Search Count: 1,546
All
Selected
 |
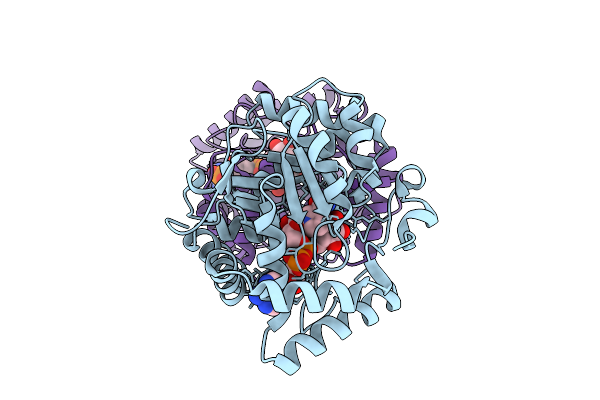
Crystal Structure Of L-Galactose Dehydrogenase From Luteolibacter Sp. Strain Lg18 In Complex With L-Galactose And Nadp+
Organism: Luteolibacter sp. lg18
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GIV, NAP |
 |
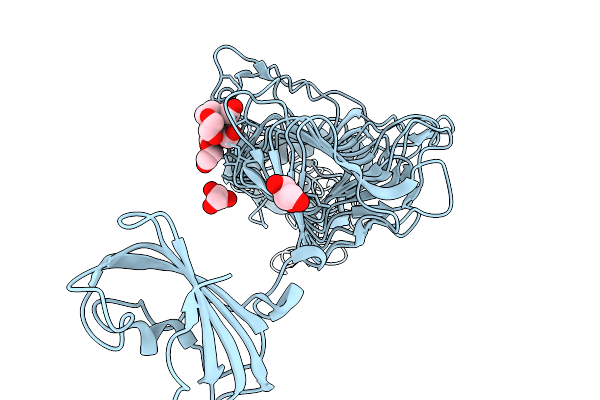
Crystal Structure Of L-Galactose Dehydrogenase From Luteolibacter Sp. Strain Lg18 In Complex With L-Glucose And Nadp+
Organism: Luteolibacter sp. lg18
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: Z8T, NAP |
 |
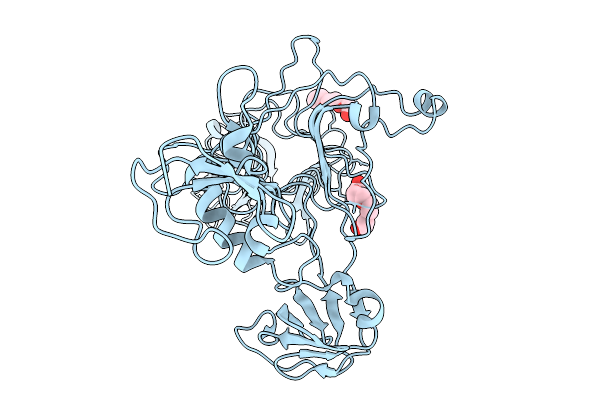
Crystal Structure Of A Tailspike Depolymerase (Belisarius_Gp86) From Acinetobacter Phage Belisarius
Organism: Acinetobacter phage
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: GOL, PGE |
 |
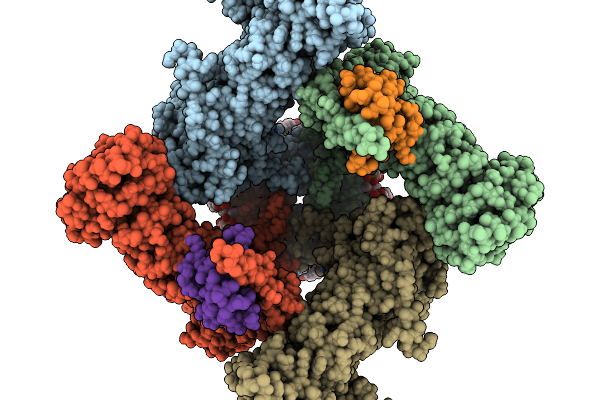
Crystal Structure Of A Tailspike Depolymerase (Solidus_Gp83) From Acinetobacter Phage Solidus
Organism: Acinetobacter phage
