Search Count: 3,360
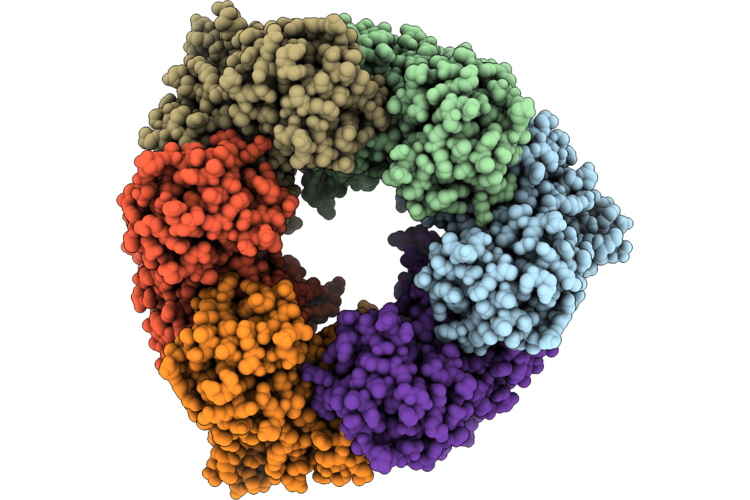 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 1
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
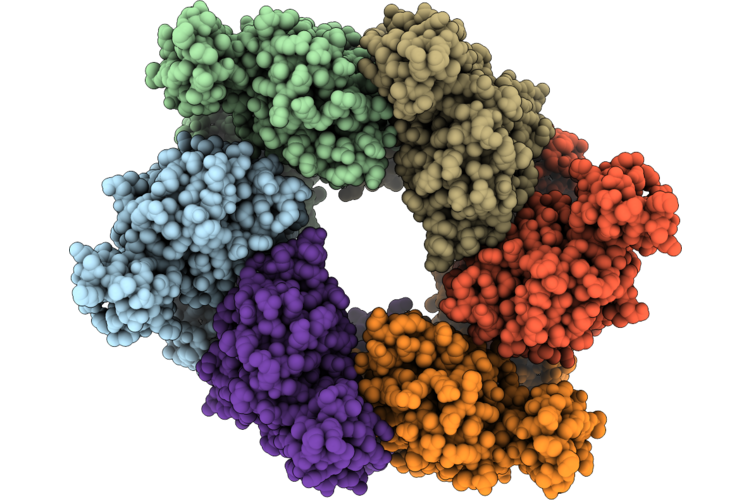 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 2
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
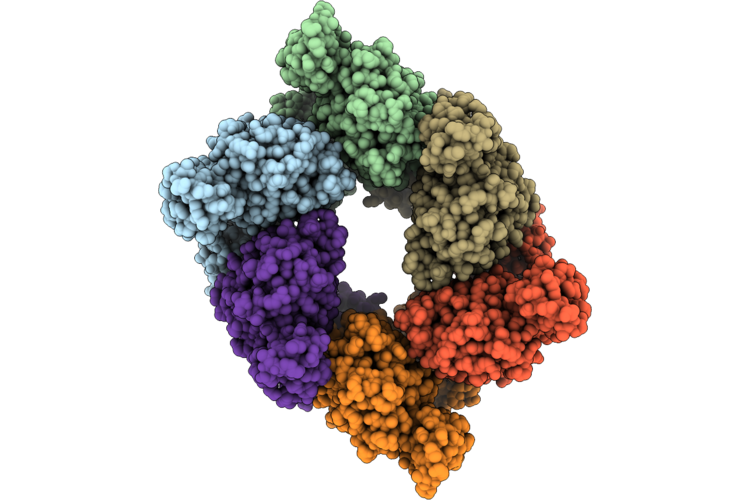 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 3
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
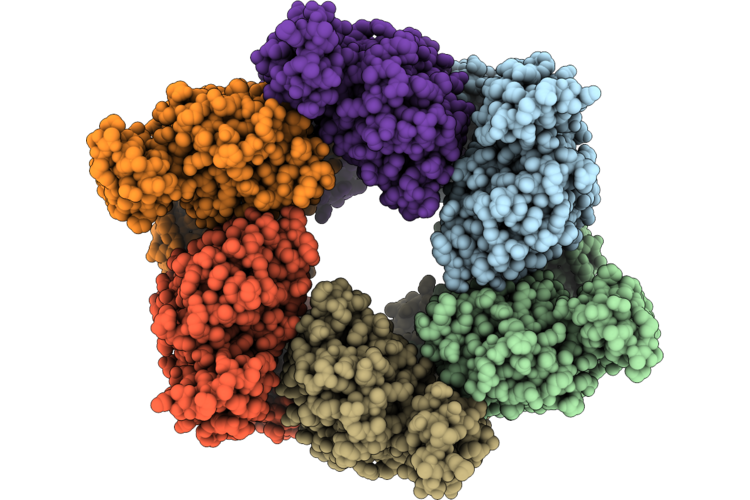 |
Single Particle Reconstruction Of Pilu From Vibrio Cholerae El Tor E7946, Form 4
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: MOTOR PROTEIN |
 |
Organism: Vibrio cholerae, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:3.64 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
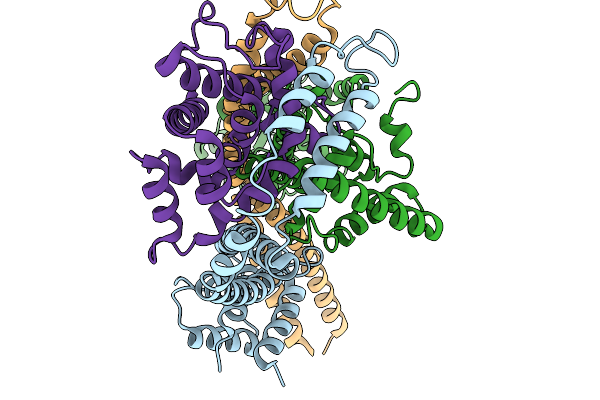 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:3.54 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
 |
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Resolution:4.07 Å Release Date: 2026-03-25 Classification: DNA BINDING PROTEIN |
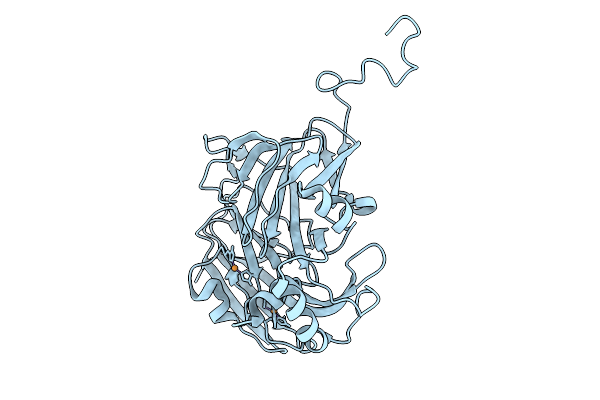 |
