Search Count: 648
All
Selected
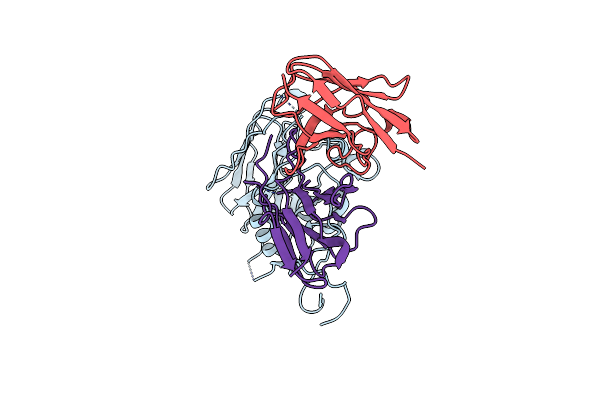 |
Structure Of Ut14 Fab In Complex With The Head Domain Of H3 (A/Singapore/Infimh-16-0019/2016)
Organism: Homo sapiens, Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2024-11-27 Classification: IMMUNE SYSTEM |
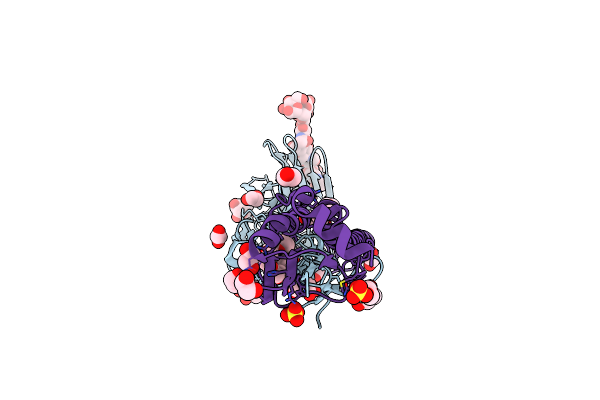 |
Crystal Structure Of The A/Ecuador/1374/2016(H3N2) Influenza Virus Hemagglutinin With Human Receptor Analog 6'-Slnln
Organism: Influenza a virus
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2024-02-14 Classification: VIRAL PROTEIN Ligands: NAG, PG4, EDO, PEG, SO4 |
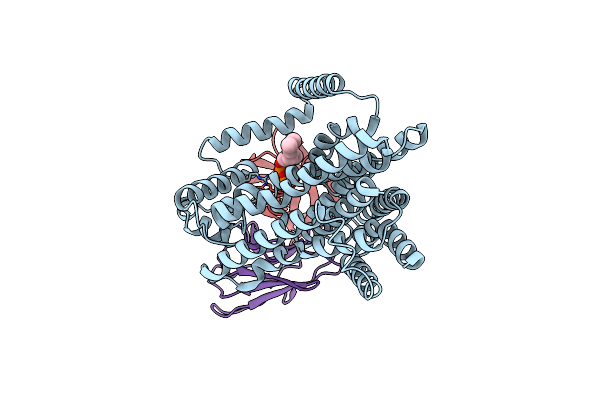 |
Single-Particle Cryo-Em Structure Of The Waal O-Antigen Ligase In Its Ligand Bound State
Organism: Cupriavidus metallidurans, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2022-04-06 Classification: MEMBRANE PROTEIN Ligands: GPP |
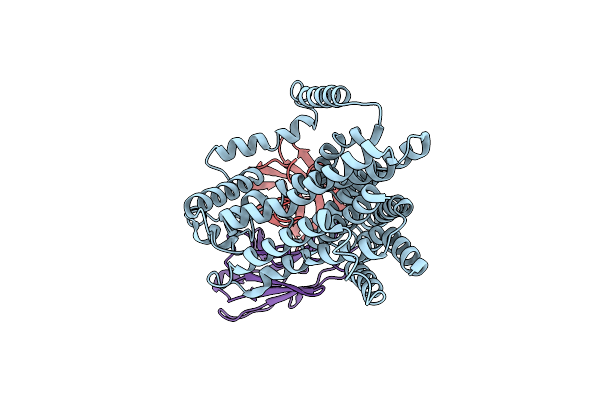 |
Single-Particle Cryo-Em Structure Of The Waal O-Antigen Ligase In Its Apo State
Organism: Cupriavidus metallidurans, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2022-04-06 Classification: MEMBRANE PROTEIN |
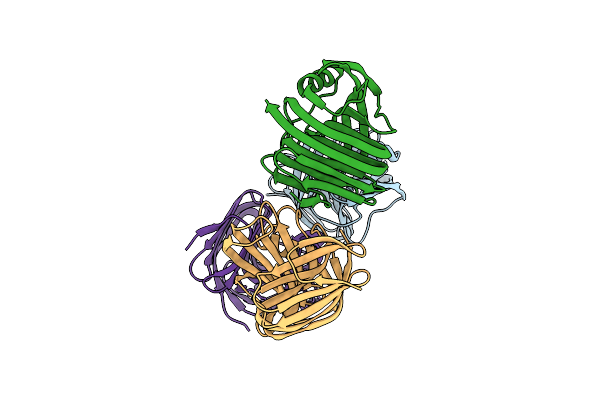 |
Crystal Structure Of Glycoside Hydrolase Family 11 Beta-Xylanase From Streptomyces Olivaceoviridis E-86
Organism: Streptomyces olivaceoviridis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2020-12-30 Classification: HYDROLASE Ligands: CL |
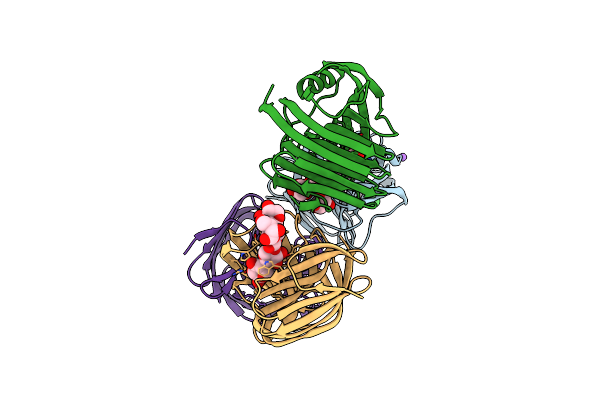 |
Crystal Structure Of Glycoside Hydrolase Family 11 Beta-Xylanase From Streptomyces Olivaceoviridis E-86 In Complex With Alpha-L-Arabinofuranosyl Xylotetraose
Organism: Streptomyces olivaceoviridis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2020-12-30 Classification: HYDROLASE Ligands: CL, NA |
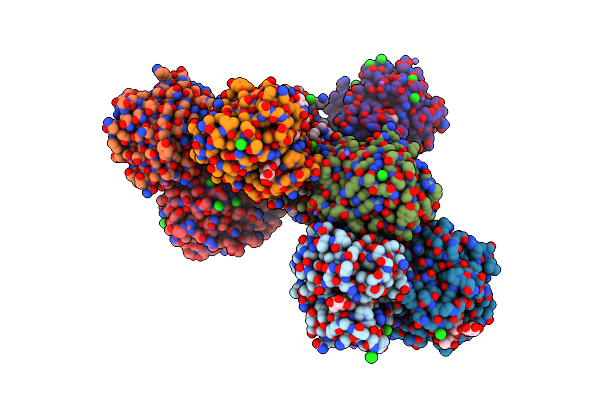 |
Crystal Structure Of Glycoside Hydrolase Family 11 Beta-Xylanase From Streptomyces Olivaceoviridis E-86 In Complex With 4-O-Methyl-Alpha-D-Glucuronopyranosyl Xylotetraose
Organism: Streptomyces olivaceoviridis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2020-12-30 Classification: HYDROLASE Ligands: CL, NA |
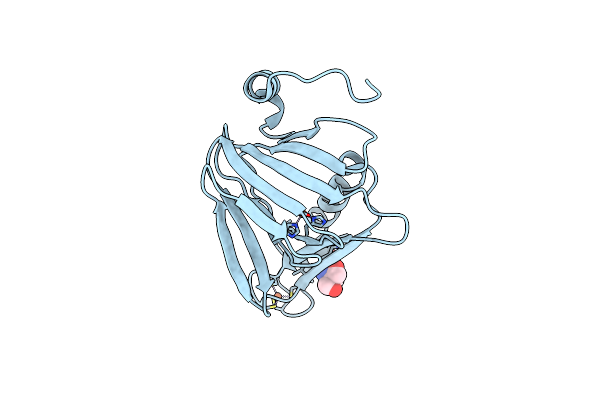 |
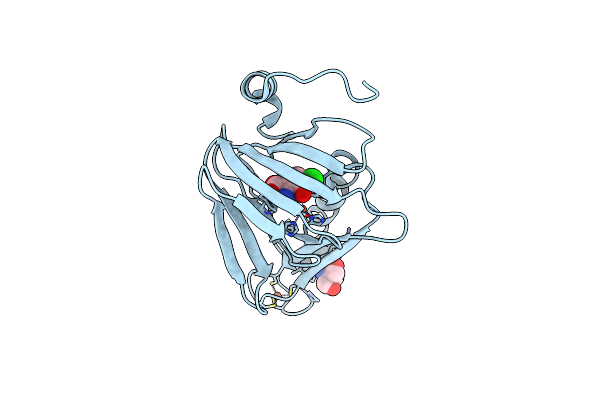
Crystal Structure Of 3-Hydroxyanthranilate-3,4-Dioxygenase N27A From Cupriavidus Metallidurans
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2018-06-06 Classification: OXIDOREDUCTASE Ligands: FE2, TRS |
 |
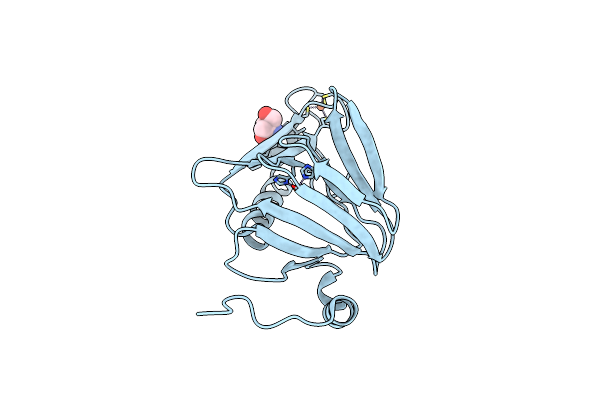
Crystal Structure Of 3-Hydroxyanthranilate-3,4-Dioxygenase N27A From Cupriavidus Metallidurans In Complex With 4-Cl-3-Haa
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2018-06-06 Classification: OXIDOREDUCTASE Ligands: FE2, 4AA, TRS |
 |
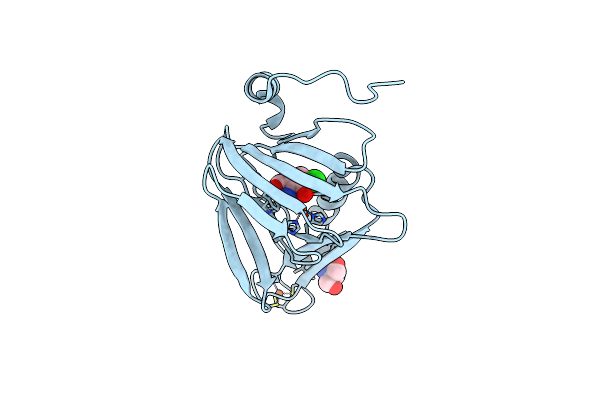
Crystal Structure Of 3-Hydroxyanthranilate-3,4-Dioxygenase I142A From Cupriavidus Metallidurans
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2018-06-06 Classification: OXIDOREDUCTASE Ligands: TRS, FE2 |
 |
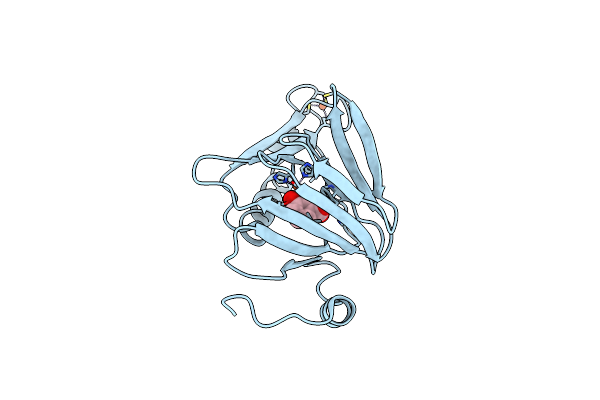
Crystal Structure Of 3-Hydroxyanthranilate-3,4-Dioxygenase I142A From Cupriavidus Metallidurans In Complex With 4-Cl-3-Haa
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2018-06-06 Classification: OXIDOREDUCTASE Ligands: 4AA, TRS, FE2 |
 |
Crystal Structure Of 3-Hydroxyanthranilate-3,4-Dioxygenase I142A From Cupriavidus Metallidurans In Complex With 3-Haa
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2018-06-06 Classification: OXIDOREDUCTASE Ligands: FE2, 3HA |
 |
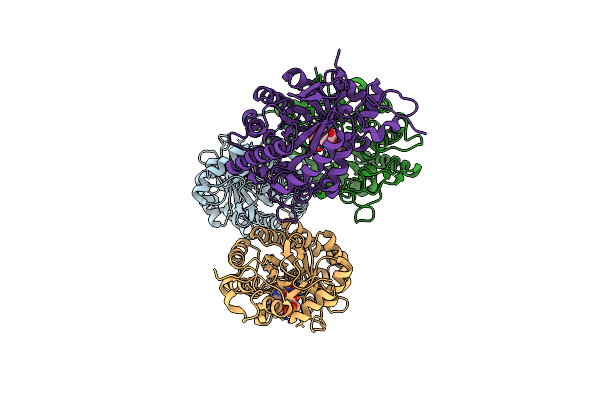
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2017-12-06 Classification: HYDROLASE Ligands: MLI |
 |
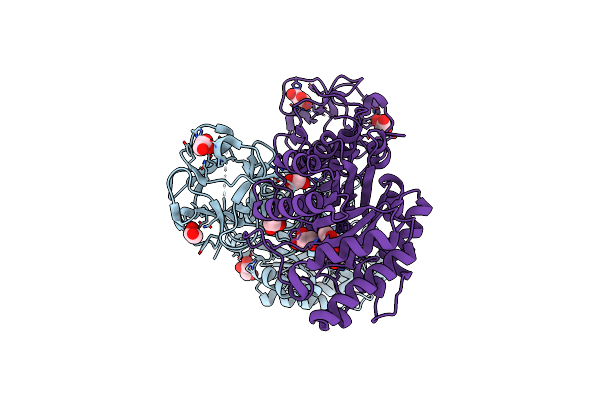
Organism: Cupriavidus metallidurans
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2017-09-06 Classification: FLAVOPROTEIN Ligands: FMN, ACT, CIT |
 |
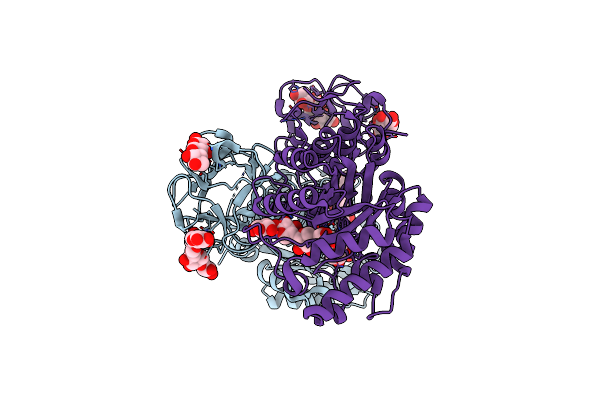
Crystal Structure Of Covalent Glycosyl-Enzyme Intermediate Of Xylanase Mutant (T82A, N127S, And E128H) From Streptomyces Olivaceoviridis E-86
Organism: Streptomyces olivaceoviridis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2017-08-09 Classification: HYDROLASE Ligands: GOL |
 |
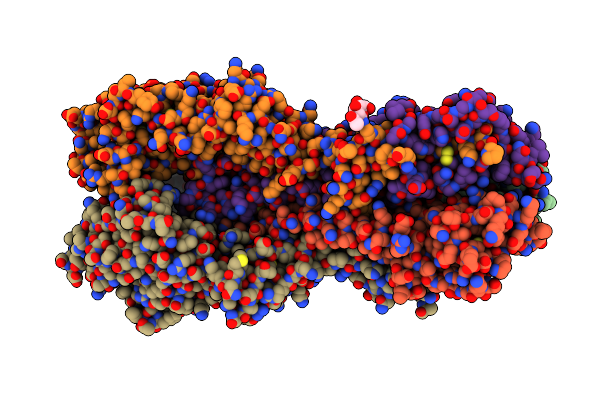
Crystal Structure Of Michaelis Complex Of Xylanase Mutant (T82A, N127S, And E128H) From Streptomyces Olivaceoviridis E-86
Organism: Streptomyces olivaceoviridis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2017-08-09 Classification: HYDROLASE |
 |
Crystal Structure Of H6 Hemagglutinin G225D Mutant From Taiwan (2013) H6N1 Influenza Virus
Organism: H6n1 subtype
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2017-06-14 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of H6 Hemagglutinin G225D Mutant From Taiwan (2013) H6N1 Influenza Virus In Complex With 6'-Sln
Organism: H6n1 subtype
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2017-06-14 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of H6 Hemagglutinin G225D Mutant From Taiwan (2013) H6N1 Influenza Virus In Complex With 3'-Sln
Organism: H6n1 subtype
Method: X-RAY DIFFRACTION Resolution:2.86 Å Release Date: 2017-06-14 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of H6 Hemagglutinin G225D Mutant From Taiwan (2013) H6N1 Influenza Virus In Complex With Lsta
Organism: H6n1 subtype
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2017-06-14 Classification: IMMUNE SYSTEM Ligands: NAG |

