Search Count: 93
All
Selected
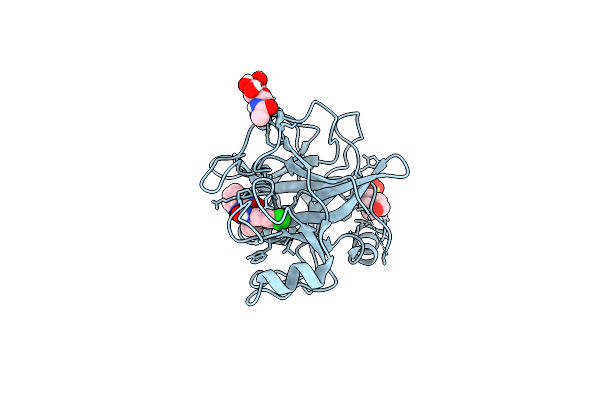 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Complex With Compound 15 ((3'R)-1'-(5-Amino-1-Phenyl-1H-Pyrazole-4-Carbonyl)-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-01 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6X, NAG |
 |
N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: A1C6O |
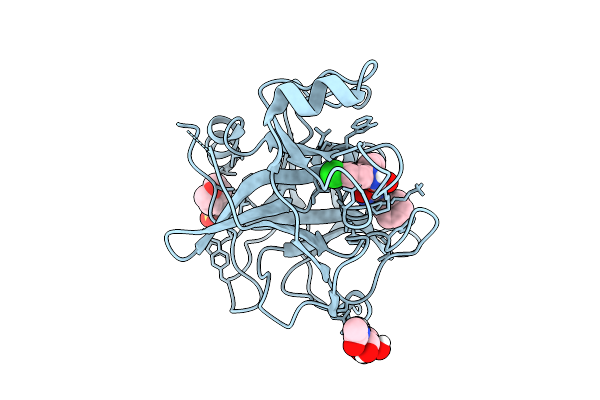 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasmakallikrein Inhibitors Complex With Compound 4 ((3'R)-1'-(5-Amino-1-Benzyl-1H-Pyrazole-4-Carbonyl)-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6Q, NAG |
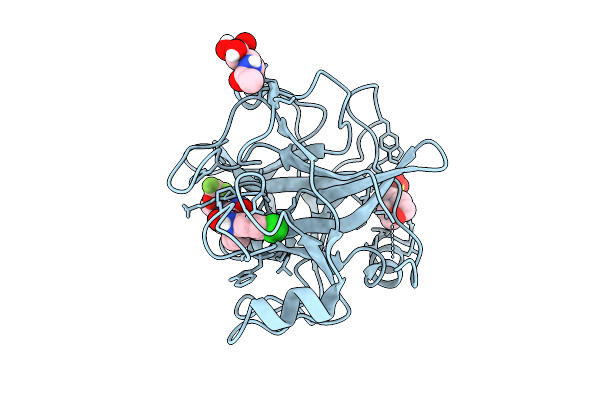 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Compound 25 ((3'R)-1'-{(1P)-5-Amino-1-[2-(Trifluoromethoxy)Phenyl]-1H-Pyrazole-4-Carbonyl}-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6Z, NAG |
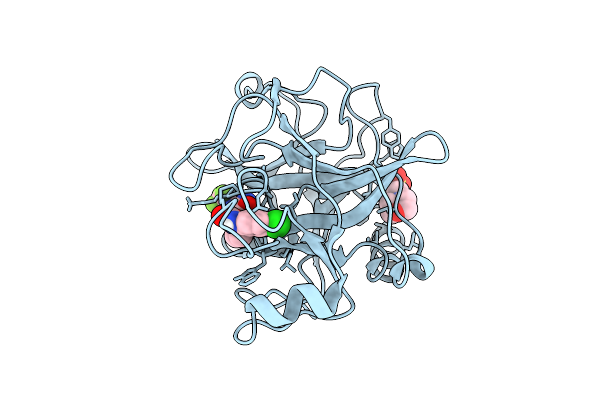 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Compound 13 ((3'R)-1'-{5-Amino-1-[(2S)-1,1,1-Trifluorobutan-2-Yl]-1H-Pyrazole-4-Carbonyl}-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C7M |
