Search Count: 1,974
 |
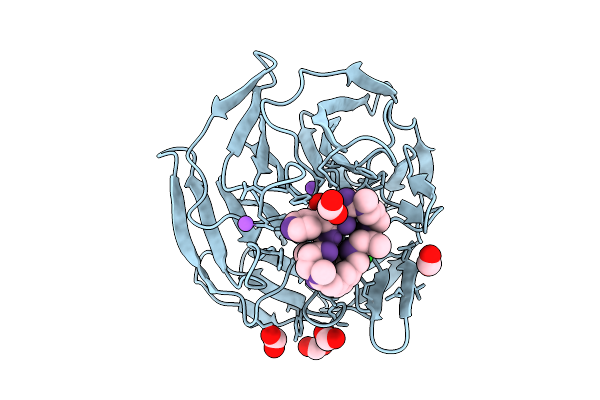
Human Kras G12D (Gdp-Bound) In Complex With Macrocyclic Peptide Inhibitor Ap6252
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-05-27 Classification: SIGNALING PROTEIN Ligands: GDP, EDO, FMT, MG |
 |
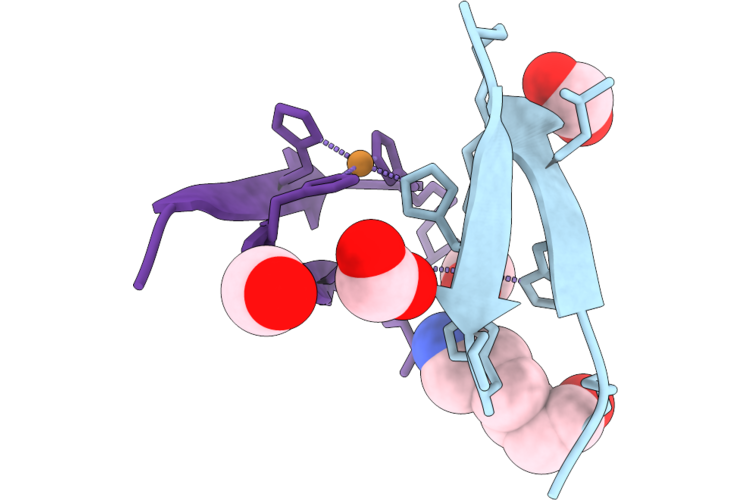
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-05-06 Classification: MOTOR PROTEIN Ligands: FMT, GOL |
 |
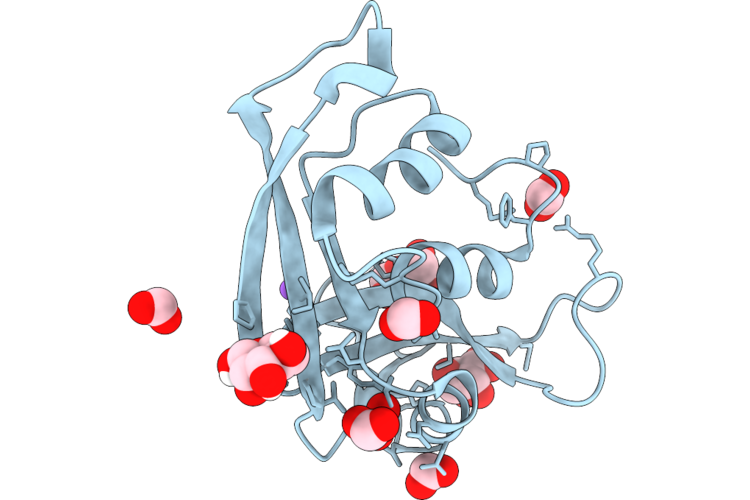
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-04-22 Classification: DE NOVO PROTEIN Ligands: CU, FMT, ACA |
 |
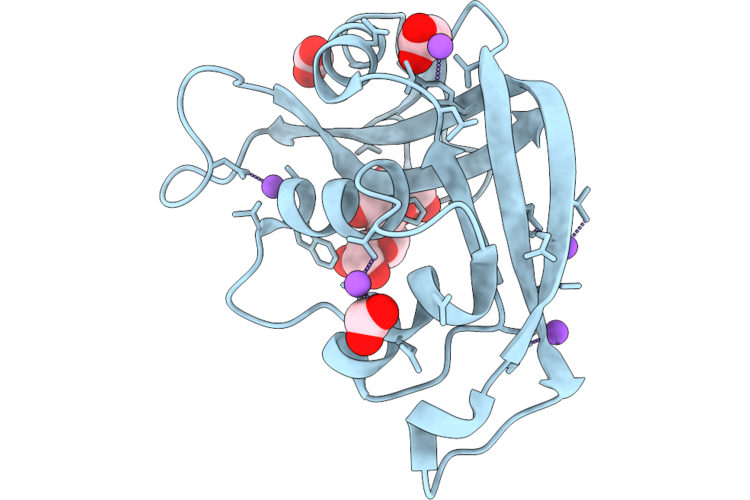
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: MAN, FMT, BMA, NA |
 |
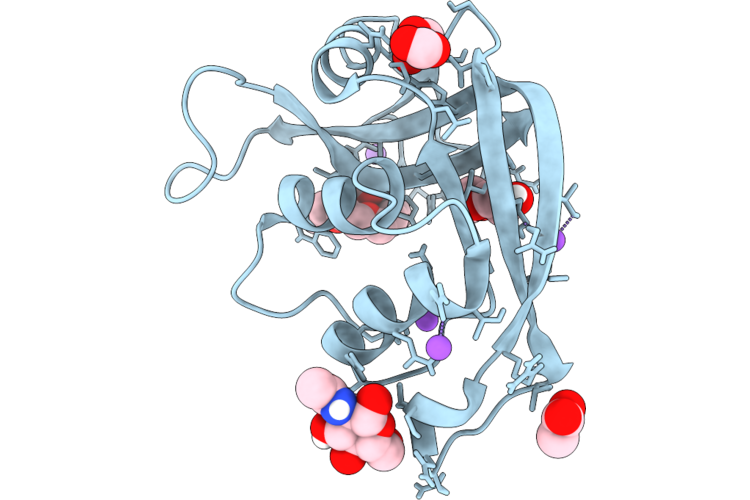
