Search Count: 346
All
Selected
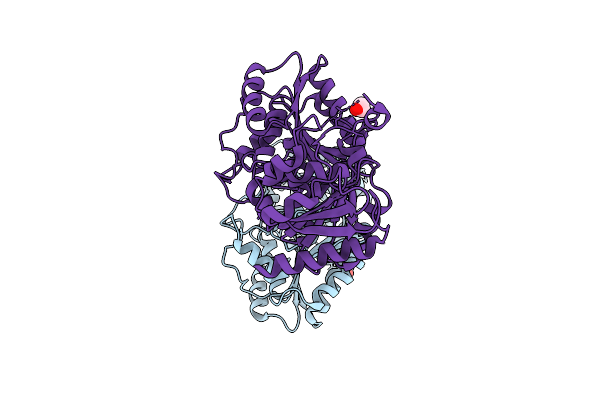 |
Organism: Agrobacterium tumefaciens
Method: X-RAY DIFFRACTION Resolution:3.13 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: EDO |
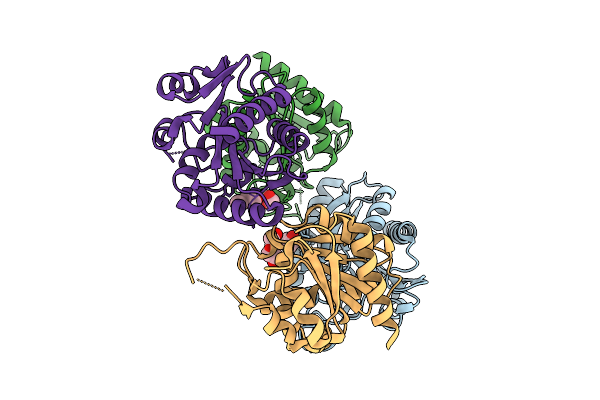 |
Crystal Structure Of The C-Terminal Domain Of Acvb From Agrobacterium Tumefaciens
Organism: Agrobacterium tumefaciens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: PEG |
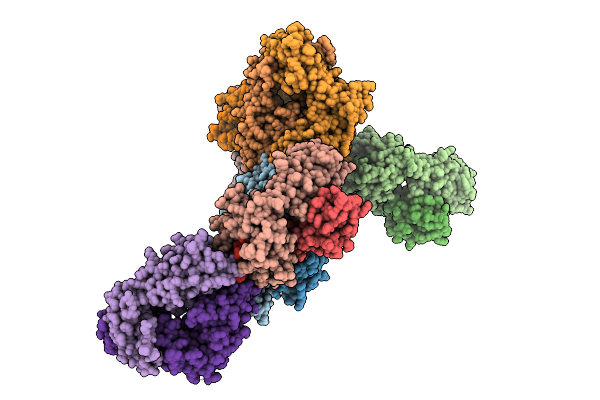 |
Human T Cell Receptor (Trav24*01/Trbv27*01) In Complex With Hla-C*1202 And Iy11 Peptide
Organism: Homo sapiens, Human immunodeficiency virus type 1 (z2/cdc-z34 isolate)
Method: X-RAY DIFFRACTION Resolution:2.98 Å Release Date: 2025-09-17 Classification: IMMUNE SYSTEM |
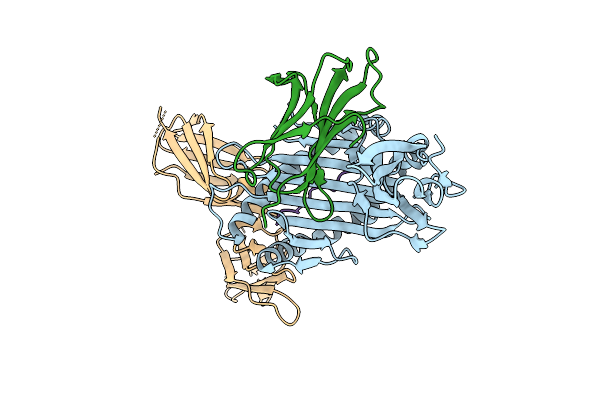 |
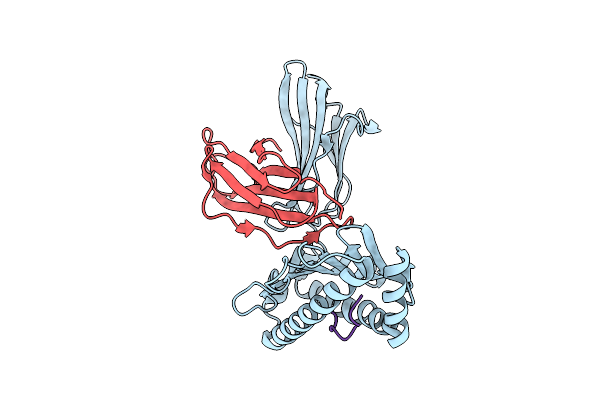
Killer Cell Immunoglobulin-Like Receptor 2Dl2 In Complex With Hla-C*1202 And Iy10 Peptide
Organism: Homo sapiens, Human immunodeficiency virus type 1 (z2/cdc-z34 isolate)
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-09-17 Classification: IMMUNE SYSTEM |
 |
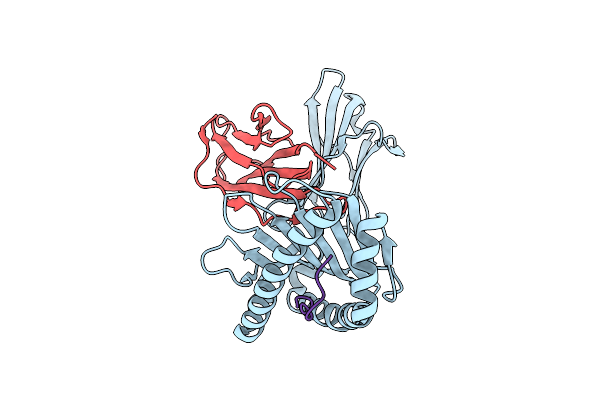
Organism: Homo sapiens, Human immunodeficiency virus type 1 (z2/cdc-z34 isolate)
