Search Count: 1,279
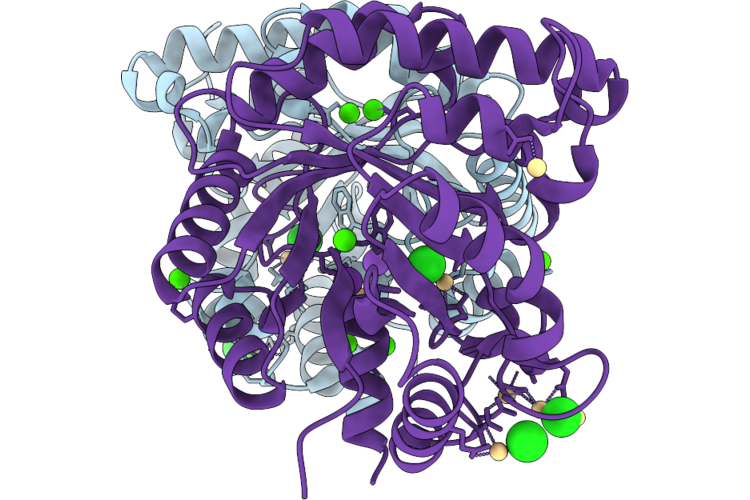 |
Organism: Cellulomonas fimi
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-05-06 Classification: UNKNOWN FUNCTION Ligands: CD, CA, K, CL |
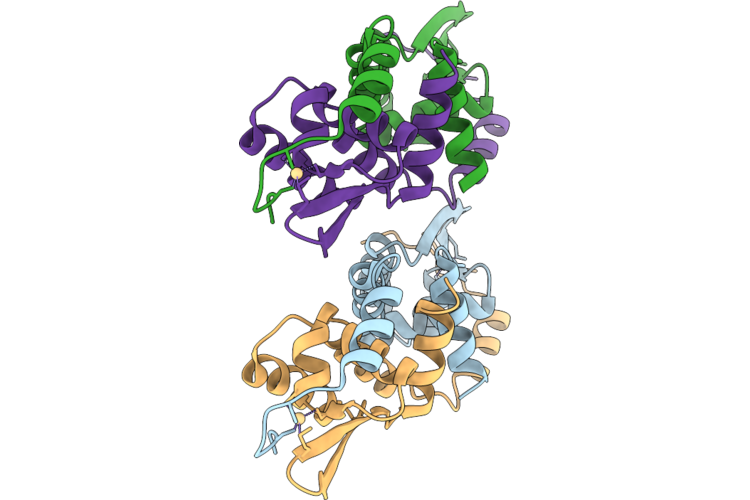 |
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: CD |
 |
Crystal Structure Of Human S-Adenosyl-L-Homocysteine Hydrolase In Complex With Adenosine And Cadmium Ions.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: NAD, ADN, CD, K |
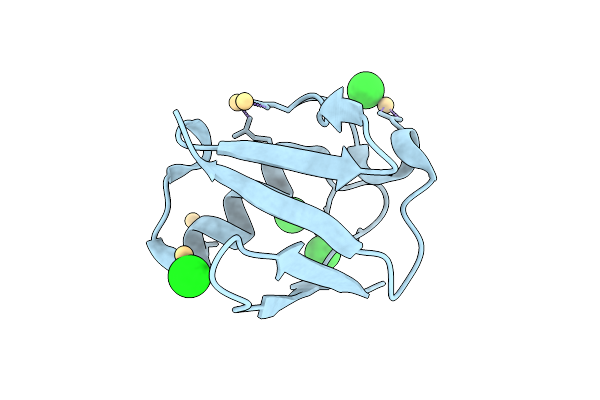 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-15 Classification: SIGNALING PROTEIN Ligands: CD, CL |
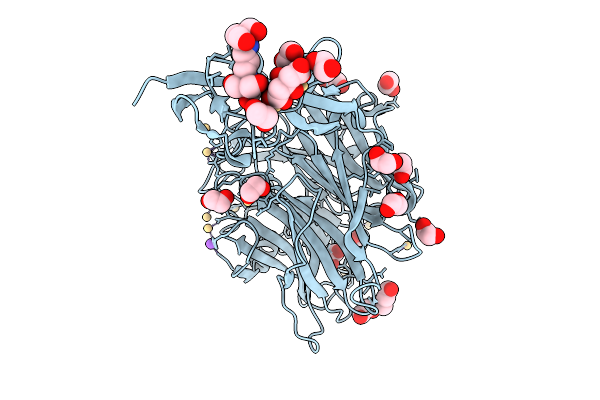 |
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: CD, NA, B3P, CIT, EDO, PGE |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) Bound To Neu5Ac2En (Dana)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: DAN, EDO, PEG, CD |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) Bound To Neu5Ac (Nana)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: CD, SLB, EDO, PEG, PGE |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) D165A Mutant