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: PG4, 1PE |
 |
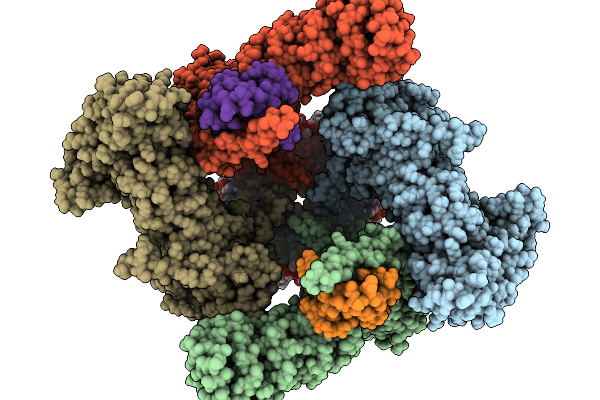
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, CA, D39 |
 |
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, NCA, CA, D39 |
 |
Structure Of Nanchung-Inactive-Calmodulin In Complex With Nicotinamide, Edta
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, NCA, D39 |
 |
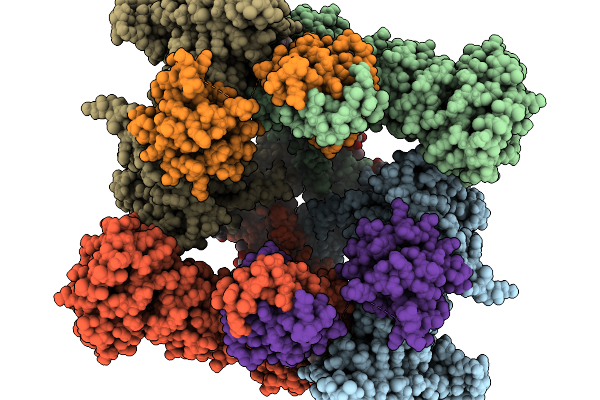
Structure Of Nanchung-Inactive-Calmodulin In Complex With Afidopyropen And Calcium
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, A1B32, CA, D39 |
 |
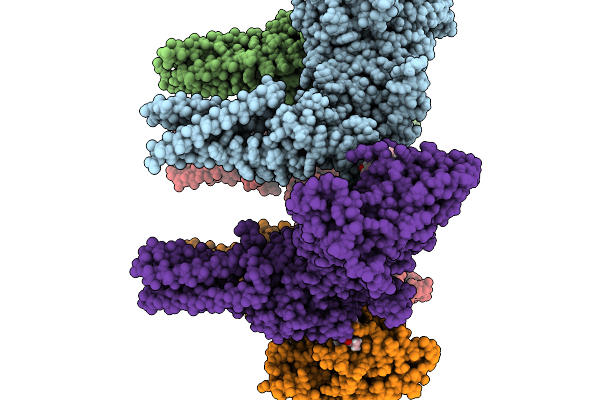
Structure Of Nanchung-Inactive-Calmodulin In Complex With Afidopyropen, Edta
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, A1B32, D39 |
 |
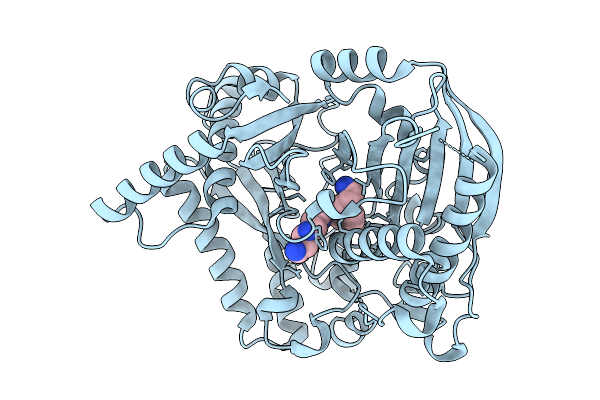
Organism: Halyomorpha halys
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: A1B32 |
 |
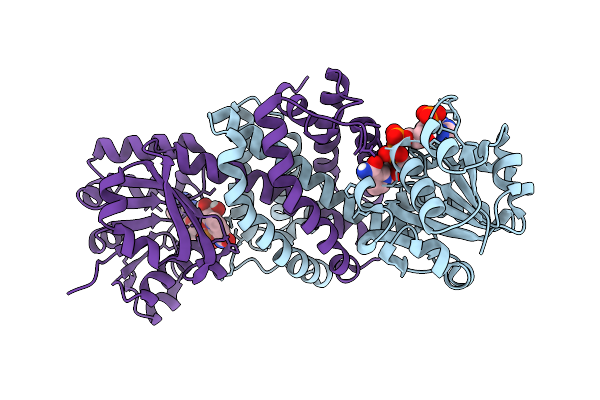
