Search Count: 47
All
Selected
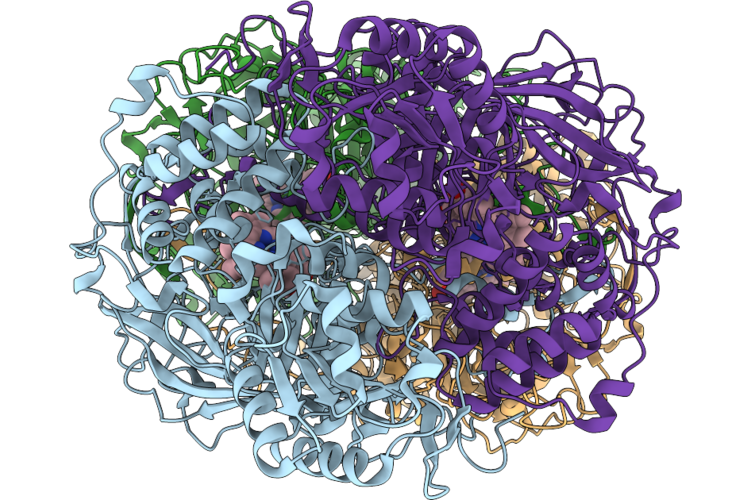 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
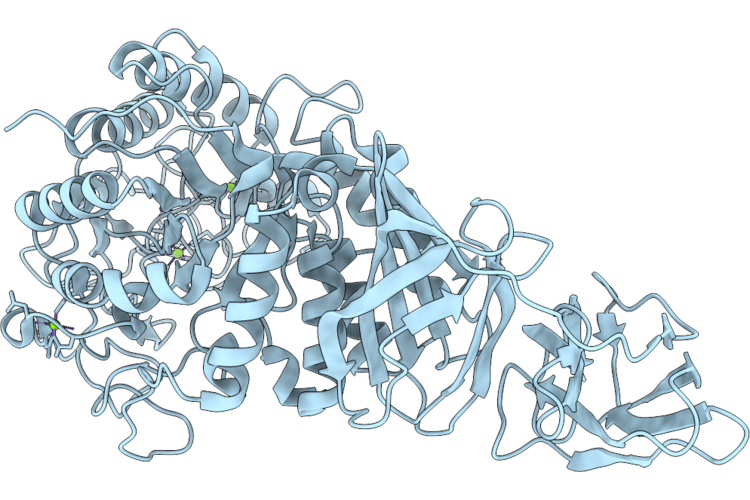
Crystal Structure Of The C-Terminal Of An Alpha-Amylase Family Glycosyl Hydrolase From Vibrio Parahaemolyticus
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: GOL, MG |
 |
Crystal Structure Of The N-Terminal Of An Alpha-Amylase Family Glycosyl Hydrolase From Vibrio Parahaemolyticus
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: MG |
 |
Organism: Streptomyces lividans tk24
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-04-22 Classification: HYDROLASE |
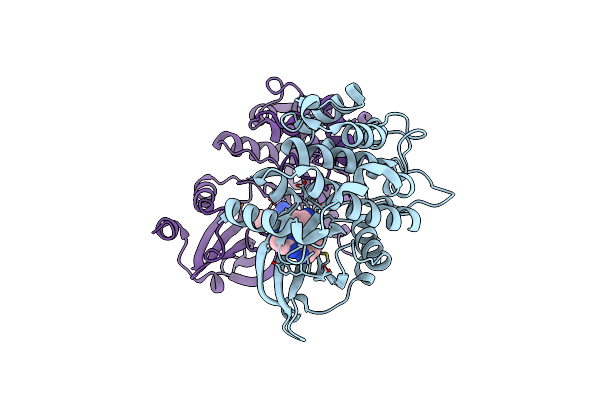 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2P |
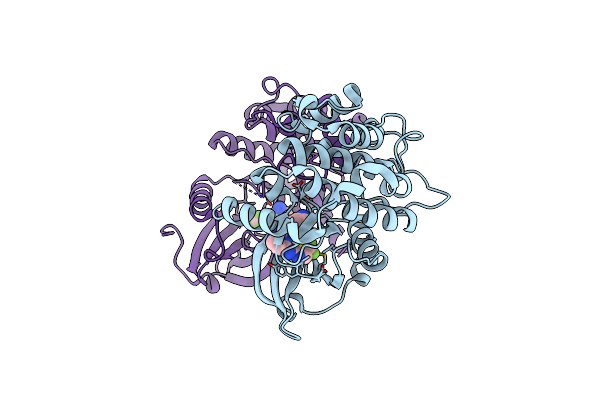 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2Q |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2S |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2S |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2T |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2Q |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2R, EDO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.47 Å Release Date: 2025-04-02 Classification: TRANSFERASE Ligands: A1A2P |
 |
Organism: Vibrio parahaemolyticus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-09-25 Classification: HYDROLASE |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.16 Å Release Date: 2024-03-06 Classification: RNA BINDING PROTEIN |
 |
