Search Count: 3,272
All
Selected
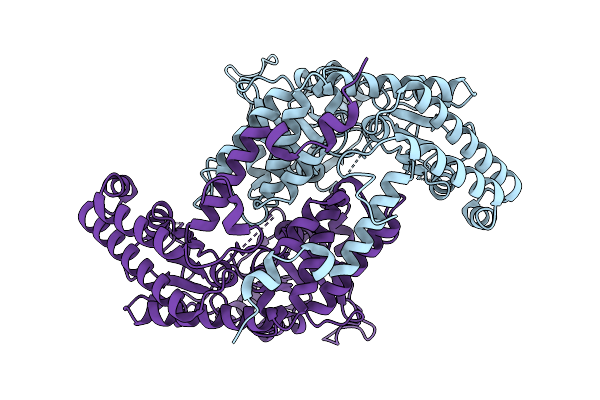 |
Cryo-Em Structure Of The A149T Dimer Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Essex In Complex With Plp
Organism: Glycine max
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE |
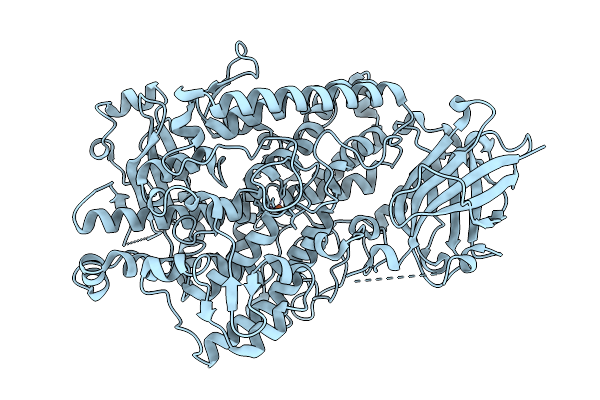 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: FE |
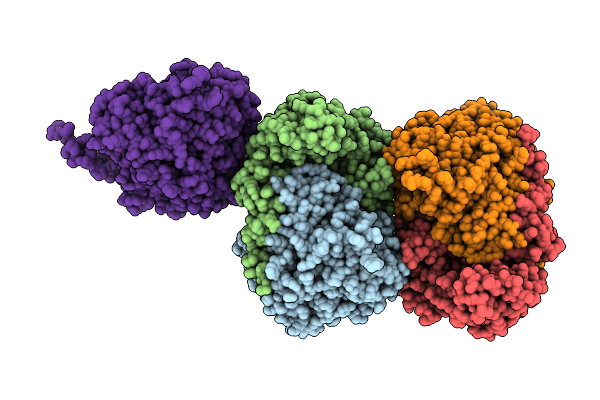 |
Crystal Structure Of A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Essex In Complex With Plp-Glycine
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PLG |
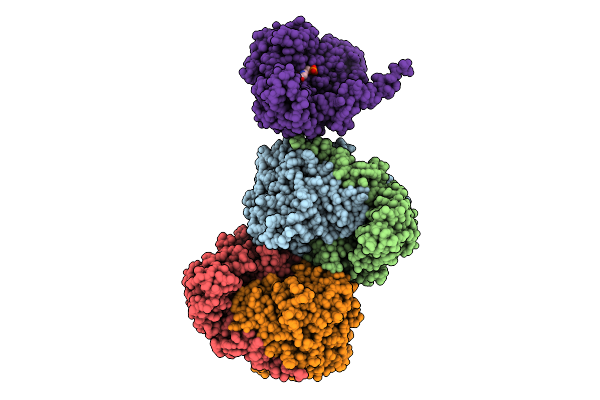 |
Crystal Structure Of The A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Forrest In Complex With Plp-Glycine
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PLG |
 |
Crystal Structure Of The A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Forrest In Complex With Plp
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Crystal Structure Of The A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Essex In Complex With Plp
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, PO4 |
 |
Organism: Micrococcus flavus
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN Ligands: GOL |
 |
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Alanylated Rna Microhelices Analogues Mimicking Ala-Trna-Ala Substrate
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.61 Å Release Date: 2026-03-11 Classification: RNA BINDING PROTEIN |
 |
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: MG, TRT, A1BYH |