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:0.94 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, FMT, NA |
 |
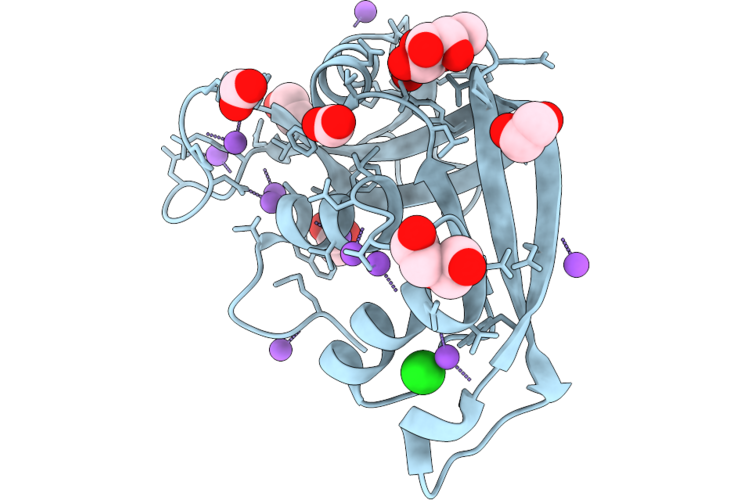
Structural Characterization Of A N-Acetyl-D-Glucosamine-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, NAG, PG4, FMT, PEG, NA |
 |
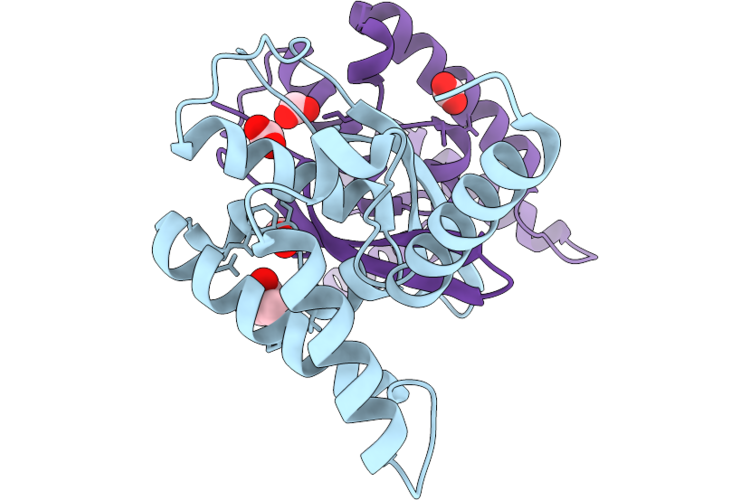
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: GLA, ACT, FMT, PEG, EDO, NA, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: FMT, GOL |
 |
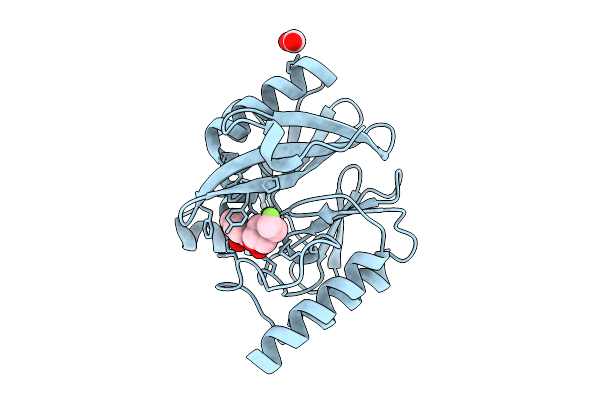
Structural Characterization Of A N-Acetylmuramic Acid-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-04-15 Classification: PROTEIN BINDING Ligands: MUB, AMU, ACT, FMT, NA, CL |
 |
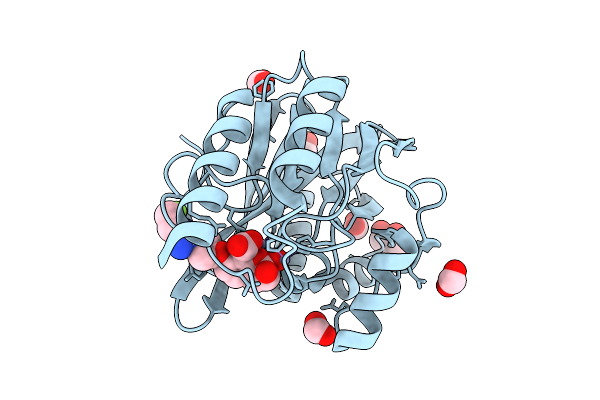
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: ZN, A1EOX, FMT |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: ZN, A1EO0, GOL, FMT |