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-09-17 Classification: IMMUNE SYSTEM |
 |
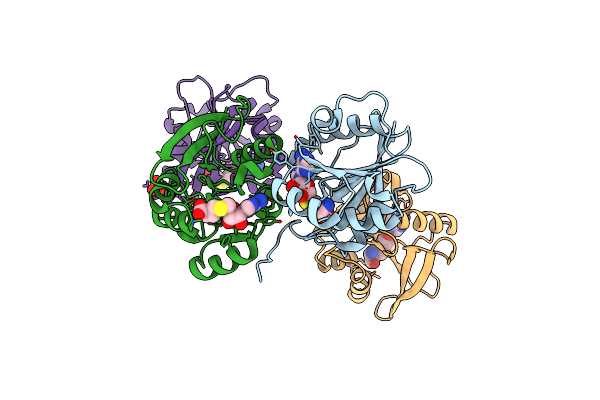
Organism: Homo sapiens, Human immunodeficiency virus type 1 (z2/cdc-z34 isolate)
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-09-17 Classification: IMMUNE SYSTEM |
 |
Organism: Agrobacterium tumefaciens (strain c58)
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-04-23 Classification: TRANSFERASE Ligands: SAH, SO4 |
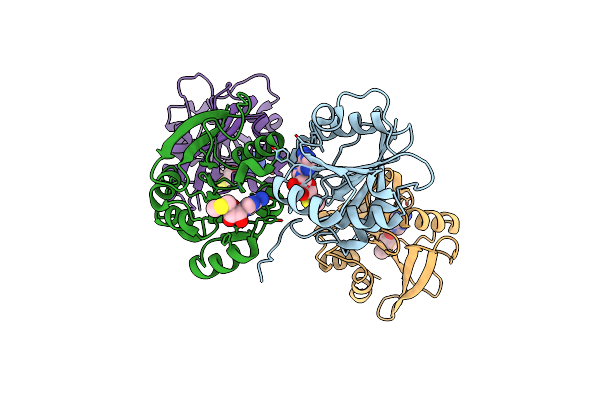 |
Crystal Structure Of Agrobacterium Tumefaciens Pmta With 5'-Methylthioadenosine
Organism: Agrobacterium tumefaciens (strain c58)
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2025-04-23 Classification: TRANSFERASE Ligands: MTA |
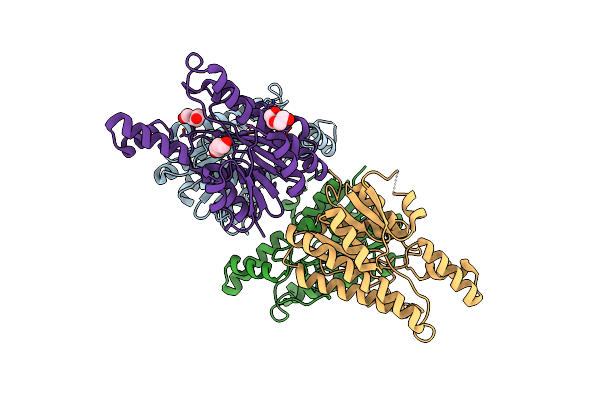 |
Crystal Structure Of L-2-Keto-3-Deoxyfuconate 4-Dehydrogenase From Herbaspillum Huttiense (Apo Form)
Organism: Herbaspirillum huttiense
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2024-07-10 Classification: OXIDOREDUCTASE Ligands: PEG |
 |
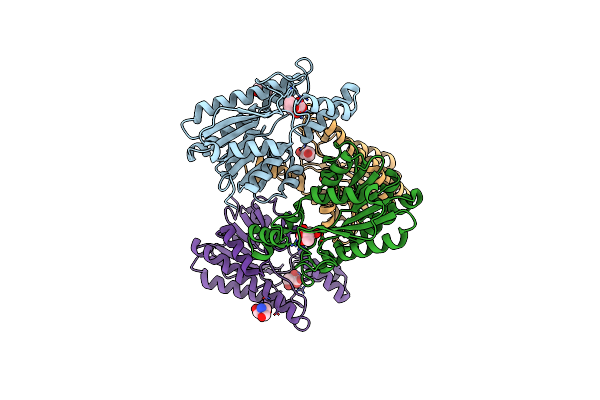
Crystal Structure Of L-2-Keto-3-Deoxyfuconate 4-Dehydrogenase Bound To Nad(H) And Sulfate Ion
Organism: Herbaspirillum huttiense