 |
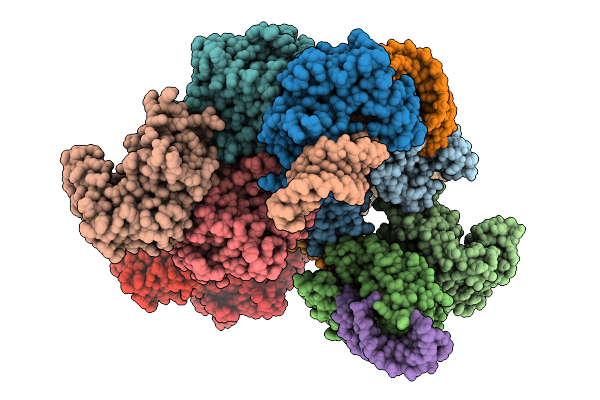
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Ala Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.29 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
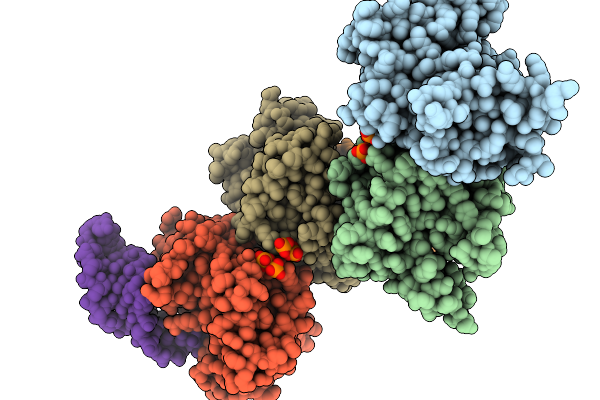
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Glu Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.52 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
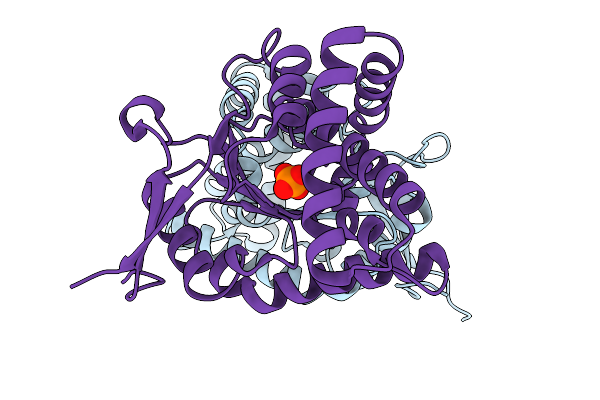
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Glu Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.43 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
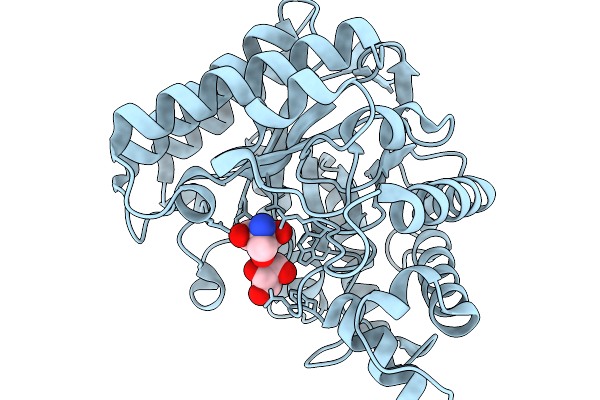
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps)
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-18 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: TRS, GOL |
 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2026-02-04 Classification: HYDROLASE |
 |
Organism: Enterococcus faecium do
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-01-28 Classification: CELL ADHESION Ligands: EDO, SO4 |
 |
Organism: Enterococcus faecium do
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-01-28 Classification: CELL ADHESION |
 |
Crystal Structure Of Nodd-Ebd (Effector Binding Domain) In Complex With Hesperetin From Rhizobium Leguminosarum Bv. Vicae 3841
Organism: Rhizobium leguminosarum
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-12-24 Classification: TRANSCRIPTION Ligands: GOL, 6JP |