Crystal Structure Of Isocitrate Dehydrogenase From Campylobacter Corcagiensis
Organism: Campylobacter corcagiensis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-07-05 Classification: OXIDOREDUCTASE Ligands: NAP, MN, SO4, PG4, EDO |
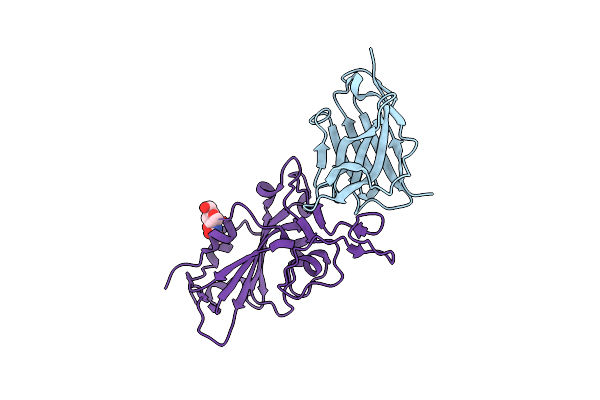 |
Organism: Vicugna pacos, Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-08-10 Classification: VIRAL PROTEIN/ANTIVIRAL PROTEIN Ligands: NAG |
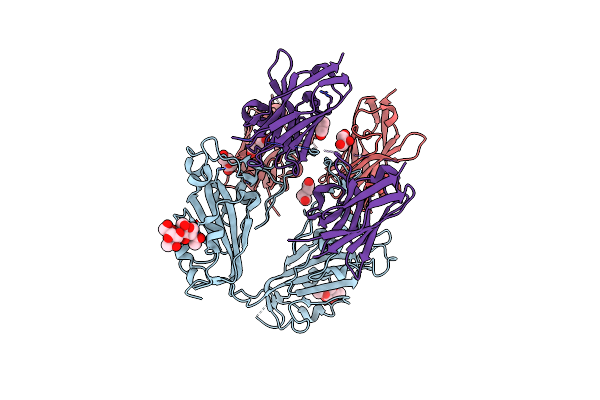 |
Crystallographic Structure Of Two Neutralizing Nanobodies In Complex With Sars-Cov-2 Spike Receptor-Binding Domain (Rbd)
Organism: Severe acute respiratory syndrome coronavirus 2, Vicugna pacos
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2022-08-03 Classification: ANTIVIRAL PROTEIN Ligands: NAG, PEG, GOL |
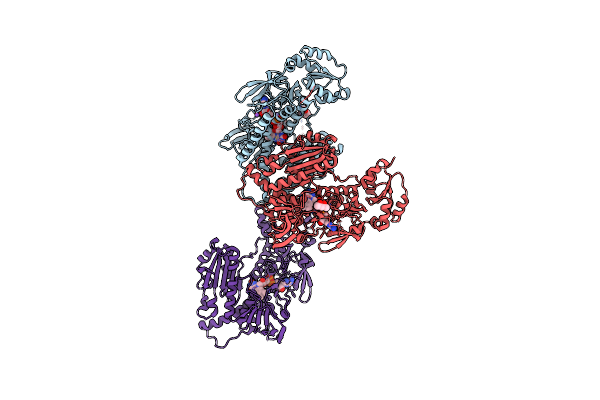 |
Organism: Brugia malayi
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2022-04-06 Classification: OXIDOREDUCTASE Ligands: FAD, MPD, NA, CL |
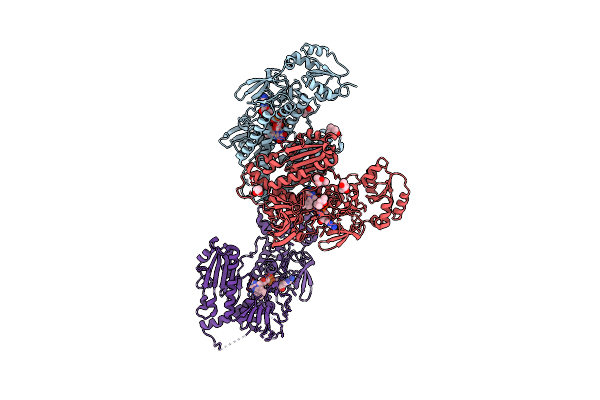 |
Crystal Structure Of Thioredoxin Reductase From Brugia Malayi In Complex With Nadp(H)
Organism: Brugia malayi
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2022-04-06 Classification: FLAVOPROTEIN Ligands: FAD, MPD, ATR |
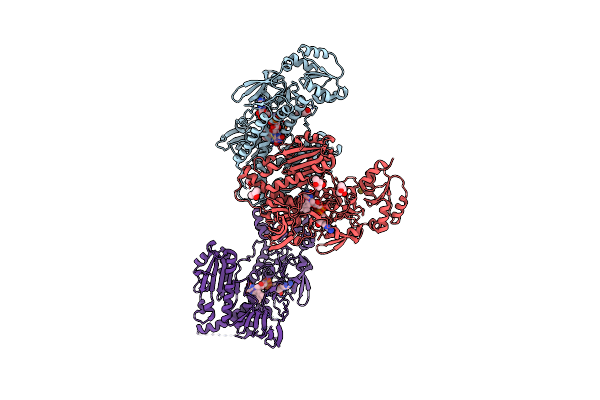 |
Crystal Structure Of Thioredoxin Reductase From Brugia Malayi In Complex With Auranofin
Organism: Brugia malayi
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2022-04-06 Classification: FLAVOPROTEIN Ligands: FAD, ATR, MPD, NA, AU |

