Search Count: 3,252
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-15 Classification: HORMONE Ligands: CRS, ZN, CL |
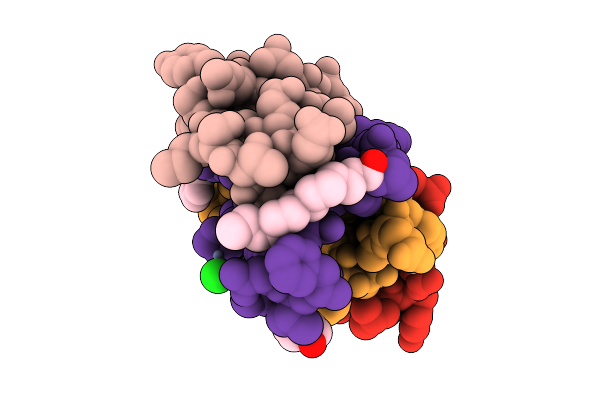 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-04-15 Classification: HORMONE Ligands: IPH, ZN, CL, MYR |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
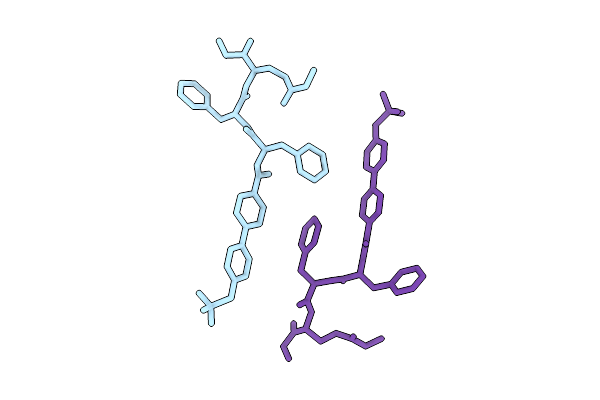 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
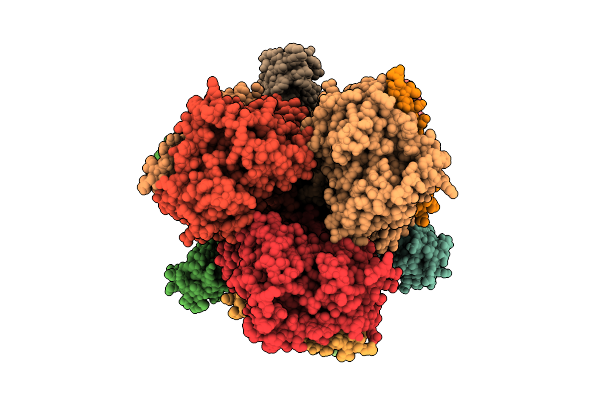 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.17 Å Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: CL |
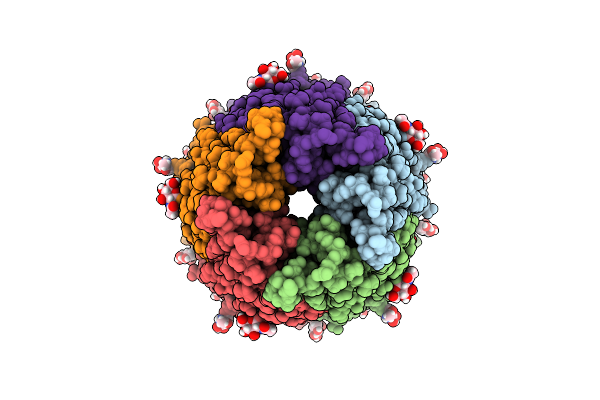 |
Organism: Octopus bimaculoides
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: NAG, STR |
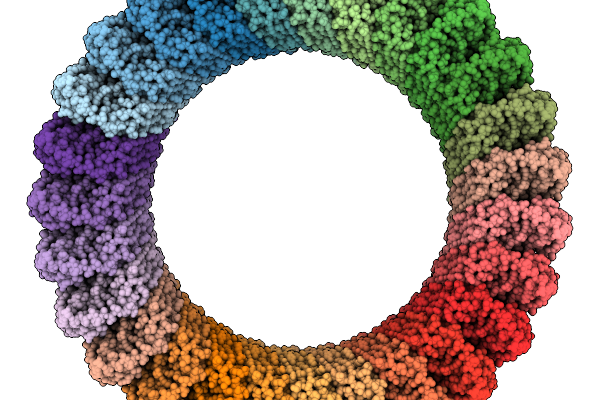 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.64 Å Release Date: 2026-03-25 Classification: IMMUNE SYSTEM |
 |
Structure Of The Ybjp Lipoprotein Bound To The Macab-Tolc Tripartite Efflux Pump
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.48 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: CL, Z41 |
 |
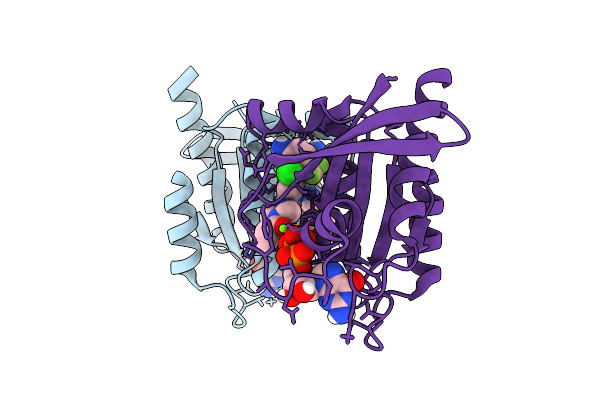
Structure Of Kras-G12C Bound To 1-[(4Ar,10P,13R)-10-[5-Amino-4-Fluoro-3-Methyl-2-(Trifluoromethyl)Phenyl]-11-Chloro-9-Fluoro-1,2,4A,5-Tetrahydropyrazino[1',2':4,5][1,4]Oxazino[2,3-C]Quinolin-3(4H)-Yl]Prop-2-En-1-One (Compound 15)
