Search Count: 86
All
Selected
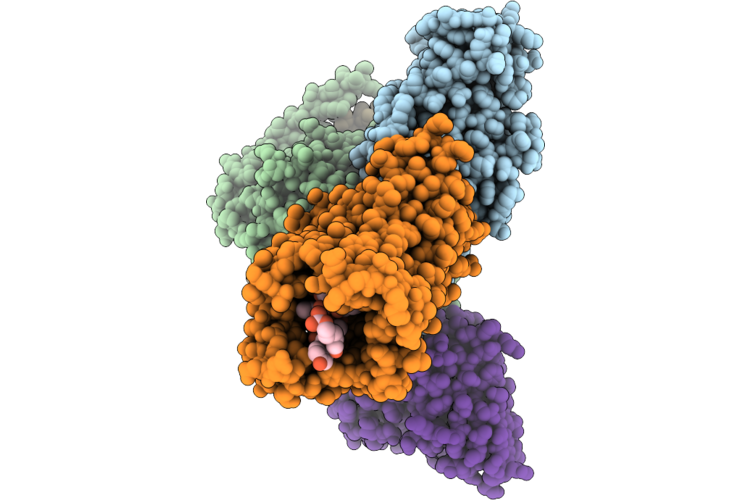 |
Organism: Homo sapiens, Escherichia coli, Oplophorus gracilirostris
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-29 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: CLR |
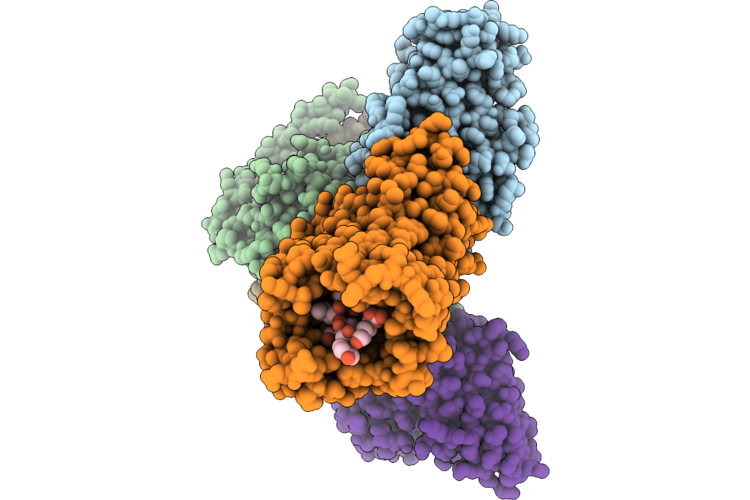 |
Organism: Homo sapiens, Escherichia coli, Oplophorus gracilirostris
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-04-29 Classification: SIGNALING PROTEIN Ligands: CLR |
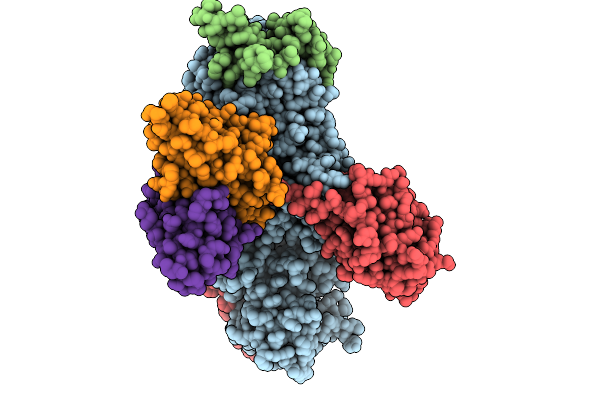 |
Organism: Human gammaherpesvirus 4, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
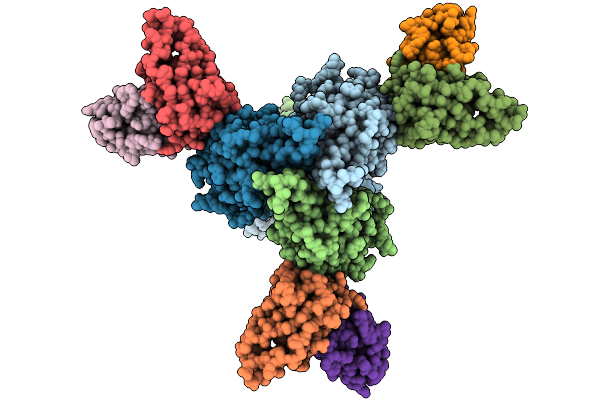 |
Organism: Mus musculus, Murid herpesvirus 68
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Murid gammaherpesvirus 4
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN |
 |
Organism: Macacine gammaherpesvirus 4, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN Ligands: ZN, EDO |
