Search Count: 23,039
All
Selected
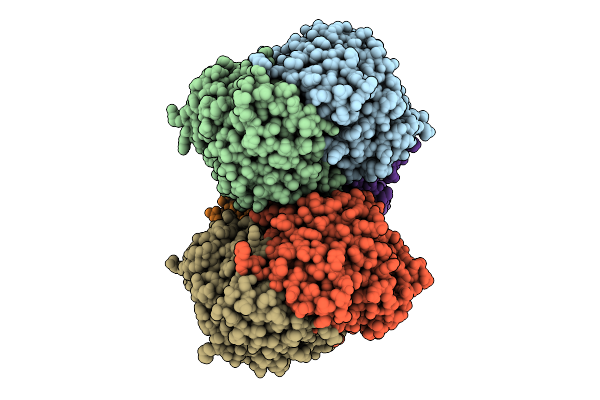 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
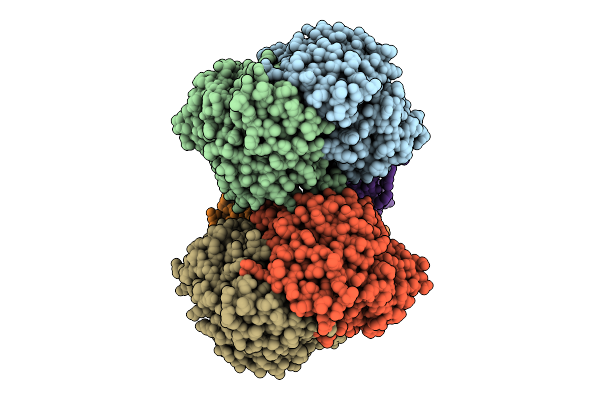 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
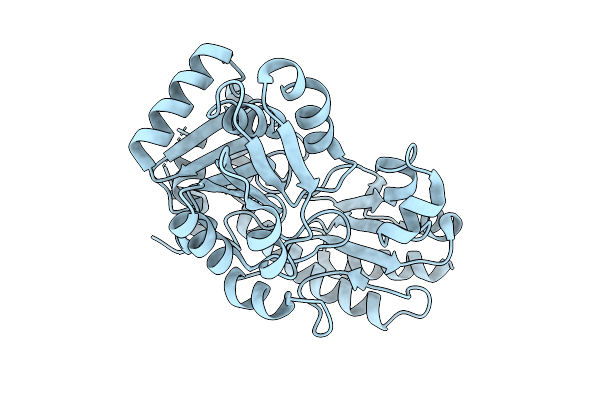 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2026-04-15 Classification: LYASE |
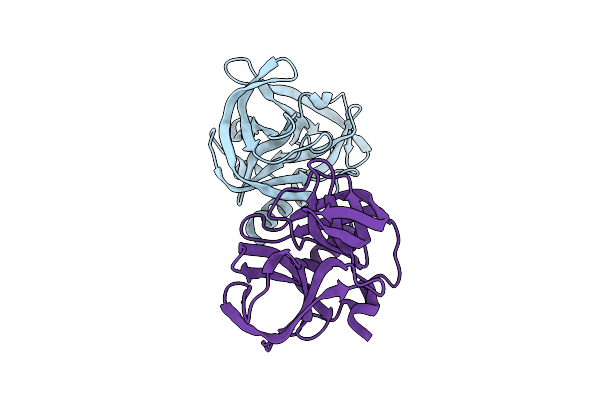 |
Crystal Structure Of Enterovirus D68 3C Protease Determined Via Sulfur Phasing
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN Ligands: CL |
 |
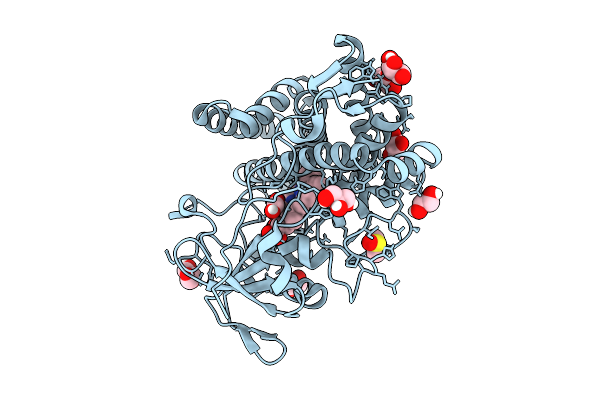
Fphi, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases I, In Complex With Carbamoyl Fluoride Compound 18
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-01 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: A1BYJ, CA |
 |
Crystal Structure Of The Cyp105D18 Double Mutant F184A/F191A From Streptomyces Laurentii
Organism: Streptomyces laurentii
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, GOL, DMS, EDO |
 |
Crystal Structure Of The Cyp105D18 Mutant F191A From Streptomyces Laurentii
Organism: Streptomyces laurentii
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, GOL |
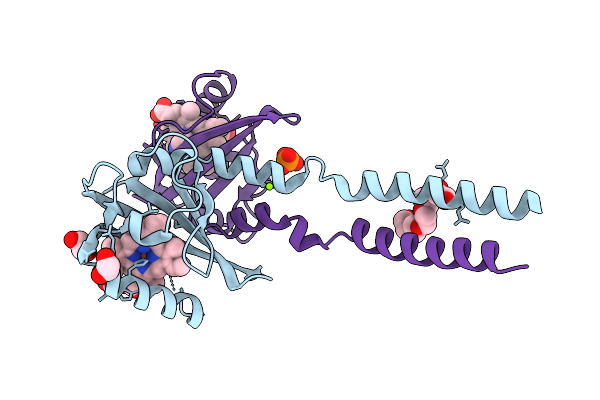 |
Crystal Structure Of Heme Binding Pas Domain From One Component Transcription Factor, Fg214
Organism: Fimbriimonas ginsengisoli gsoil 348
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-25 Classification: TRANSCRIPTION Ligands: HEM, IMD, PO4, CL, PG4, EDO, MG, FMT, POL |
 |
Crystal Structure Of Heme Binding Pas Domain From One Component Transcription Factor, Fg214 Reduced With Dithionite