Organism: Lycium barbarum
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: SPD |
 |
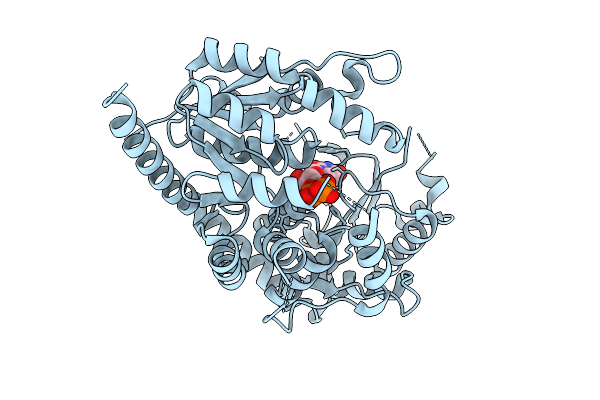
Organism: Streptomyces aurantiacus ja 4570
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2025-08-13 Classification: OXIDOREDUCTASE Ligands: NAP |
 |
Organism: Platycodon grandiflorus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-07-09 Classification: TRANSFERASE Ligands: UDP |
 |
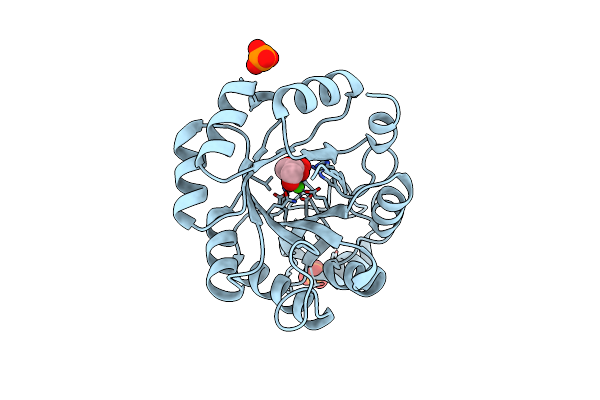
Organism: Pseudomonas
Method: X-RAY DIFFRACTION Resolution:0.99 Å Release Date: 2025-06-11 Classification: OXIDOREDUCTASE Ligands: HEM, IMD, NA, CL |
 |
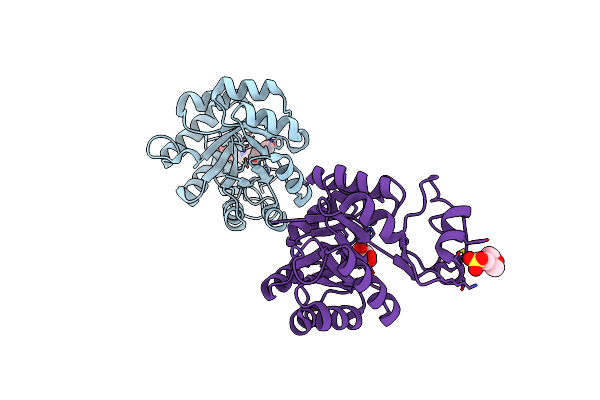
Structure Of The Bacteriophage K Gp155 C-Terminal Domain, Seleno-Methionine Modified Version
Organism: Staphylococcus phage k
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2025-05-14 Classification: VIRAL PROTEIN Ligands: G3P, CA, PO4 |
 |
Organism: Staphylococcus phage k