Atomic Resolution (1.05 A) Xfel Structure Of Chemically-Reduced Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.05 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU |
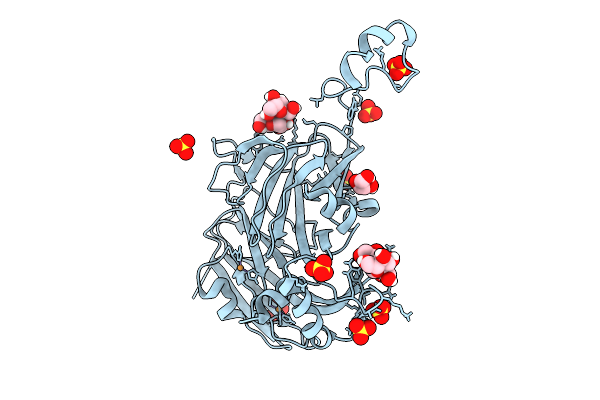 |
Atomic Resolution (1.15 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At Low Ph (5.5) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4, GOL |
 |
Joint X-Ray/Neutron Structure Of Wild-Type Bacillus Halodurans Rnase H1 In The Apo-Form
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.48 Å, 1.90 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: SO4, DOD |
 |
Joint X-Ray/Neutron Structure Of D132N Bacillus Halodurans Rnase H1 In The Apo-Form
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.4 Å, 2.20 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: SO4, DOD |
 |
Room-Temperature X-Ray Structure Of D132N Bacillus Halodurans Rnase H1 In Complex With Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MG, PO3 |
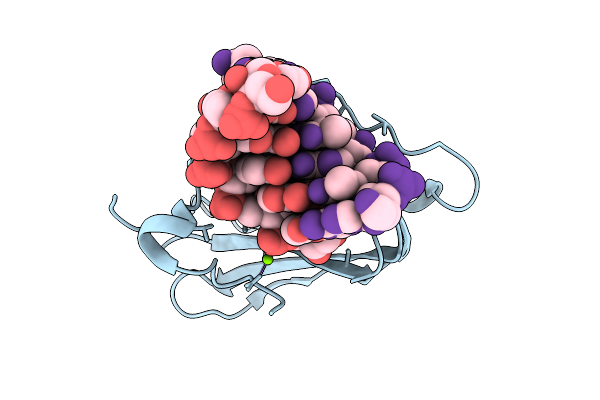 |
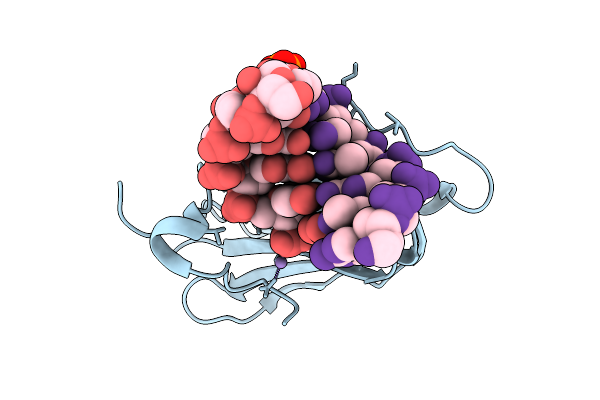
Room-Temperature X-Ray Structure Of D132N Bacillus Halodurans Rnase H1 In Complex With Complementary Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MG |
