Search Count: 763
All
Selected
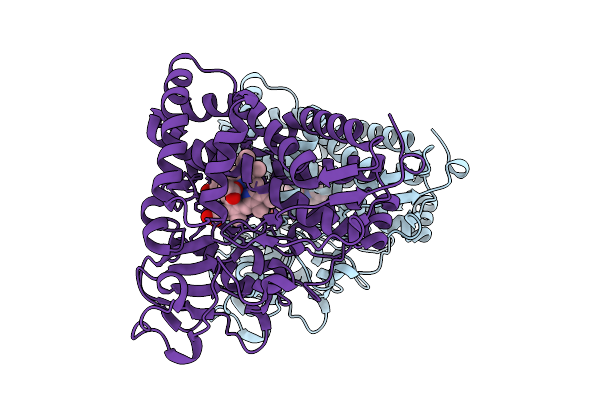 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: HEM, ACT |
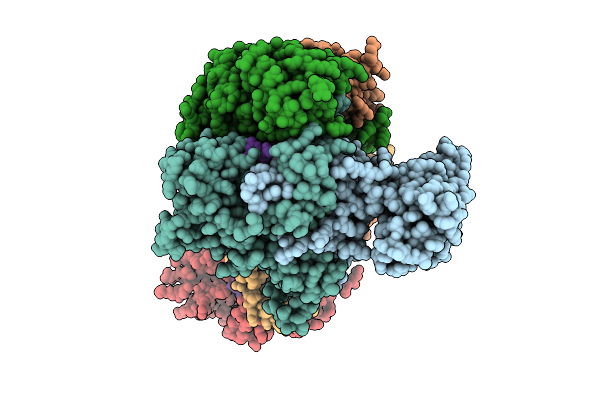 |
Organism: Pseudomonas aeruginosa ucbpp-pa14
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Pseudomonas aeruginosa ucbpp-pa14
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN/RNA |
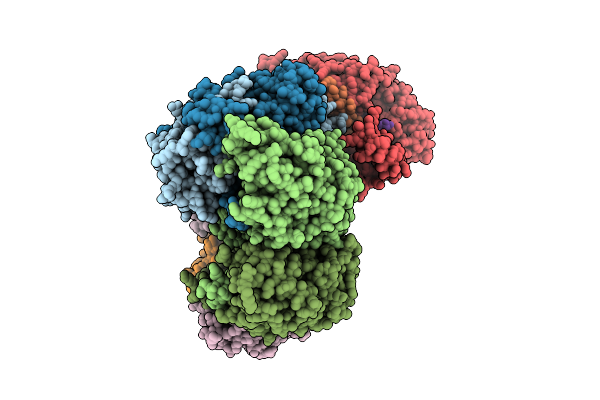 |
Organism: Pseudomonas aeruginosa ucbpp-pa14
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN/RNA |
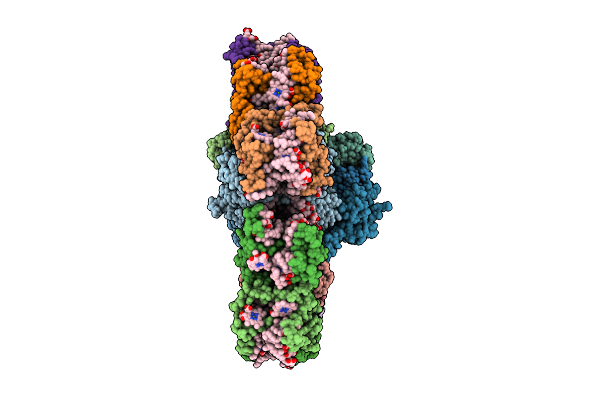 |
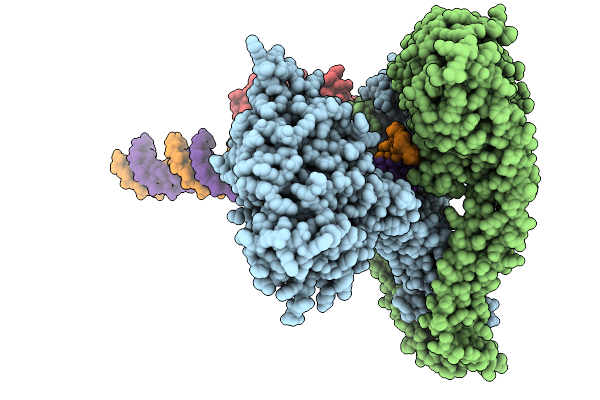
Organism: Euglena gracilis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: BCR, CLA, PQN, DD6, DGD, LHG, LMU, LMG, SF4, SQD, CHL, NEX, CL0 |
 |
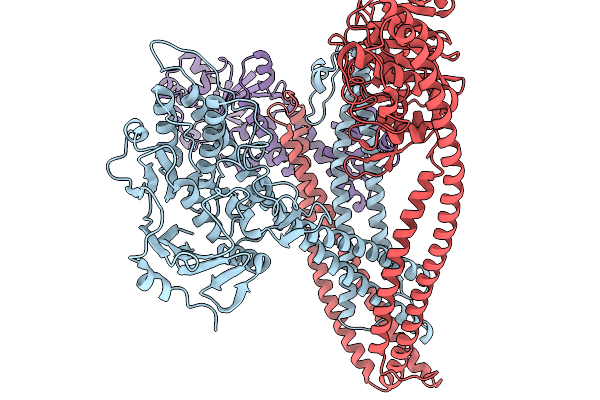
Organism: Bacillus sp. nio-1130, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.73 Å Release Date: 2026-03-25 Classification: IMMUNE SYSTEM |
 |
