Search Count: 5,020
All
Selected
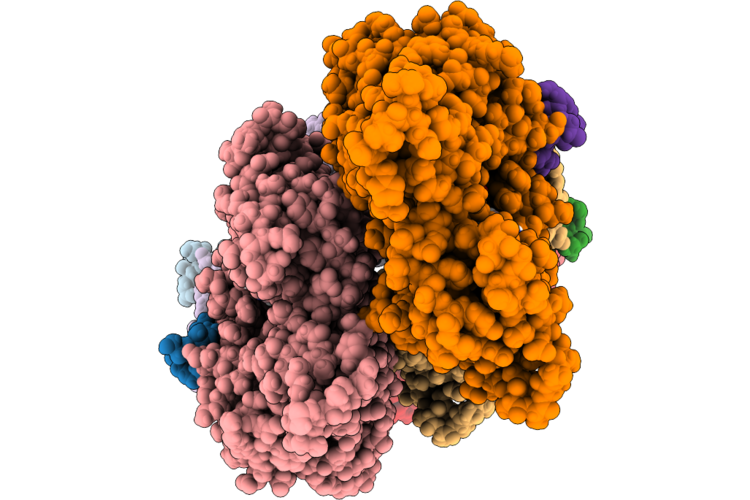 |
Organism: Homo sapiens, Gallus gallus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: STRUCTURAL PROTEIN Ligands: MG, ADP |
 |
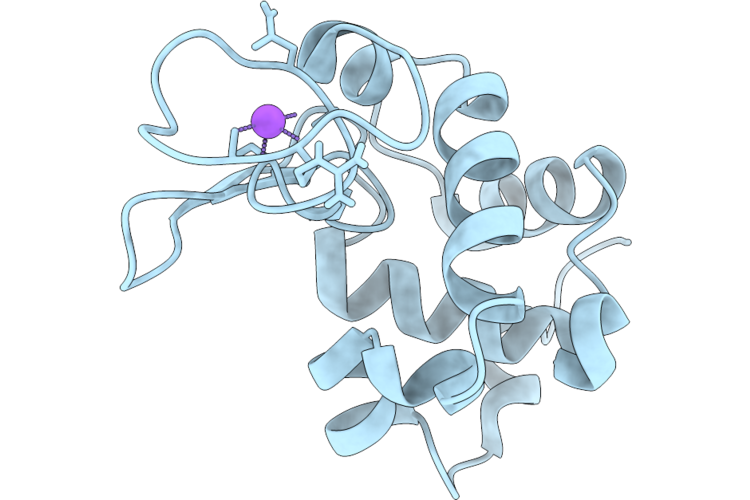
Organism: Gallus gallus
Method: ELECTRON CRYSTALLOGRAPHY Resolution:0.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ACT, CL |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CL, NA |
 |
Organism: Gallus gallus
Method: ELECTRON MICROSCOPY Resolution:3.06 Å Release Date: 2026-04-15 Classification: IMMUNE SYSTEM Ligands: NAG, BMA, MAN |
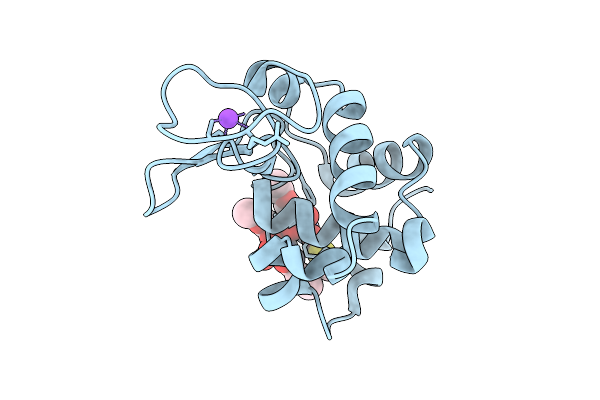 |
X-Ray Structure Of Lysozyme Obtained Upon Reaction With The Dioxidovanadium(V) Complex With Thiophene-2-Carboxylic Acid (3-Ethoxy-2-Hydroxybenzylidene)Hydrazide
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-04-01 Classification: HYDROLASE |
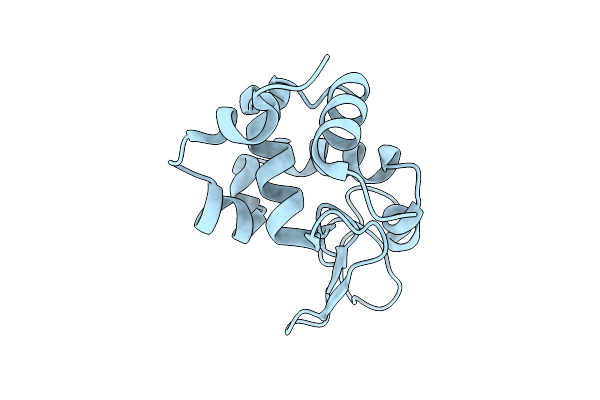 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: CL |
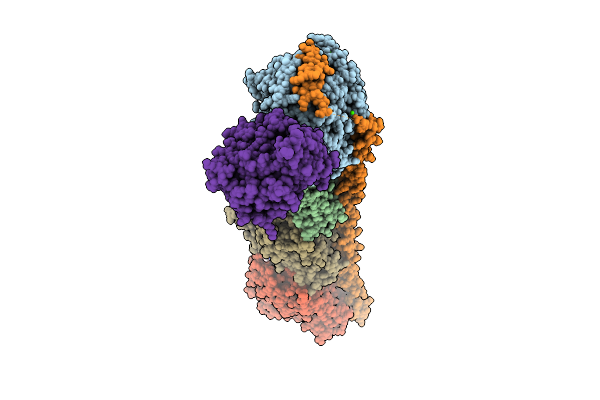 |
Organism: Rattus norvegicus, Gallus gallus, Sus scrofa
Method: X-RAY DIFFRACTION Resolution:2.93 Å Release Date: 2026-04-01 Classification: CELL CYCLE Ligands: GTP, MG, CA, MES, A1L80, GDP, ACP |
 |
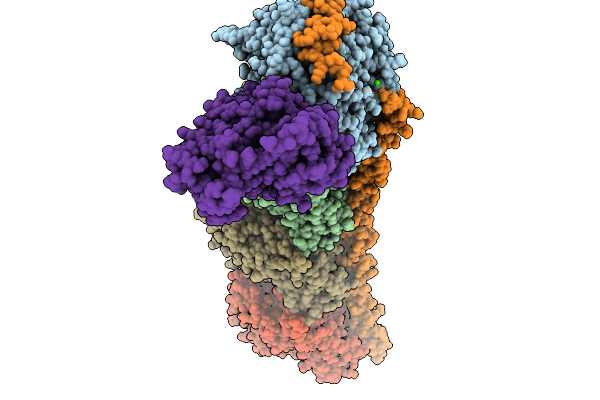