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2026-03-11 Classification: ONCOPROTEIN Ligands: MG, GDP, A1CII, PEG, GOL, DMS, ACT |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-03-11 Classification: ONCOPROTEIN Ligands: A1CQ2, GDP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2026-03-11 Classification: ONCOPROTEIN Ligands: GDP, MG, DMS, EDO, A1AWR, PEG |
 |
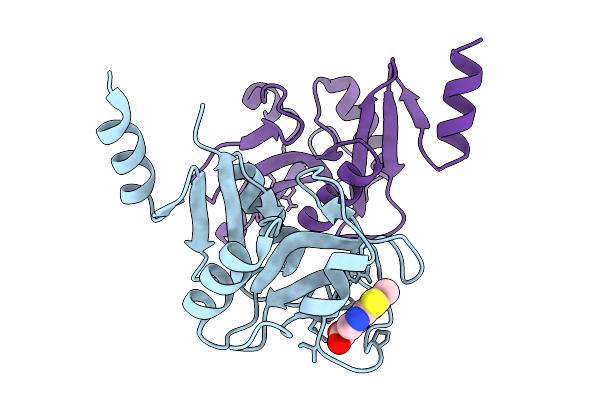
Cryo-Em Structure Of Self-Assembled Zymomonas Mobilis Levansucrase Nanotube
Organism: Zymomonas mobilis subsp. mobilis atcc 10988
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSFERASE |
 |
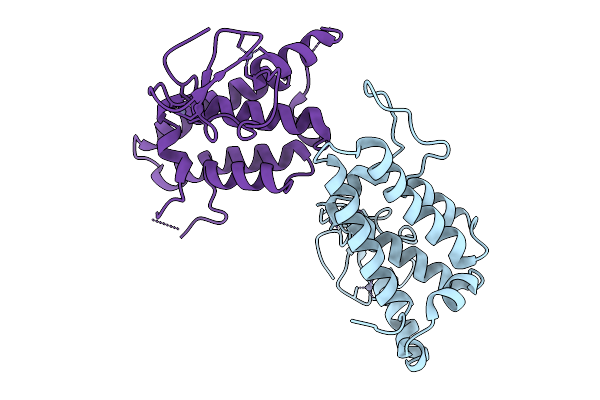
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2026-02-25 Classification: SUGAR BINDING PROTEIN Ligands: A1JHV, CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION Ligands: ZN |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: CHAPERONE Ligands: MG, ATP, K, A1AS6 |
 |
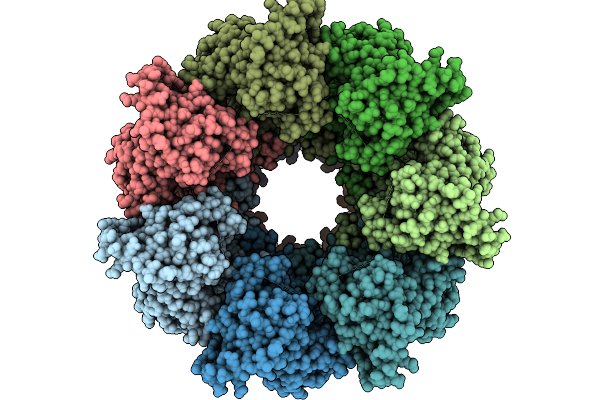
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: CHAPERONE |
 |
E. Coli Sr1 Single-Ring Groel Templated Into Pseudo-Double-Ring Complex With Pbz1587 Inhibitor
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: CHAPERONE Ligands: A1AS6 |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: CHAPERONE Ligands: K, ATP, MG, ADP |

