Search Count: 7,473
All
Selected
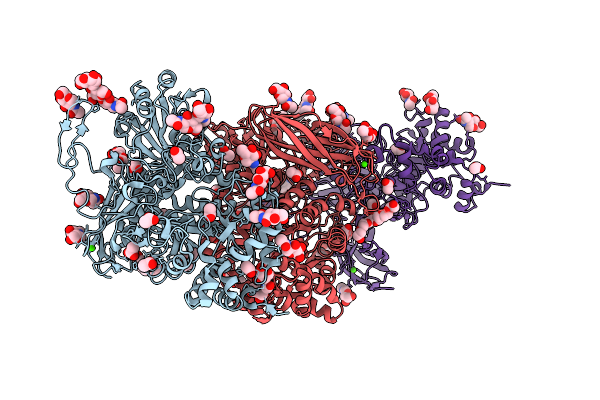 |
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
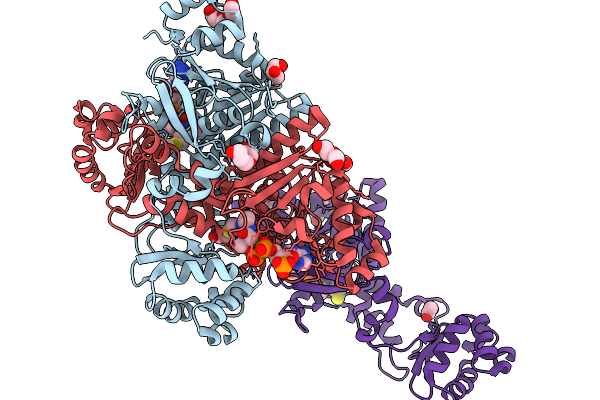 |
Crystal Structure Of Formyl-Coenzyme A Transferase From Brucella Melitensis In Complex With Coa
Organism: Brucella melitensis 16m1w
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: PGE, GOL, ACT, COA, PEG |
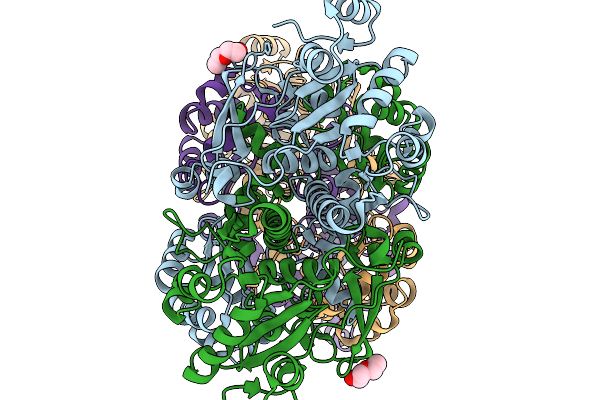 |
Crystal Structure Of Formyl-Coenzyme A Transferase From Brucella Melitensis In Complex With Zinc
Organism: Brucella melitensis 16m1w
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: PEG, ZN |
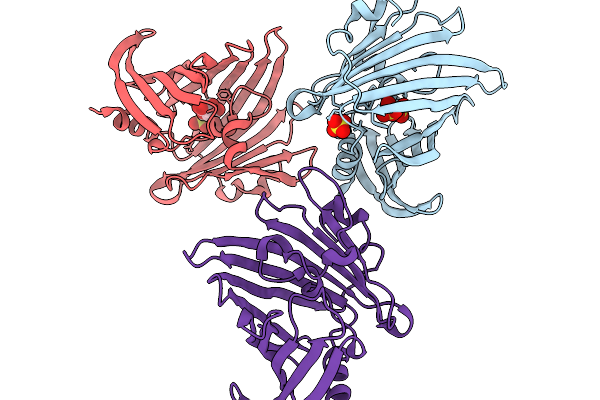 |
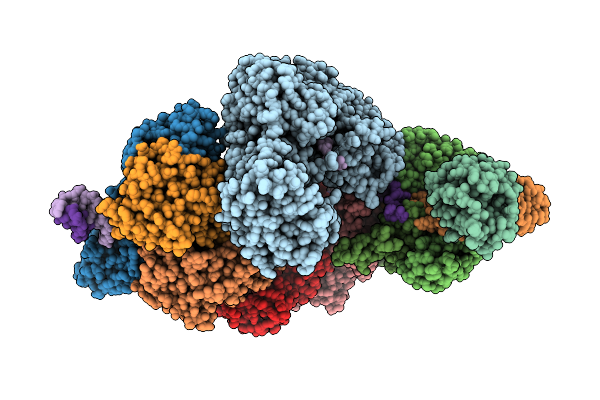
Organism: Porphyromonas gingivalis atcc 33277
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2026-02-18 Classification: TRANSPORT PROTEIN Ligands: SO4, TLA |
 |
Organism: Shewanella putrefaciens cn-32
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: ANTIVIRAL PROTEIN Ligands: MG |
 |
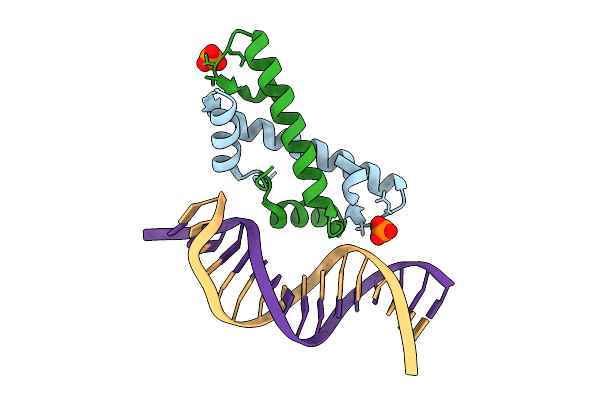
Organism: Leptospira perolatii
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN Ligands: SO4 |
 |
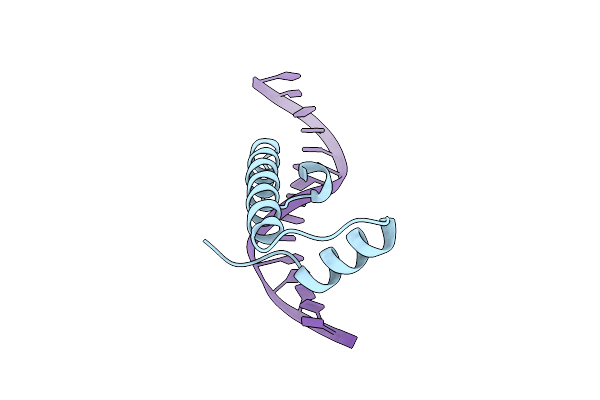
Organism: Leptospira perolatii, Dna molecule
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN Ligands: PO4 |
 |
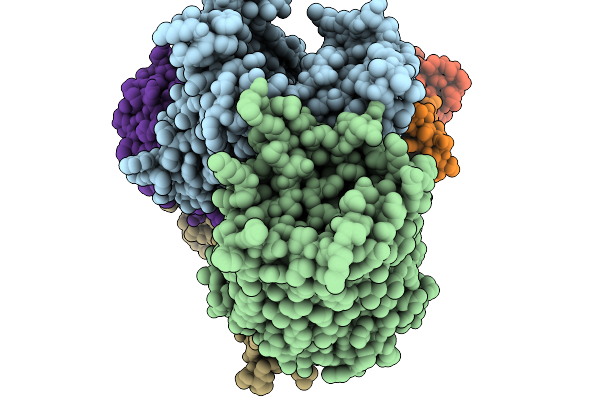
Organism: Leptospira perolatii, Dna molecule
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-24 Classification: DNA BINDING PROTEIN |
 |
Crystal Structure Of Formyl-Coenzyme A Transferase From Brucella Melitensis In Complex With Succinate
Organism: Brucella melitensis 16m1w
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2025-12-03 Classification: TRANSFERASE Ligands: SIN |
 |
Organism: Porphyromonas gingivalis atcc 33277