Organism: Bacillus sp. nio-1130
Method: ELECTRON MICROSCOPY Resolution:3.71 Å Release Date: 2026-03-25 Classification: IMMUNE SYSTEM |
 |
Organism: Vibrio cholerae, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
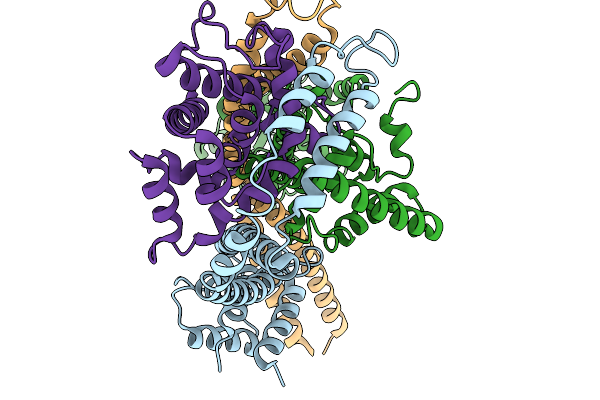
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:3.64 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
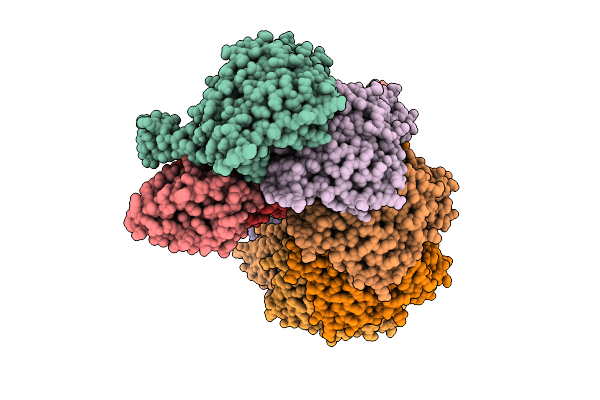
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:3.54 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:4.07 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
Organism: Bifidobacterium pseudocatenulatum, Nitratidesulfovibrio vulgaris str. hildenborough
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: RNA BINDING PROTEIN/RNA |
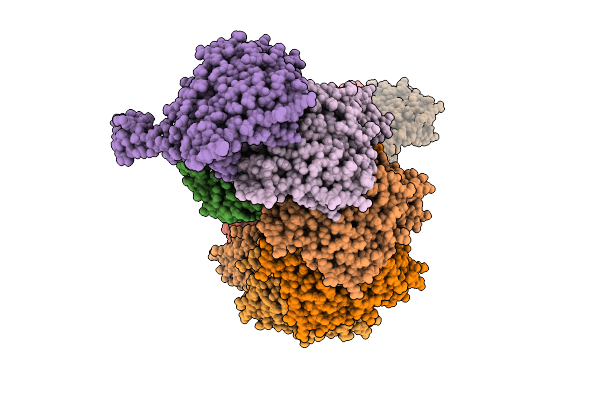 |
Organism: Nitratidesulfovibrio vulgaris str. hildenborough, Bifidobacterium pseudocatenulatum
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: RNA BINDING PROTEIN/RNA |