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-03-04 Classification: TRANSFERASE |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-08 Classification: HYDROLASE Ligands: AMP, MG, NA |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-08 Classification: HYDROLASE Ligands: MG, DCM, DGP, CO |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2023-11-08 Classification: HYDROLASE Ligands: EPE, SO4, NH4, C5P |
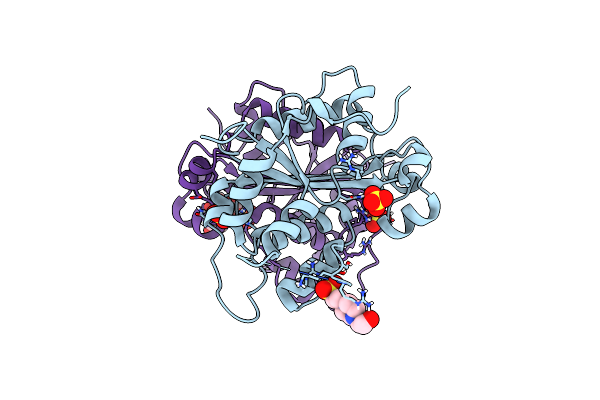 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-11-08 Classification: HYDROLASE Ligands: EPE, SO4, U5P |
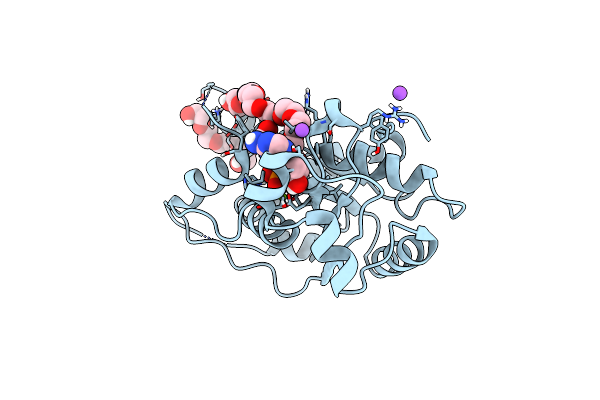 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-11-08 Classification: HYDROLASE Ligands: 1PE, NA, D5M, MG |
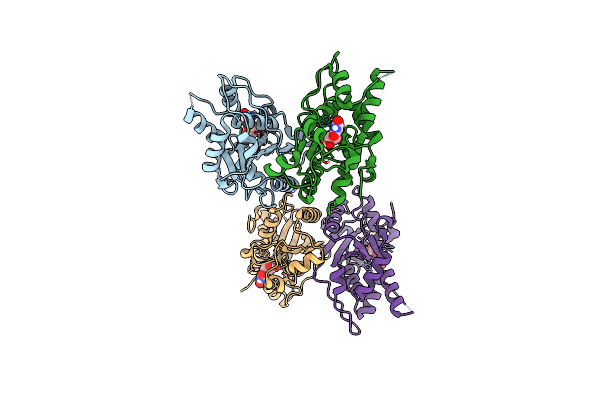 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-11-08 Classification: HYDROLASE Ligands: MG, URI |
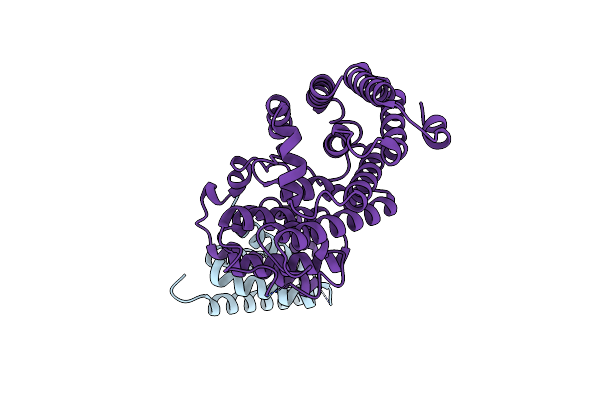 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-09-27 Classification: SIGNALING PROTEIN |