 |
Crystal Structure Of Nfia In Complex With Dna Containing The Tggca(N3)Tgcca Motif
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN Ligands: ZN, EDO |
 |
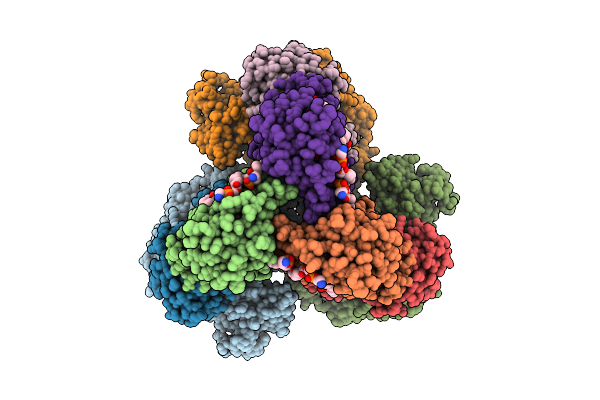
Cryo-Em Structure Of The Apo-Form Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
 |
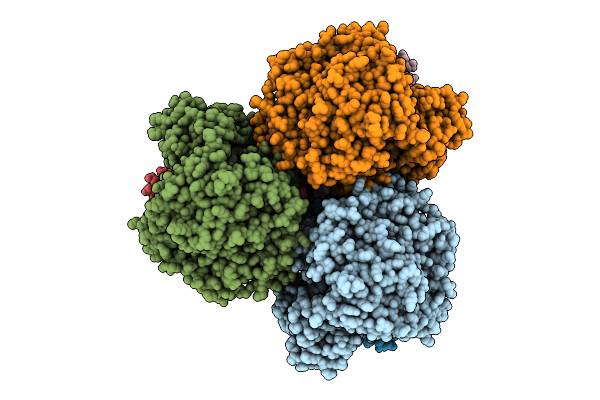
Cryo-Em Structure Of The Lipid-Bound Succiante Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
 |
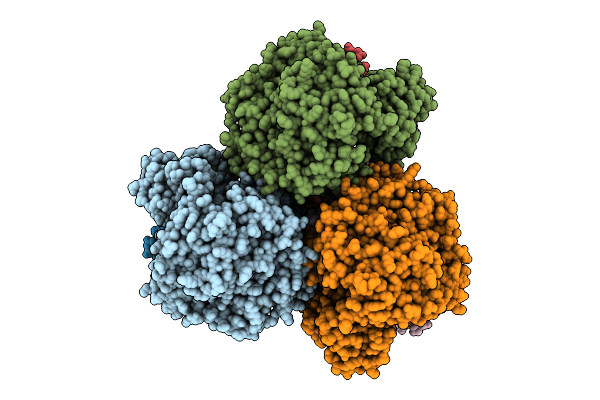
Cryo-Em Structure Of The Mk7-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, MQ7 |
 |
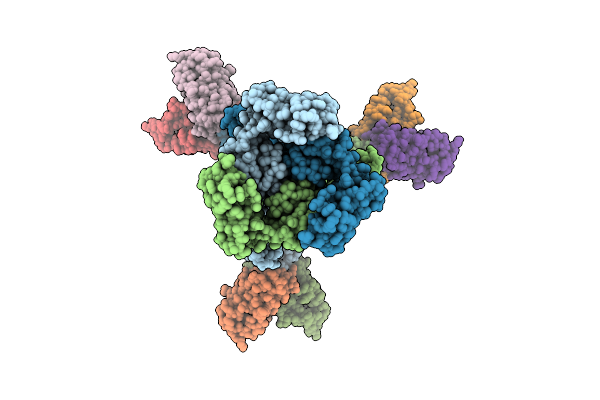
Cryo-Em Structure Of The Mk4-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, 1L3 |
 |
Structure Of A Murine Monoclonal Antibody Fab5 Targeting Epstein-Barr Virus Gb
Organism: Homo sapiens, Human gammaherpesvirus 4
Method: ELECTRON MICROSCOPY Release Date: 2025-10-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