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: MEMBRANE PROTEIN Ligands: PLM, Z41 |
 |
Organism: Magnetospirillum gryphiswaldense msr-1
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: METAL BINDING PROTEIN Ligands: HEM, FE |
 |
Organism: Shewanella putrefaciens cn-32
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2025-07-30 Classification: HYDROLASE Ligands: MG |
 |
Organism: Streptantibioticus cattleyicolor (strain atcc 35852 / dsm 46488 / jcm 4925 / nbrc 14057 / nrrl 8057)
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Streptantibioticus cattleyicolor (strain atcc 35852 / dsm 46488 / jcm 4925 / nbrc 14057 / nrrl 8057)
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: A1D90, SO4 |
 |
Organism: Streptantibioticus cattleyicolor (strain atcc 35852 / dsm 46488 / jcm 4925 / nbrc 14057 / nrrl 8057)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: A1D91 |
 |
Organism: Marchantia polymorpha, Phytophthora infestans
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-05-21 Classification: PEPTIDE BINDING PROTEIN |
 |
Organism: Shewanella putrefaciens cn-32
Method: X-RAY DIFFRACTION Resolution:3.01 Å Release Date: 2025-03-26 Classification: SIGNALING PROTEIN |
 |
Organism: Cutibacterium acnes sk137
Method: SOLUTION NMR Release Date: 2025-03-05 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Deinococcus geothermalis (strain dsm 11300 / cip 105573 / ag-3a)
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2025-01-22 Classification: CARBOHYDRATE |
 |
Cryo-Em Structure And Rational Engineering Of A Novel Efficient Ochratoxin A-Detoxifying Amidohydrolase
Organism: Pseudoxanthomonas wuyuanensis
Method: ELECTRON MICROSCOPY Resolution:2.37 Å Release Date: 2024-12-18 Classification: HYDROLASE Ligands: ZN, 97U |