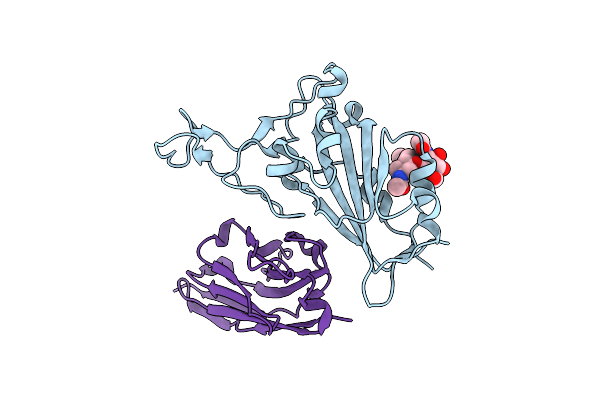 |
Crystal Structure Of The Sars-Cov-2 Neutralizing Vhh 7A9 Bound To The Spike Receptor Binding Domain
Organism: Severe acute respiratory syndrome coronavirus 2, Lama glama
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-08-16 Classification: ANTIVIRAL PROTEIN |
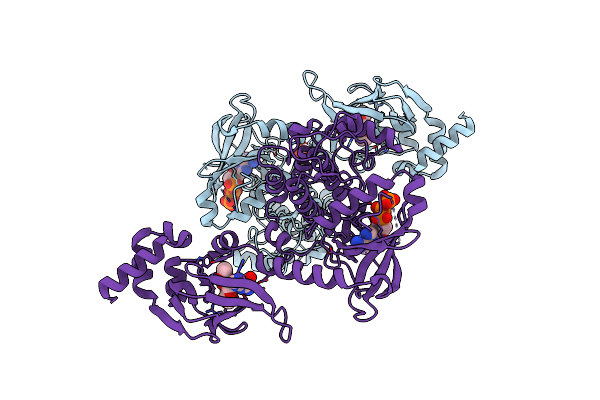 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2023-06-14 Classification: SIGNALING PROTEIN Ligands: PCG, ATP, EDO, CL |
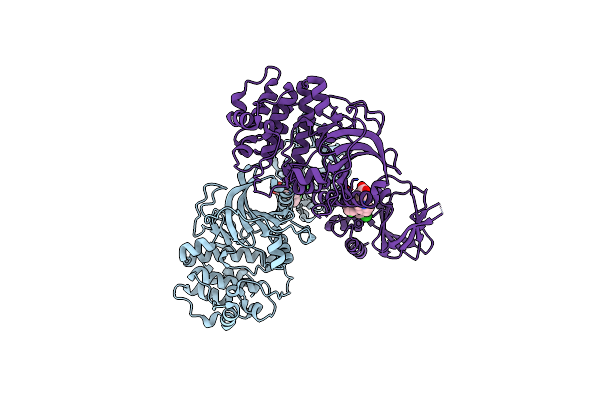 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2023-06-14 Classification: TRANSFERASE Ligands: EX6 |
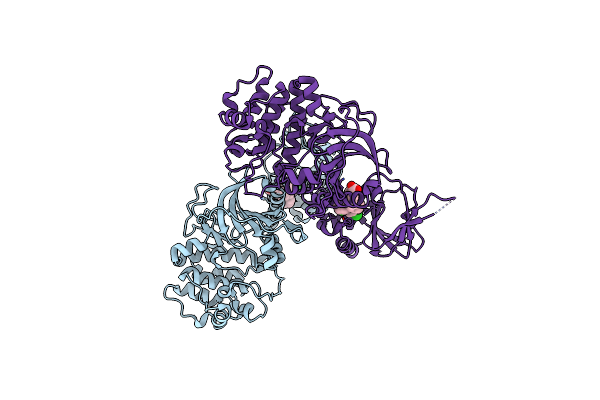 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2023-06-14 Classification: TRANSFERASE Ligands: EXZ, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2023-06-14 Classification: TRANSFERASE Ligands: EZJ, CL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-04-26 Classification: SIGNALING PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2022-08-24 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: B4I |