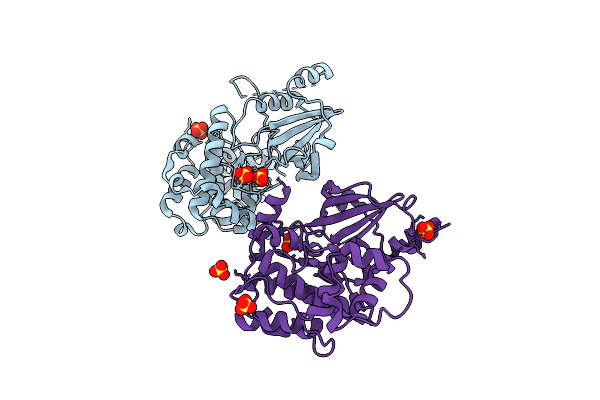 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2025-08-13 Classification: TRANSFERASE Ligands: SO4 |
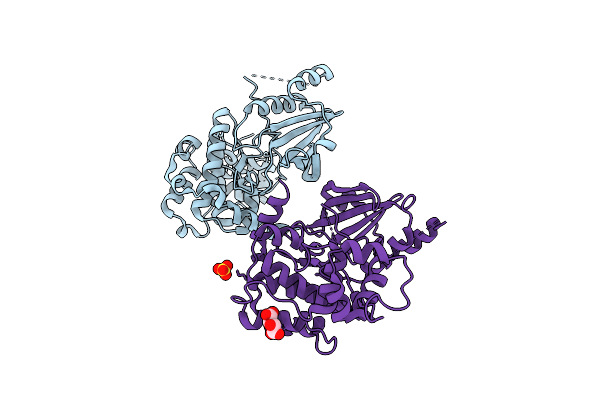 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2025-08-13 Classification: TRANSFERASE Ligands: MLT, SO4 |
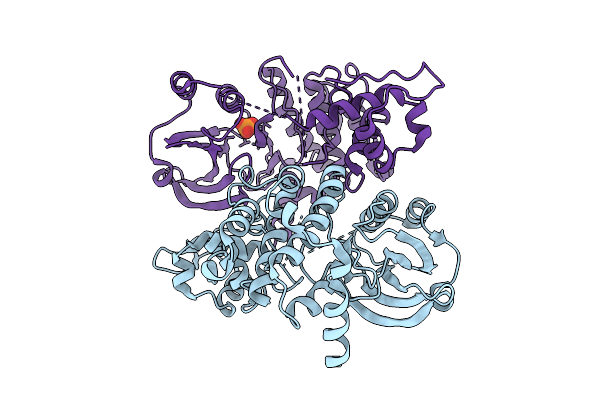 |
The Crystal Structure Of Full-Length Pak2 Containing K278R And D368N Mutants
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.02 Å Release Date: 2025-08-13 Classification: TRANSFERASE Ligands: PO4 |
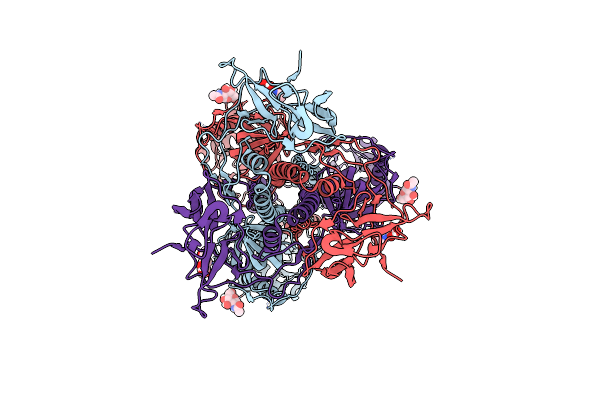 |
Organism: Human herpesvirus 6 strain z29
Method: ELECTRON MICROSCOPY Release Date: 2025-06-25 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: MEMBRANE PROTEIN Ligands: POV, EIC, ACD, D21, K |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: MEMBRANE PROTEIN Ligands: POV, EIC, K |

