Search Count: 302
All
Selected
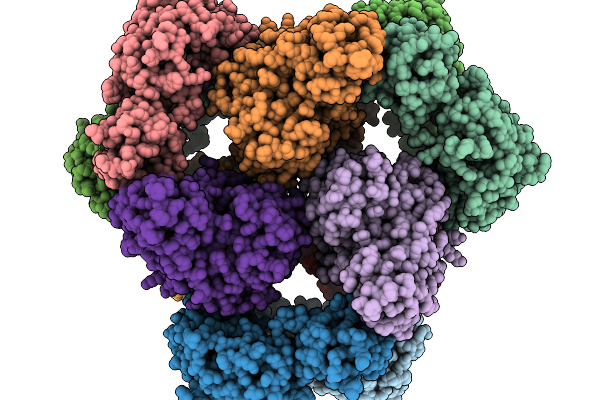 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSFERASE |
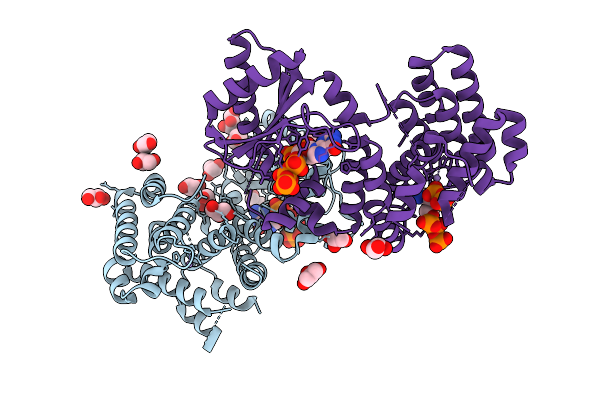 |
Crystal Structure Of The Relseq N-Terminal Domain From Streptococcus Equisimilis In Complex With Pppgpp
Organism: Streptococcus dysgalactiae subsp. equisimilis
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: GOL, MN, 0O2, CL |
 |
Organism: Aquimarina sp. 2-a2
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-18 Classification: SUGAR BINDING PROTEIN |
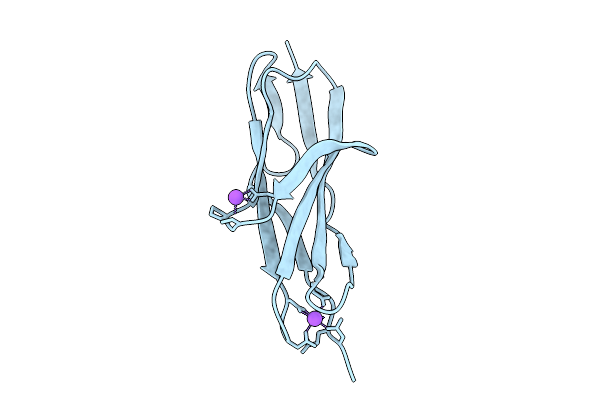 |
Organism: Aquimarina sp. 2-a2
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: SUGAR BINDING PROTEIN |
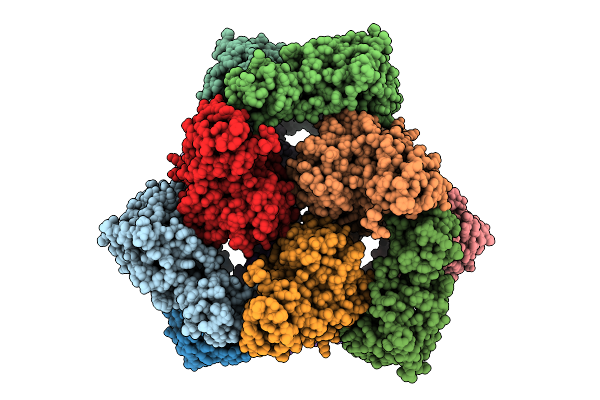 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: TRANSFERASE |
 |
Cryogenic Temperature Crystal Structure Of Nmar_1308 Protein At 2.46 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2025-07-09 Classification: STRUCTURAL PROTEIN |
 |
Cryogenic Temperature Crystal Structure Of Nmar_1308 Protein At 2.71 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2025-07-09 Classification: STRUCTURAL PROTEIN |
 |
Cryogenic Temperature Crystal Structure Of Nmar_1308 Protein At 3.29 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:3.29 Å Release Date: 2025-07-09 Classification: STRUCTURAL PROTEIN |
 |
Ambient Temperature Crystal Structure Of Nmar_1308 Protein At 3.33 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:3.33 Å Release Date: 2025-07-09 Classification: STRUCTURAL PROTEIN |
 |
Ambient Temperature Crystal Structure Of Nmar_1308 Protein At 3.40 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2025-07-09 Classification: STRUCTURAL PROTEIN |
 |
Proline Utilization A With The Fadh- N5 Atom Covalently Modified By Proline
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-06-04 Classification: OXIDOREDUCTASE Ligands: NAD, PGE, FDA, A1CAB, PRO, MG, SO4, A1AT4 |
 |
Proline Utilization A With The Covalent Acyl-Enzyme Intermediate In The Aldehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-06-04 Classification: OXIDOREDUCTASE Ligands: NAD, SO4, PGE, A1AUG, MG, FDA, PEG |
 |
Proline Utilization A Complexed With The Substrate L-Glutamate Gamma-Semialdehyde In The Aldehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti sm11
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2025-06-04 Classification: OXIDOREDUCTASE Ligands: NAD, FDA, PGE, PEG, A1AUG, FMT, MG, SO4 |
 |
Cryogenic Temperature Crystal Structure Of Nmar_1308 Protein At 2.96 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:2.96 Å Release Date: 2025-04-30 Classification: STRUCTURAL PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: BIOSYNTHETIC PROTEIN |
 |
Proline Utilization A Complexed With The Product L-Glutamate In The Aldehyde Dehydrogenase Active Site
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-03-12 Classification: OXIDOREDUCTASE Ligands: GGL, SO4, PGE, FAD, PEG |
 |
Structure Of Proline Utilization A Complexed With 2,3-Dihydro-1,4-Benzodioxine-5-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, 53E, FMT, NAD, SO4, MG, PGE, PG4 |
 |
Structure Of Proline Utilization A Complexed With Quinoline-2-Carboxylic Acid
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, QNC, PEG, FMT, SO4, MG, PG4 |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: PGE, PEG, PYS, FMT, SO4, FAD |
 |
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: FAD, PGE, FMT, PEG, PG4, A1BDA, SO4 |