Bound To 3'-Sialyllactose (Only Neu5Ac Visible)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: SIA, GOL, PEG, CD, NA |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |
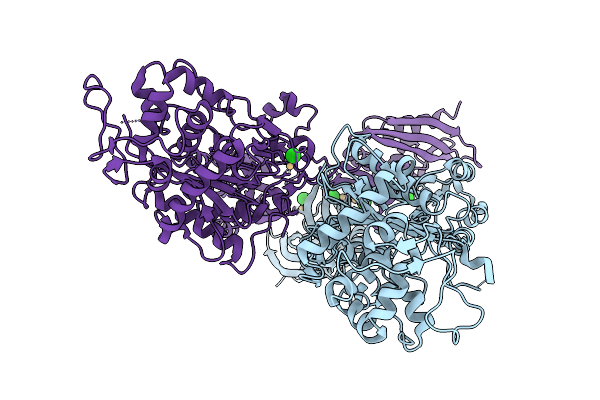 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |
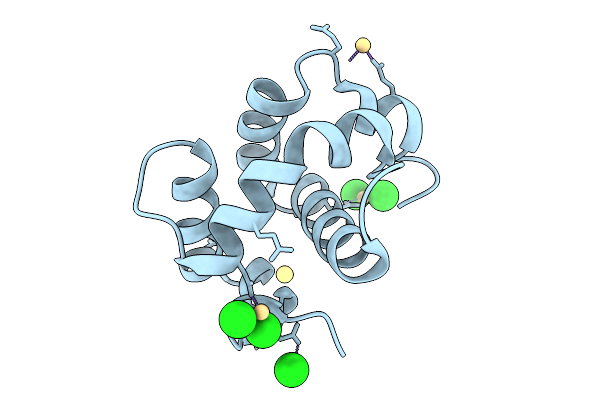 |
Organism: Anopheles culicifacies
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-01-28 Classification: LIPID BINDING PROTEIN Ligands: CD, CL |
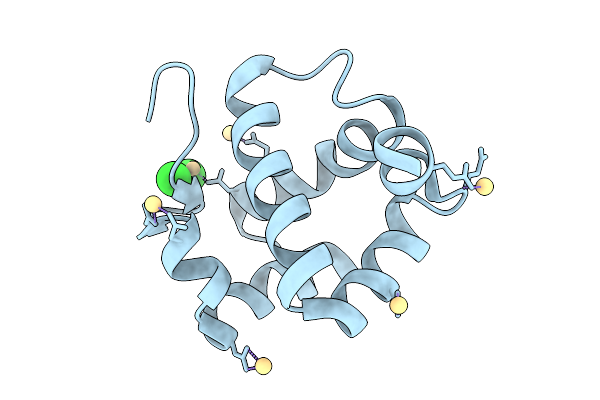 |
Crystal Structure Sensory Appendage Proteins 2 From Anopheles Culicifacies In Space Group P212121
Organism: Anopheles culicifacies
Method: X-RAY DIFFRACTION Resolution:1.14 Å Release Date: 2026-01-28 Classification: LIPID BINDING PROTEIN Ligands: CD, CL |
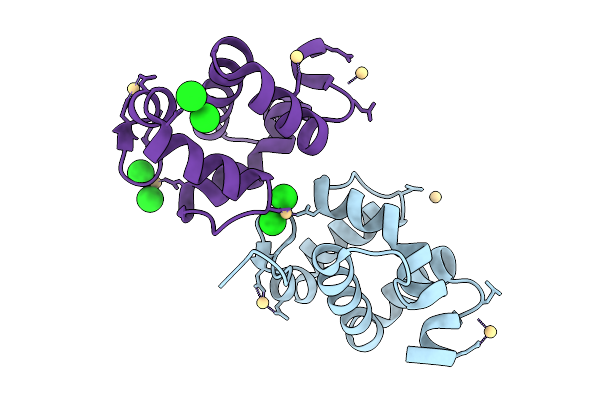 |
Crystal Structure Sensory Appendage Proteins 2 From Anopheles Culicifacies In Space Group P212121 And 2 Molecule Per Asu
Organism: Anopheles culicifacies
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-01-28 Classification: LIPID BINDING PROTEIN Ligands: CD, CL |
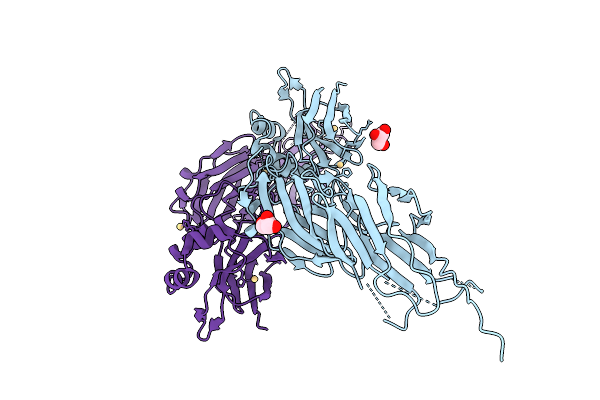 |
Organism: Streptococcus sanguinis
