Search Count: 26
All
Selected
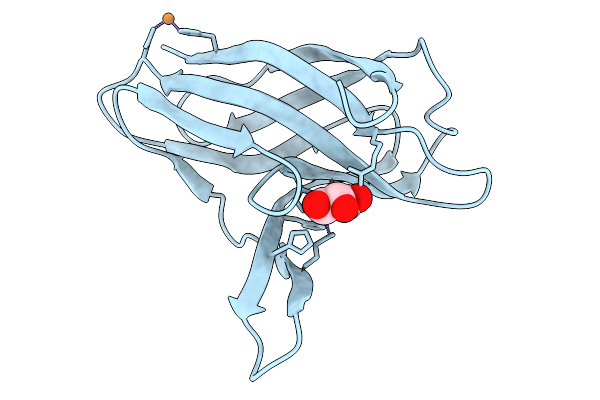 |
Organism: Neisseria meningitidis serogroup a
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-03-25 Classification: METAL BINDING PROTEIN Ligands: CU, GOL |
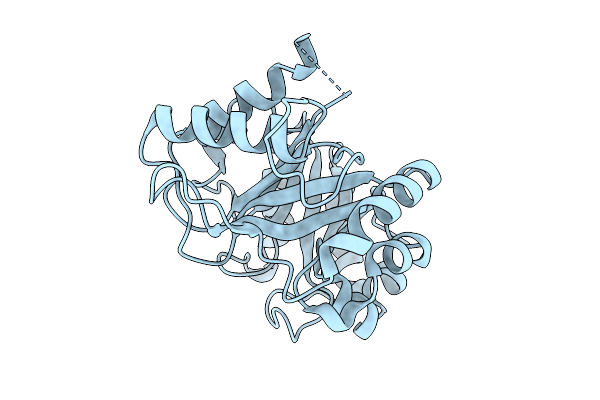 |
Organism: Oryza sativa subsp. japonica
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-09-03 Classification: PLANT PROTEIN |
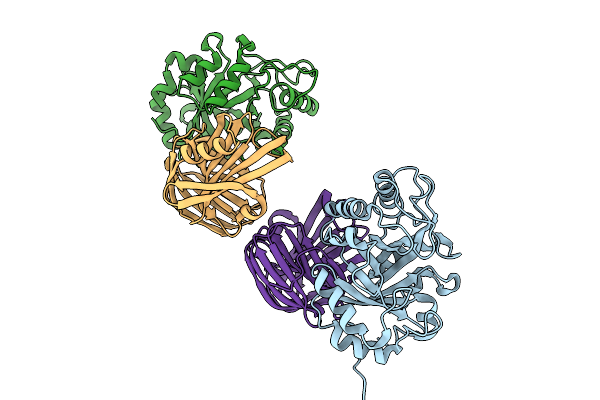 |
Organism: Rhizopus arrhizus, Oryza sativa subsp. japonica
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-09-03 Classification: PROTEIN BINDING |
 |
Crystal Structure Of The Type I-B Crispr-Associated Protein, Csh2 From Thermobaculum Terrenum
Organism: Thermobaculum terrenum
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2023-05-31 Classification: TRANSFERASE |
 |
Organism: Streptococcus mutans ua159
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2023-01-11 Classification: TRANSCRIPTION Ligands: CIT |
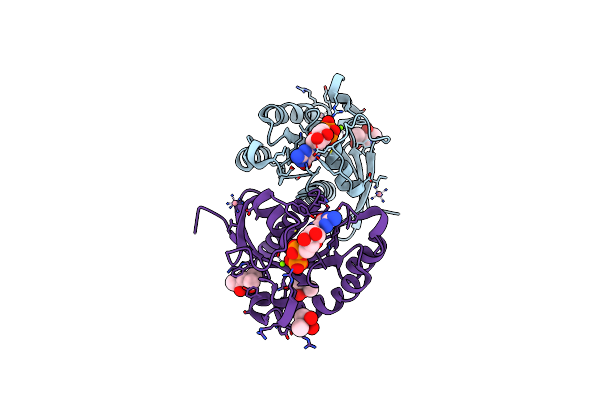 |
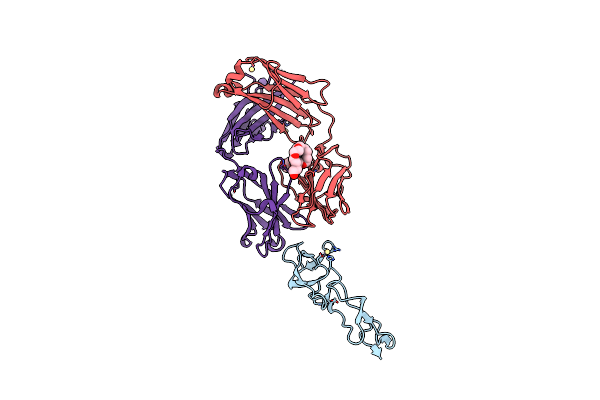
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2022-11-02 Classification: CELL CYCLE Ligands: GDP, MG, NCO, MPD, CAC |
 |
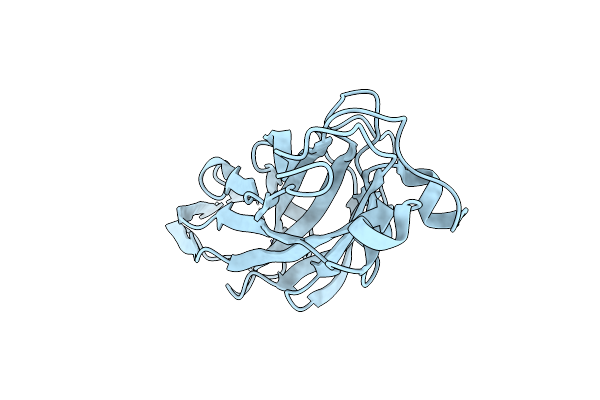
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2022-09-28 Classification: IMMUNE SYSTEM Ligands: CD, 2PE |
 |
Organism: Proteus mirabilis