Organism: Fimbriimonas ginsengisoli gsoil 348
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-25 Classification: TRANSCRIPTION Ligands: HEM, FMT, PO4, POL |
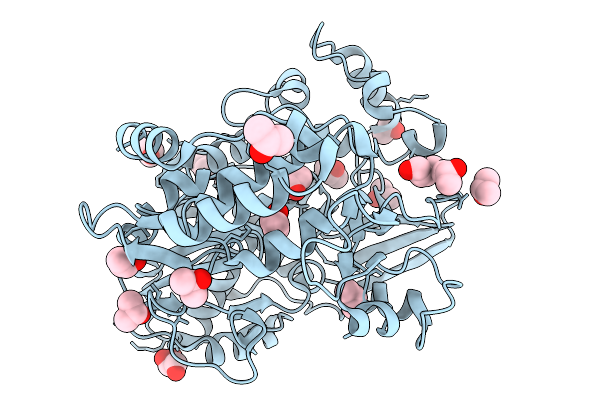 |
Pandda Analysis Group Deposition Of Ground-State Model Of Enterovirus D68 3Dpol
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: IPA, PEG, GOL |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z111507846
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, AWP, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1143279263
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, UR7, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1198177230
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, K6U, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1201620232
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, NZ1, MG |
 |
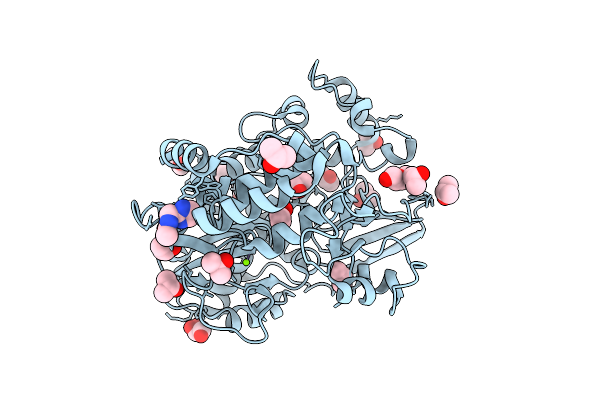
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1203107138
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, JH1, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1203191681
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, A1AJ1, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1220452176
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, HWH, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1222424326
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, S7J, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1230130478
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, A1C9Q, MG |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z1230795916
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, A1BMO, MG |