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-01-21 Classification: CELL ADHESION Ligands: CD, GOL |
 |
Organism: Acinetobacter baylyi adp1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-11-12 Classification: PROTEIN BINDING Ligands: ACT, CD |
 |
Crystal Structure Of The Cyp154C5 F92A/R114A/T248D/E282A Variant From Nocardia Farcinica
Organism: Nocardia farcinica ifm 10152
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-11-05 Classification: OXIDOREDUCTASE Ligands: TES, HEM, CD, CL, K, GOL |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-22 Classification: DE NOVO PROTEIN Ligands: CD, CL |
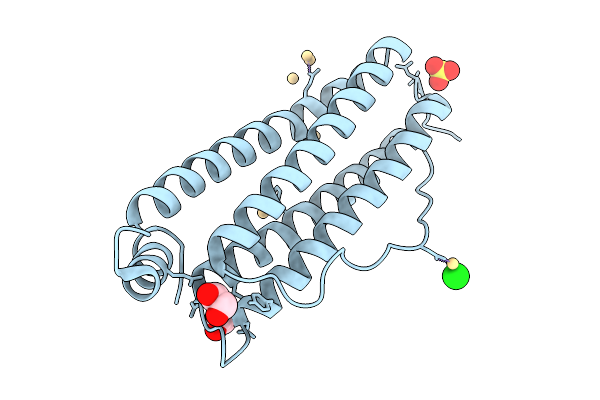 |
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-09-24 Classification: METAL BINDING PROTEIN Ligands: EDO, CD, SO4, CL |
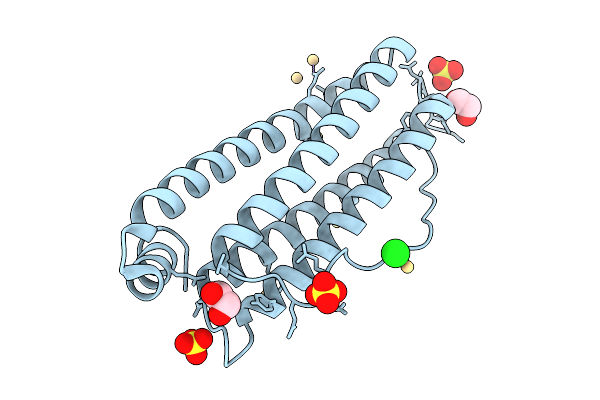 |
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-09-24 Classification: METAL BINDING PROTEIN Ligands: EDO, CD, SO4, CL |
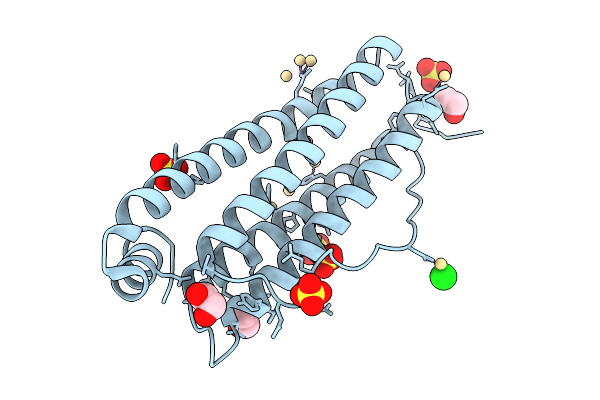 |
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-09-24 Classification: METAL BINDING PROTEIN Ligands: EDO, CD, SO4, CL |