Organism: Rattus norvegicus, Gallus gallus, Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-03-25 Classification: CELL CYCLE Ligands: GTP, MG, GOL, CA, CL, GDP, MES, A1I6L, IMD, ACP |
 |
Organism: Rattus norvegicus, Gallus gallus, Sus scrofa
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2026-03-25 Classification: CELL CYCLE Ligands: GTP, MG, CA, MES, GDP, A1L8Z |
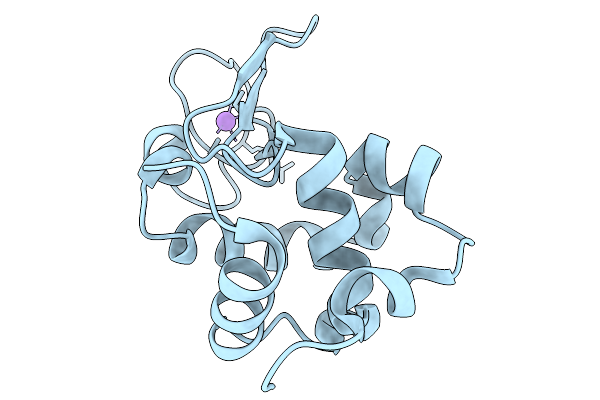 |
In Situ Structure Of Egg-White Lysozyme Using A Goniometer-Compatible Chip-Based Platform
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-03-25 Classification: HYDROLASE |
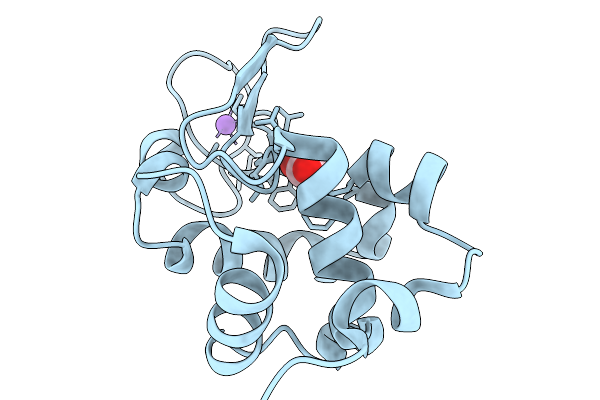 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-25 Classification: HYDROLASE |
 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 2 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 5 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 9.7 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 1.3 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 5.0 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 7.5 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 9.7 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 452 Mm N-Acetyl-D-Glucosamine At 1.3 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |

