Search Count: 2,318
All
Selected
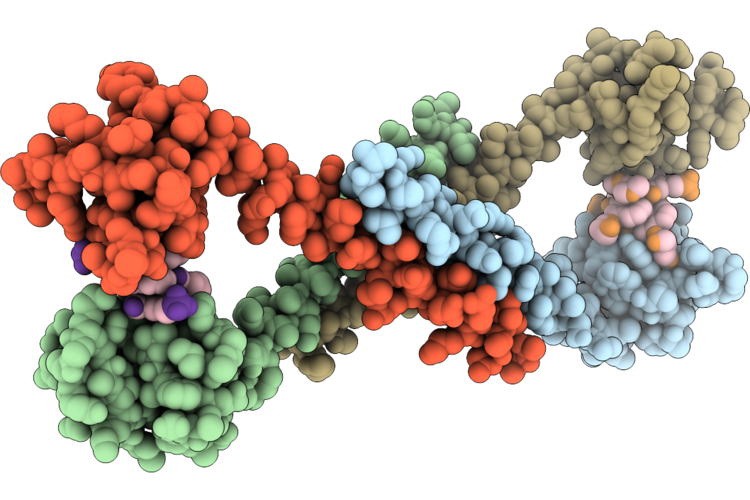 |
Crystal Structure Of Affitin C10 Fused To A Coiled-Coil Domain In Complex With A Quinoline Oligoamide Foldamer
Organism: Sulfolobus acidocaldarius, Mus musculus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-22 Classification: DE NOVO PROTEIN |
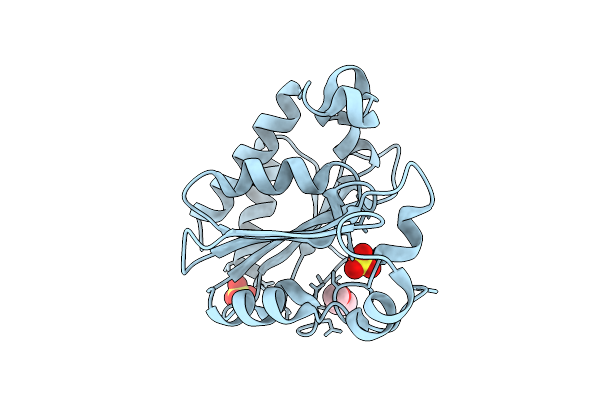 |
Crystal Structure Of Poly-Gamma-Glutamate Hydrolase From Bacillus Phage Pm1
Organism: Bacillus phage pm1
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: PEG, SO4 |
 |
Crystal Structure Of Poly-Gamma-Glutamate Hydrolase From Bacillus Phage Pm1 In Complex With A Zinc Ion
Organism: Bacillus phage pm1
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: PEG, ZN |
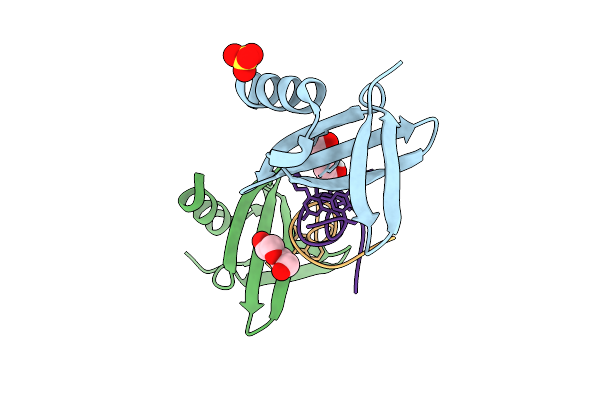 |
Crystal Structure Of Nanofitin C10 In Complex With A A Double-Helical Aromatic Oligoamide Foldamer
Organism: Sulfolobus acidocaldarius, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-08 Classification: DE NOVO PROTEIN Ligands: SO4, PEG, CL |
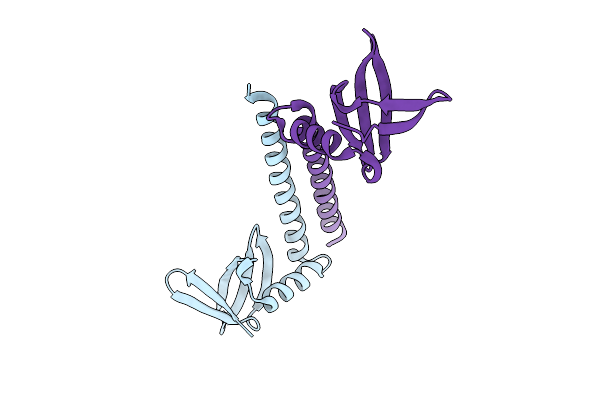 |
Crystal Structure Of The Aromatic Oligoamide Foldamer Binder Nanofitin C10 Fused To A Coiled-Coil Domain
Organism: Sulfolobus acidocaldarius
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-03-18 Classification: DE NOVO PROTEIN |
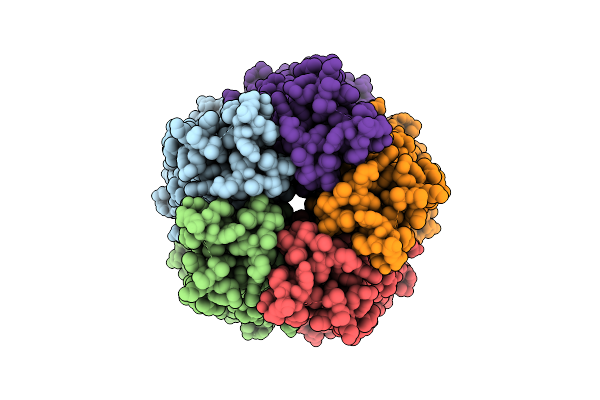 |
Organism: Sarcoptes scabiei
Method: ELECTRON MICROSCOPY Resolution:4.24 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
 |
Organism: Sarcoptes scabiei
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: IVM |
 |
Organism: Sarcoptes scabiei
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: CL |
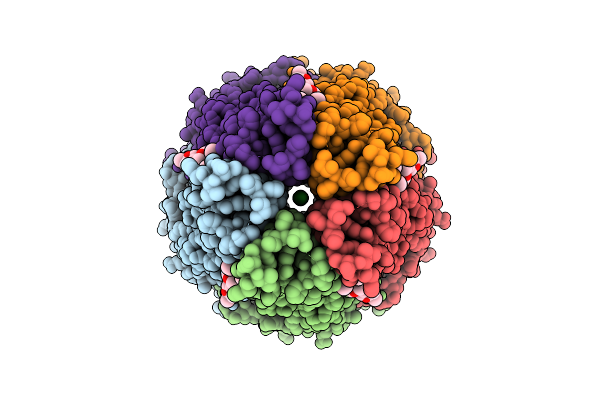 |
Organism: Sarcoptes scabiei
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: IVM, CL |
 |
Crystal Structure Of Retro-Aldolase Ra95.5-8 Mutant With A Covalently Bound Cyclohexenone Derivative
Organism: Saccharolobus solfataricus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: SO4, EDO |
 |
Crystal Structure Of Retro-Aldolase T53L/K210H Ra95.5-8 With A Covalently Bound Cyclohexenone Derivative
Organism: Saccharolobus solfataricus
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: SO4, EDO |
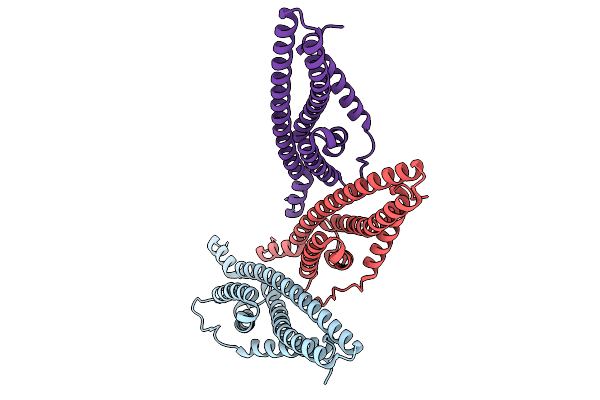 |
Organism: Sulfolobus acidocaldarius
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
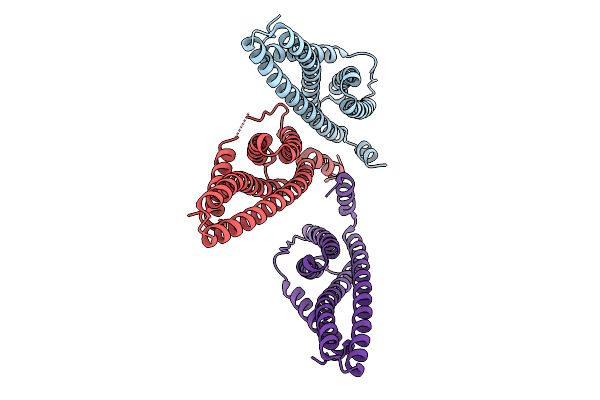 |
Organism: Sulfolobus acidocaldarius
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
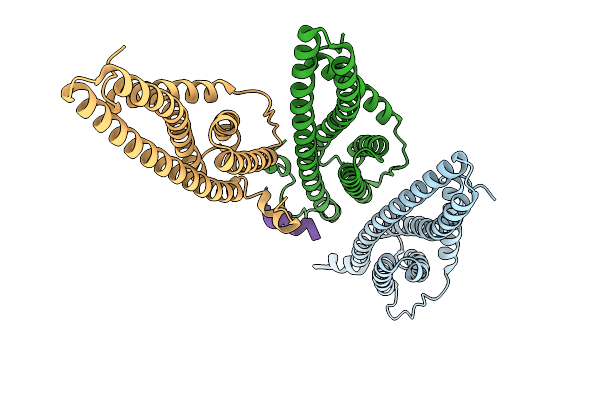 |
Organism: Sulfolobus acidocaldarius
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: CELL CYCLE |
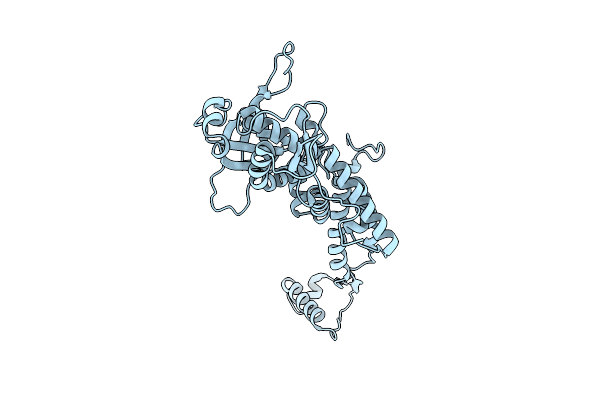 |
Organism: Shigella phage sf11 smd-2017
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2025-11-12 Classification: VIRUS |
 |
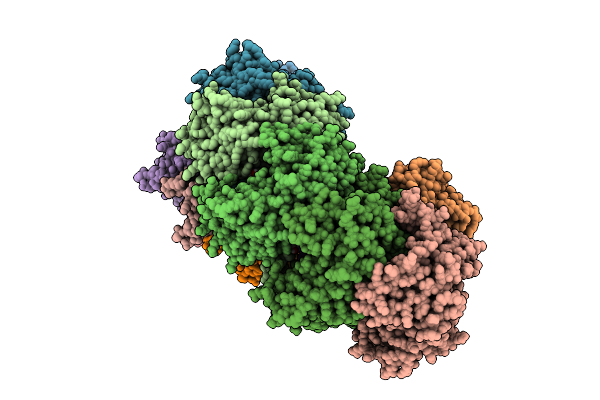
Organism: Trichoplax adhaerens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: HYDROLASE |
 |
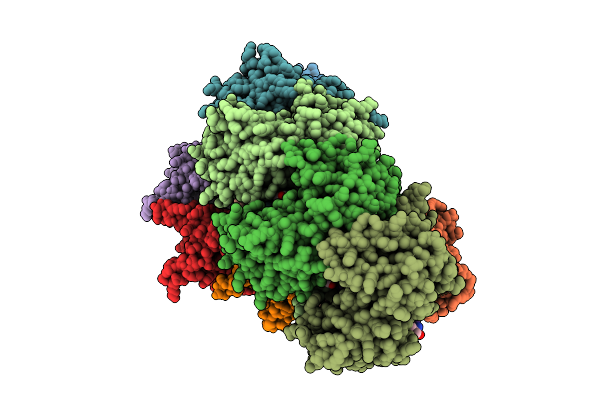
Organism: Saccharolobus solfataricus p2, Saccharolobus solfataricus
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: RNA BINDING PROTEIN/RNA Ligands: TRS |
 |
Organism: Saccharolobus solfataricus, Saccharolobus solfataricus p2
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: TRS |
 |
Organism: Saccharolobus solfataricus, Saccharolobus solfataricus p2
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: RNA BINDING PROTEIN/RNA Ligands: TRS |
 |
Organism: Flock house virus
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-08-13 Classification: VIRUS |