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-07-10 Classification: OXIDOREDUCTASE Ligands: NAD, SO4, PEG, TRS, GOL |
 |
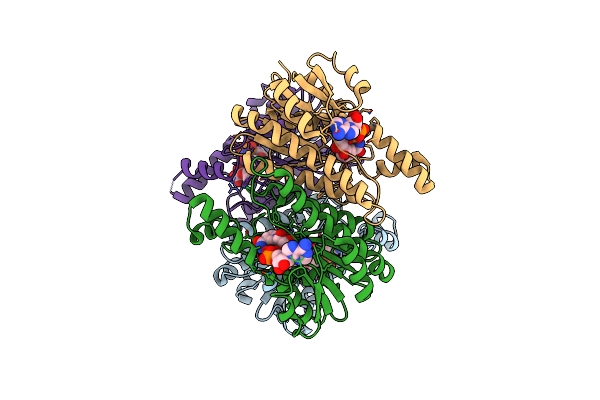
Crystal Structure Of L-2-Keto-3-Deoxyfuconate 4-Dehydrogenase Bound To L-Kdf Or L-2,4-Dkdf
Organism: Herbaspirillum huttiense
Method: X-RAY DIFFRACTION Resolution:1.29 Å Release Date: 2024-07-10 Classification: OXIDOREDUCTASE Ligands: TRS, A1LXA, A1LXB, PEG |
 |
Crystal Structure Of L-2-Keto-3-Deoxyfuconate 4-Dehydrogenase Bound To L-2,4-Dkdf And Nadh
Organism: Herbaspirillum huttiense
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2024-07-10 Classification: OXIDOREDUCTASE Ligands: NAD, A1LXB |
 |
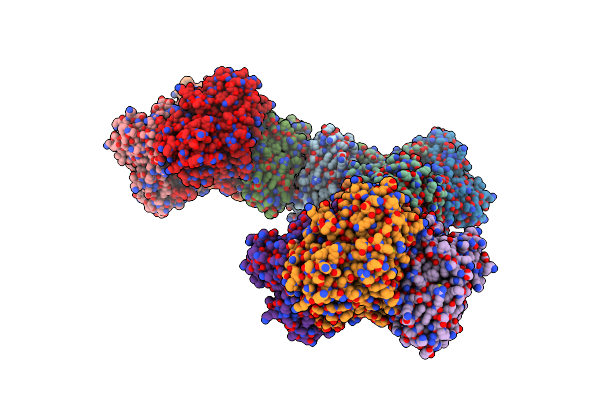
Crystal Structure Of L-2-Keto-3-Deoxyfuconate 4-Dehydrogenase Bound To D-Kdp
Organism: Herbaspirillum huttiense
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2024-07-10 Classification: OXIDOREDUCTASE Ligands: TRS, A1LXD, PEG |
 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-02-14 Classification: LYASE Ligands: TVJ, GOL, EDO |
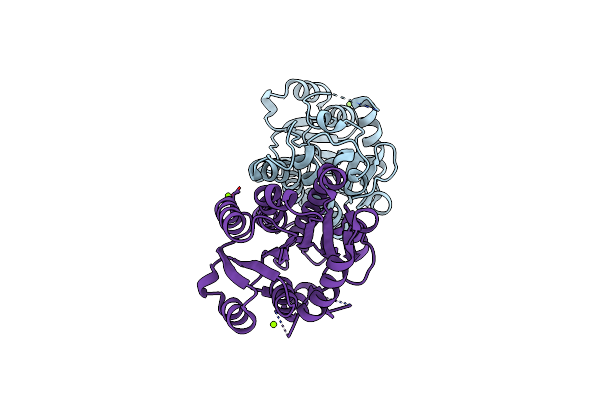 |
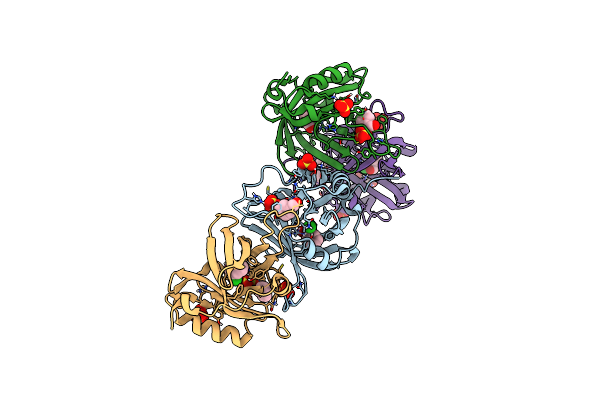
