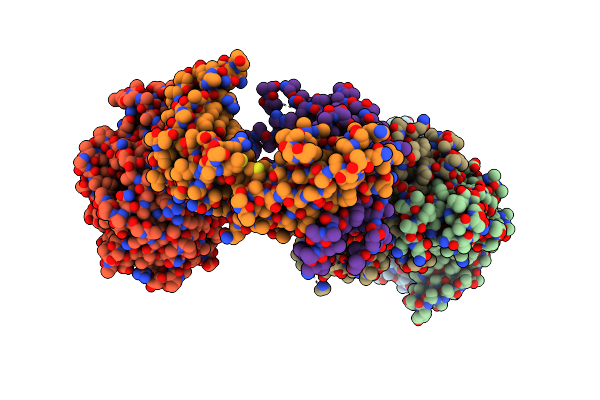
Capsular Polysaccharide Synthesis Multienzyme Of Actinobacillus Pleuropneumoniae Serotype 3
Organism: Actinobacillus pleuropneumoniae
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2024-07-03 Classification: BIOSYNTHETIC PROTEIN Ligands: SO4, ZN |
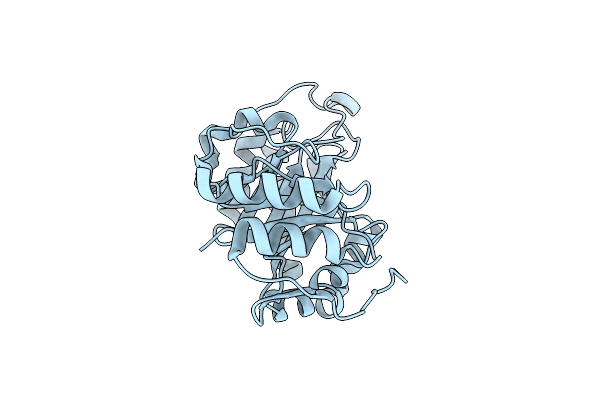 |
Crystal Structure Of Bacteroides Thetaiotaomicron Vpi-5482 Endoglycosidase Bt1285 D161A-E163A Inactive Version
Organism: Bacteroides thetaiotaomicron vpi-5482
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-05-29 Classification: HYDROLASE |
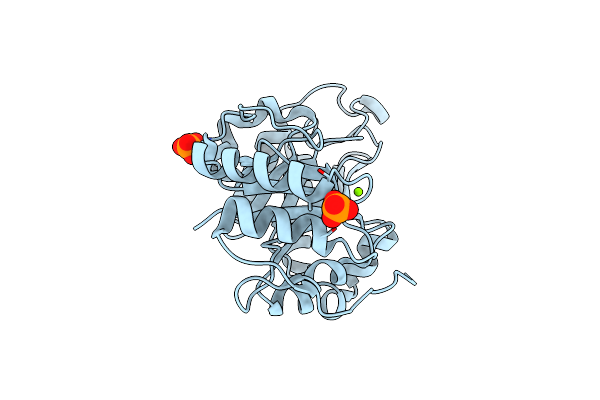 |
Crystal Structure Of Bacteroides Thetaiotaomicron Vpi-5482 Endoglycosidase Bt1285
Organism: Bacteroides thetaiotaomicron vpi-5482
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2024-05-29 Classification: HYDROLASE Ligands: PO4, MG |
 |
Crystal Structure Of Bacteroides Thetaiotamicron Bt1285 D161A-E163A Inactive Endoglycosidase In Complex With High-Mannose N-Glycan (Man9Glcnac2) Substrate
Organism: Bacteroides thetaiotaomicron vpi-5482
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-05-29 Classification: CARBOHYDRATE Ligands: PO4 |