 |
Protein Structure Of The First Glycoside Hydrolase Family 30, Subfamily 12 Endoxylanase
Organism: Anaerobacterium chartisolvens
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: GOL, FMT, ACY, EDO, NA, PEG |
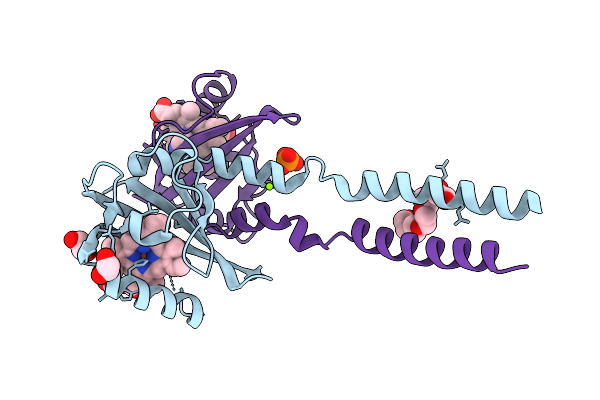 |
Crystal Structure Of Heme Binding Pas Domain From One Component Transcription Factor, Fg214
Organism: Fimbriimonas ginsengisoli gsoil 348
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-25 Classification: TRANSCRIPTION Ligands: HEM, IMD, PO4, CL, PG4, EDO, MG, FMT, POL |
 |
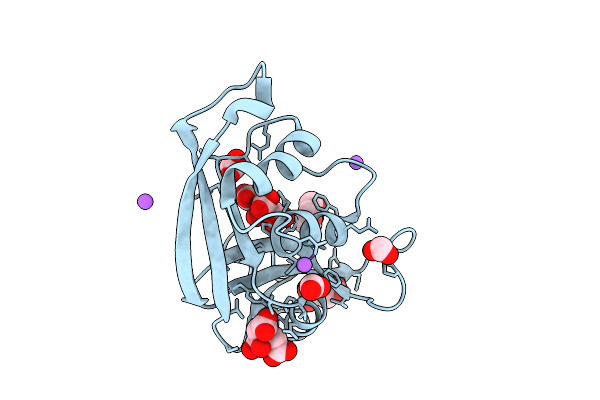
Crystal Structure Of Heme Binding Pas Domain From One Component Transcription Factor, Fg214 Reduced With Dithionite
Organism: Fimbriimonas ginsengisoli gsoil 348