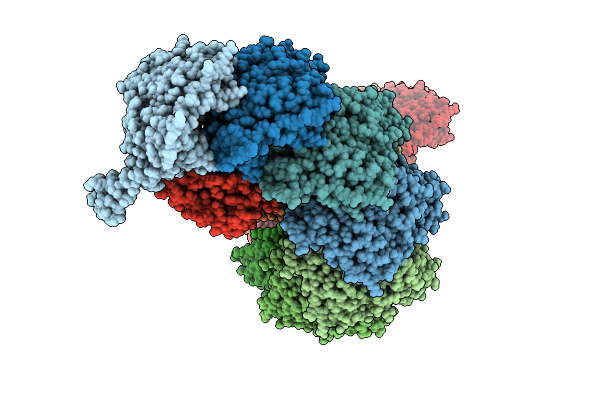 |
Organism: Nitratidesulfovibrio vulgaris str. hildenborough, Bifidobacterium pseudocatenulatum
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: RNA BINDING PROTEIN/RNA |
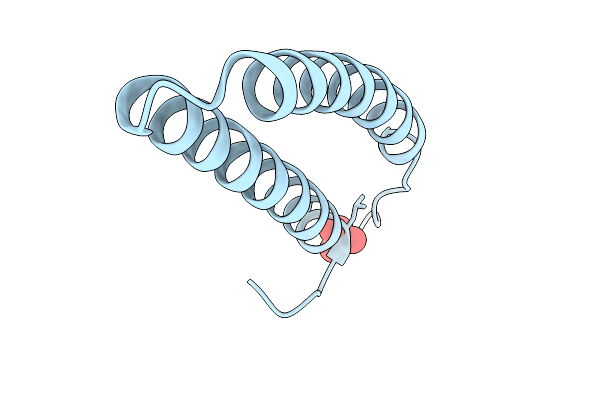 |
Organism: Escherichia phage t4
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2026-03-04 Classification: ANTIMICROBIAL PROTEIN Ligands: SO4 |
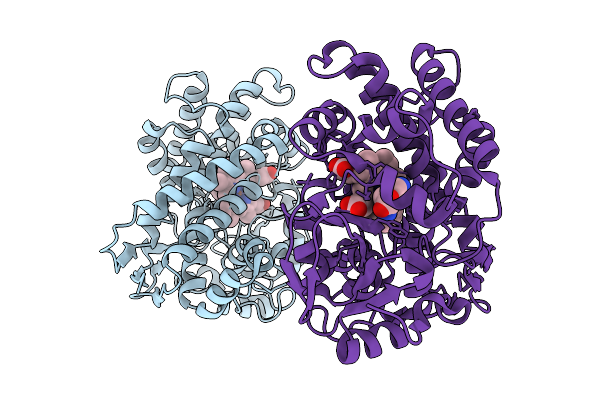 |
Organism: Streptomyces scabiei 87.22
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2026-01-28 Classification: BIOSYNTHETIC PROTEIN Ligands: TRP, HEM |
 |
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, A1EI5 |
 |
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, EDO |
 |
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: A1EI5, HEM |
 |
Crystal Structure Of P450 Bm3 F87A/V78S/L75N Mutant Complex With (-)-Ambroxide
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, A1EI5 |