Discovery And Characterization Of A New Carbonyl Reductase From Rhodotorula Toluroides Reducing Fluoroketones, And X-Ray Analysis Of The Variant By Rational Engineering
Organism: Rhodotorula toruloides
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2024-02-07 Classification: OXIDOREDUCTASE Ligands: MG |
 |
Organism: Oxidus gracilis
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-01-24 Classification: LYASE Ligands: HBX, CL, SO4, GOL |
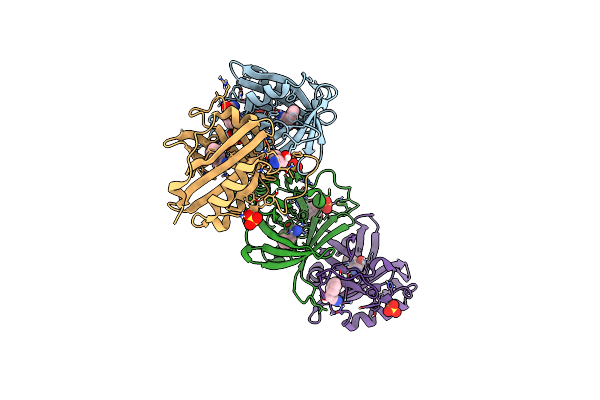 |
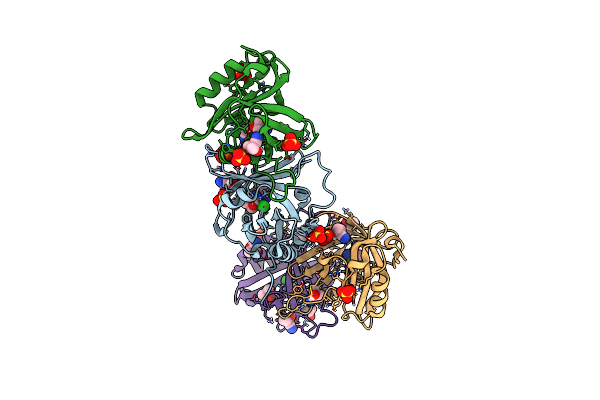
Hydroxynitrile Lyase From The Millipede, Oxidus Gracilis Bound With (R)-(+)-Alpha-Hydroxybenzene-Acetonitrile
Organism: Oxidus gracilis
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-01-24 Classification: LYASE Ligands: MXN, SO4 |
 |
Organism: Oxidus gracilis
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-01-24 Classification: LYASE Ligands: CNH, CL, SO4 |
 |
Hydroxynitrile Lyase From The Millipede, Oxidus Gracilis Complexed With (R)-2-Chloromandelonitrile
Organism: Oxidus gracilis
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-01-24 Classification: LYASE Ligands: IJ5, GOL, SO4 |
 |
Organism: Oxidus gracilis
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-01-17 Classification: LYASE Ligands: SO4, CL |