 |
100K X-Ray Structure Of Mixed Metal D132N Bacillus Halodurans Rnase H1 Complex With Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MN, MG, PO3 |
 |
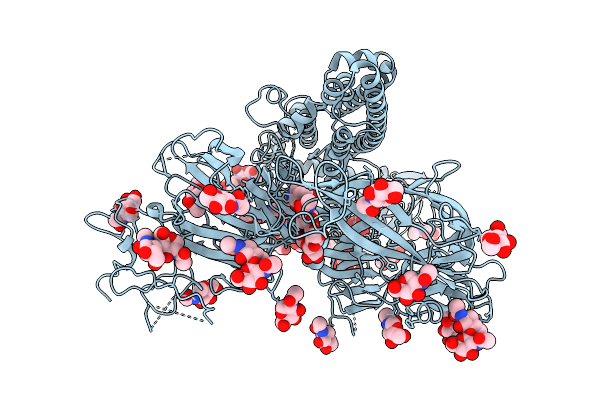
Cryo-Em Structure Of The Escherichia Phage Hk446 Rip1 In Complex With The Enterobacteria Phage T6 Small Terminase
Organism: Enterobacteria phage t6, Escherichia phage hk446
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: VIRAL PROTEIN |
 |
Organism: Feline infectious peritonitis virus (strain 79-1146), Enterobacteria phage t6
Method: ELECTRON MICROSCOPY Resolution:2.78 Å Release Date: 2026-01-28 Classification: PROTEIN BINDING Ligands: NAG |
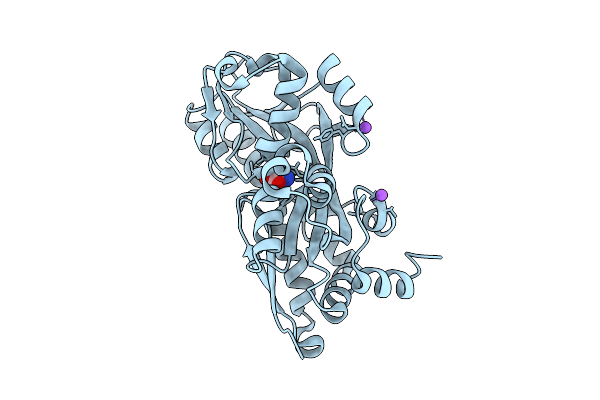 |
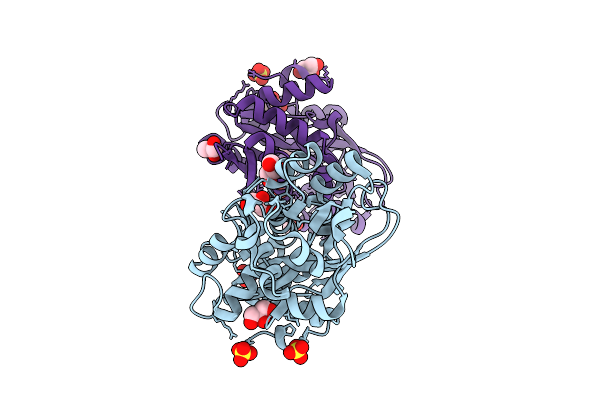
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2025-12-24 Classification: TRANSPORT PROTEIN |
 |
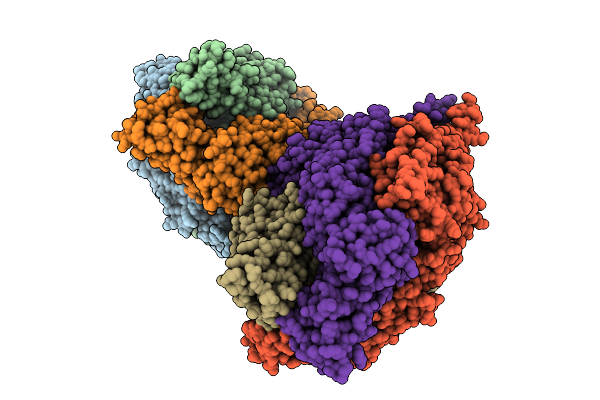
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: TRANSPORT PROTEIN Ligands: GOL, SO4 |
 |
Organism: Human respiratory syncytial virus, Enterobacteria phage t6
Method: X-RAY DIFFRACTION Resolution:2.92 Å Release Date: 2025-12-17 Classification: VIRAL PROTEIN |