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2022-07-06 Classification: SUGAR BINDING PROTEIN Ligands: CL |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2022-07-06 Classification: SUGAR BINDING PROTEIN Ligands: IOD |
 |
Organism: Proteus mirabilis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2022-07-06 Classification: SUGAR BINDING PROTEIN Ligands: FUL, CL |
 |
Crystal Structure Of The Ucad Lectin-Binding Domain In Complex With Glucose
Organism: Proteus mirabilis
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2022-07-06 Classification: SUGAR BINDING PROTEIN Ligands: BGC, CL |
 |
Crystal Structure Of The Ucad Lectin-Binding Domain In Complex With Galactose
Organism: Proteus mirabilis
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2022-07-06 Classification: SUGAR BINDING PROTEIN Ligands: GLA, CL |
 |
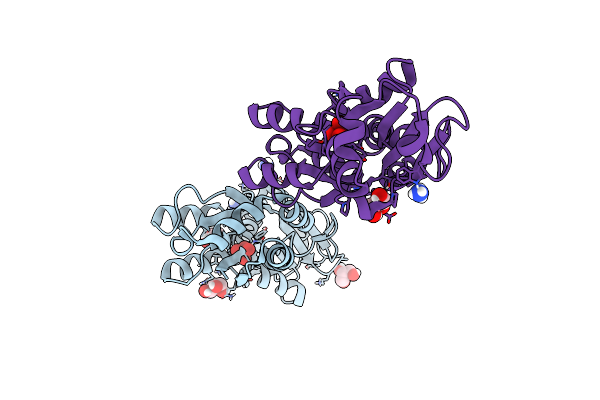
Crystal Structure Of The Molybdate-Binding Periplasmic Protein Moda From The Bacteria Pseudomonsa Aeruginosa In Ligand-Free Form
Organism: Pseudomonas aeruginosa pa1
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2022-05-04 Classification: METAL BINDING PROTEIN Ligands: GOL, NH4, SO4 |
 |
Crystal Structure Of The Molybdate-Binding Periplasmic Protein Moda From The Bacteria Pseudomonsa Aeruginosa In Chromate-Bound Form
Organism: Pseudomonas aeruginosa pa1
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-05-04 Classification: METAL BINDING PROTEIN Ligands: CQ4, NH4, GOL |
 |
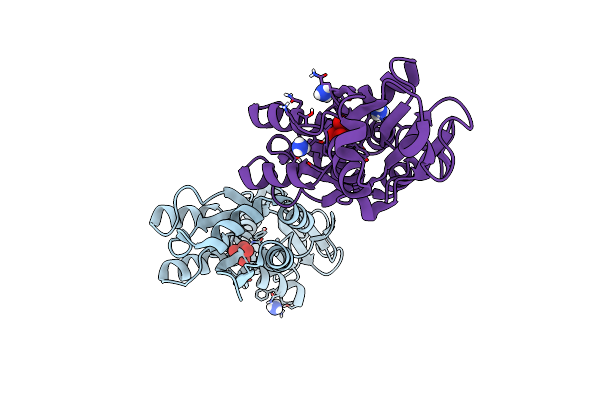
