Search Count: 61
All
Selected
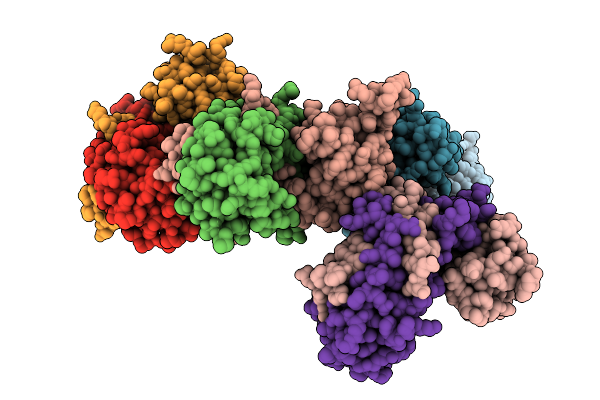 |
Structural Insights Into Hpar-Mediated Recognition Of Hrpx And Hrpg In Xanthomonas Campestris Pv. Campestris
Organism: Xanthomonas campestris pv. campestris (strain atcc 33913 / dsm 3586 / ncppb 528 / lmg 568 / p 25)
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-03-11 Classification: TRANSCRIPTION |
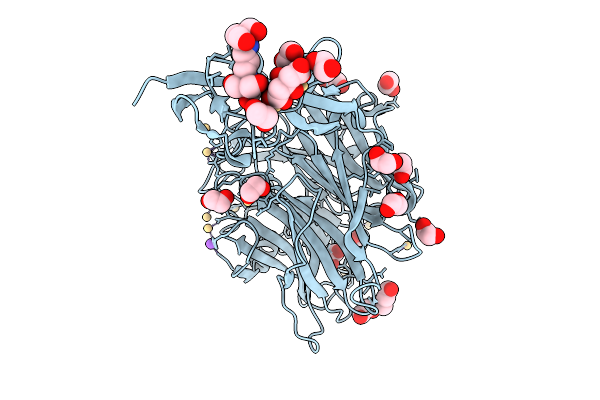 |
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: CD, NA, B3P, CIT, EDO, PGE |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) Bound To Neu5Ac2En (Dana)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: DAN, EDO, PEG, CD |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) Bound To Neu5Ac (Nana)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: CD, SLB, EDO, PEG, PGE |
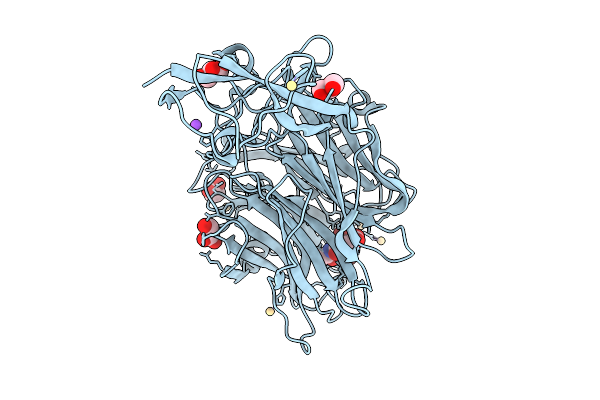 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) D165A Mutant Bound To 3'-Sialyllactose (Only Neu5Ac Visible)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: SIA, GOL, PEG, CD, NA |
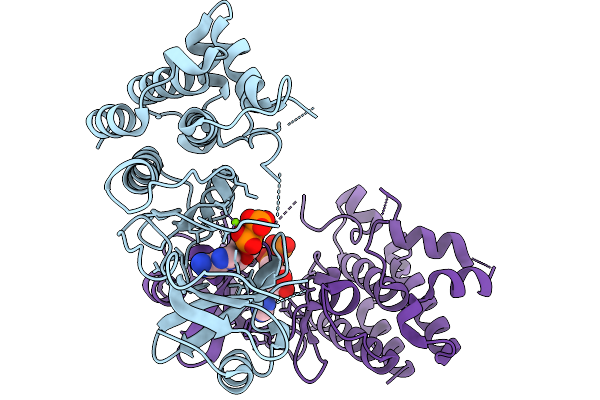 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: ADP, MG |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.52 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: ADP, MG |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.76 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: MG, ADP |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: ADP, MG |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: ANP |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: MG, ANP |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-01-28 Classification: TRANSFERASE |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: ATP, MG |
 |
Structural Basis Of Signal Activation And Transduction By Chitin Elicitor Receptor Kinase 1 In Oryza Sativa
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: ADP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2025-03-05 Classification: TRANSCRIPTION Ligands: PLM, EUI |
 |
Structural Basis Of Chorismate Isomerization By Arabidopsis Isochorismate Synthase Ics1
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-05-22 Classification: ISOMERASE Ligands: ACT, FMT, MG |
 |
Structural Basis Of Chorismate Isomerization By Arabidopsis Isochorismate Synthase Ics1
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2024-05-22 Classification: ISOMERASE Ligands: ISJ, MG, FMT |
 |
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2023-10-04 Classification: HYDROLASE Ligands: FLC, PG4, PEG, PGE |
 |
Crystal Structure Of Porphyromonas Gingivalis Sialidase (Pg_0352) Bound To Neu5Ac2En (Dana)
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2023-10-04 Classification: HYDROLASE Ligands: DAN, PGE, PEG, PG4 |
 |
Crystal Structure Of Porphyromonas Gingivalis Sialidase (Pg_0352) Bound To Neu5Ac (Nana)
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2023-10-04 Classification: HYDROLASE Ligands: SIA, PGE, PEG, PG4 |