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-25 Classification: TRANSCRIPTION Ligands: HEM, FMT, PO4, POL |
 |
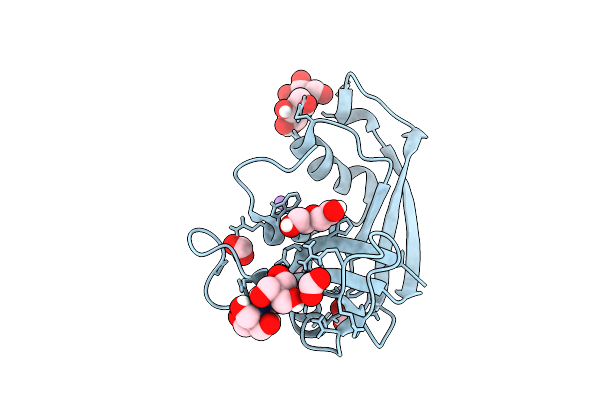
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.17 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: CIT, FMT, PEG, ACT, NA |
 |
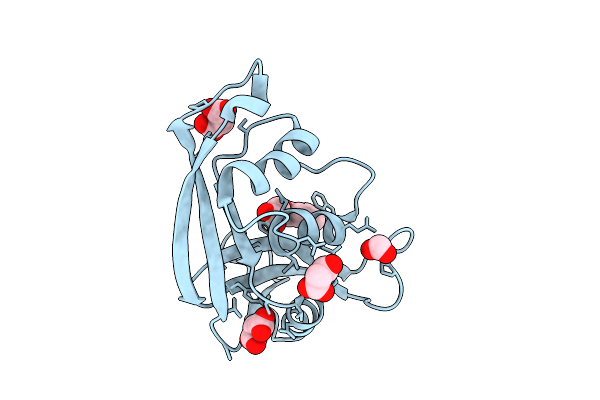
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: B3P, FMT, PEG, NA |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: MLI, PEG, ACT, FMT |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: PEG, ACT, FMT, FUL, FUC |
 |
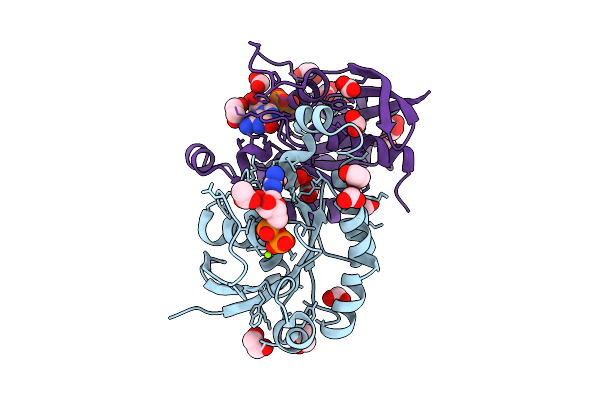
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-18 Classification: STRUCTURAL PROTEIN Ligands: FMT, CL, NA |
 |
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: GDP, MG, FMT, ACT, EDO |