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2025-04-09 Classification: VIRAL PROTEIN Ligands: MES, GOL |
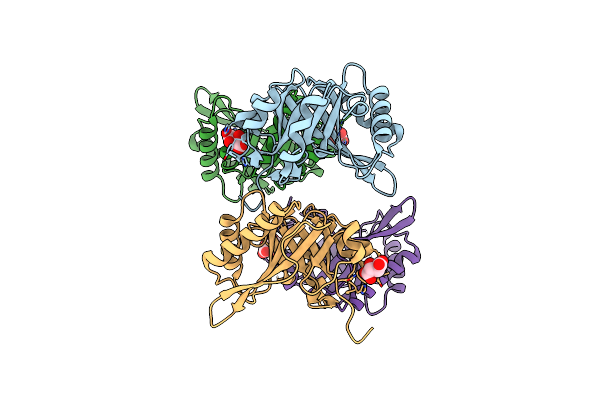 |
Organism: Yersinia pseudotuberculosis b-6862
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-02-05 Classification: HYDROLASE Ligands: TLA |
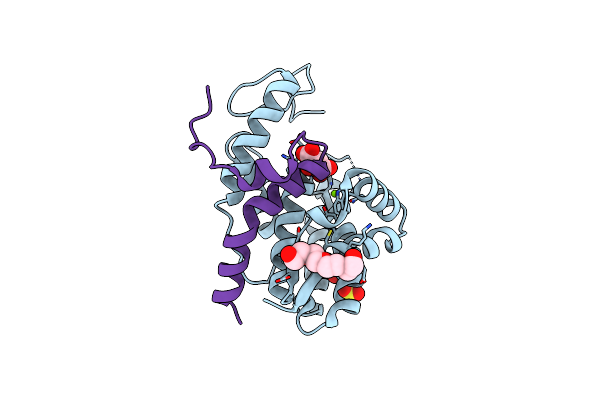 |
The Toxin-Antitoxin Complex Fic-1-Antf Is A Deampylase That Regulates The Activity Of Dna Gyrase
Organism: Pseudomonas, Pseudomonas bijieensis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-10-23 Classification: STRUCTURAL PROTEIN Ligands: PG4, TAR, MG, SO4 |
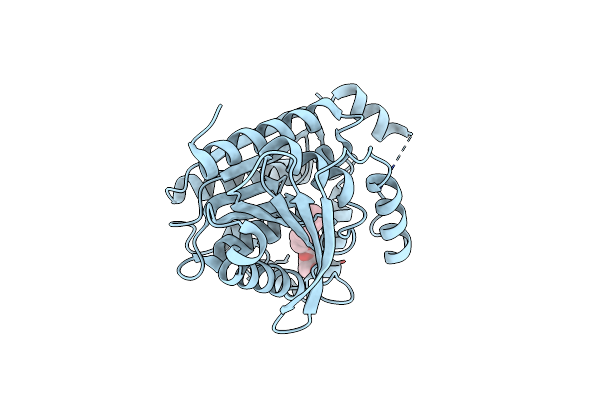 |
Organism: Saccharolobus shibatae
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-07-10 Classification: HYDROLASE Ligands: BOG |
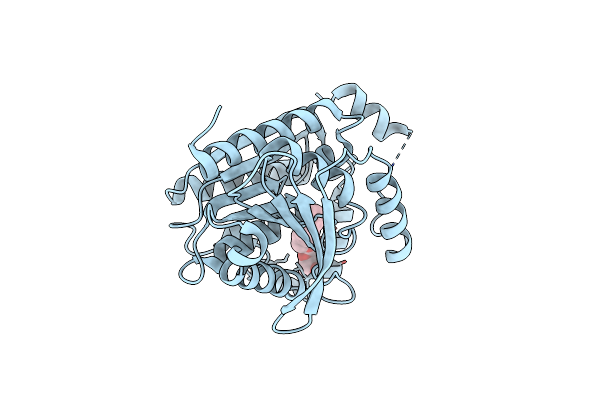 |
Organism: Saccharolobus shibatae
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2024-07-10 Classification: HYDROLASE Ligands: BOG |