Crystal Structure Of The Molybdate-Binding Periplasmic Protein Moda From The Bacteria Pseudomonsa Aeruginosa In Molybdate-Bound Form
Organism: Pseudomonas aeruginosa pa1
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2022-05-04 Classification: METAL BINDING PROTEIN Ligands: NH4, MOO, GOL |
 |
Crystal Structure Of The Molybdate-Binding Periplasmic Protein Moda From The Bacteria Pseudomonsa Aeruginosa In Tungstate-Bound Form
Organism: Pseudomonas aeruginosa pa1
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2022-05-04 Classification: METAL BINDING PROTEIN Ligands: WO4, NH4 |
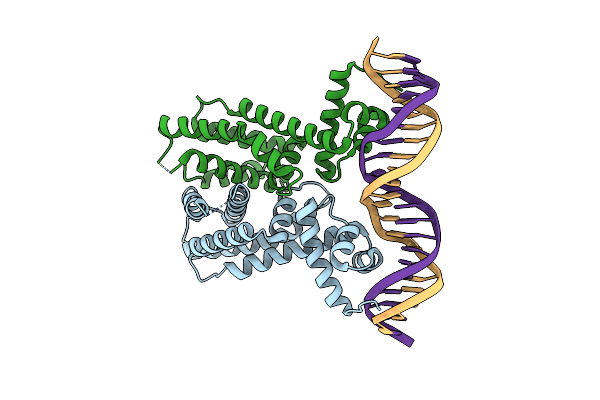 |
Organism: Vibrio alginolyticus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2021-03-31 Classification: TRANSCRIPTION |
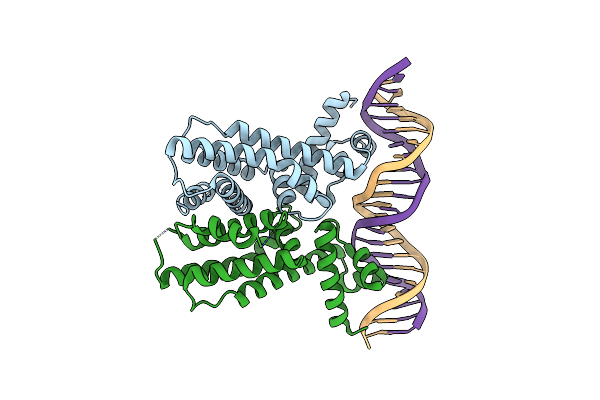 |
Organism: Vibrio alginolyticus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2021-03-31 Classification: TRANSCRIPTION |
 |
A Pre-Assembled Molecular-Helical Cascade Backbone Of Csy3 Subunits From Zymomonas Mobilis
Organism: Zymomonas mobilis subsp. mobilis
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2020-05-13 Classification: RNA BINDING PROTEIN |
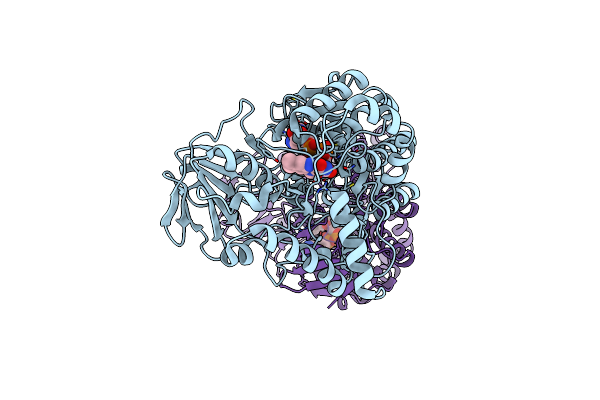 |
Crystal Structure Of Baeyer-Villiger Monooxygenase From Parvibaculum Lavamentivorans
Organism: Parvibaculum lavamentivorans (strain ds-1 / dsm 13023 / ncimb 13966)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2019-04-10 Classification: OXIDOREDUCTASE Ligands: FAD, MG |

