Search Count: 26,838
All
Selected
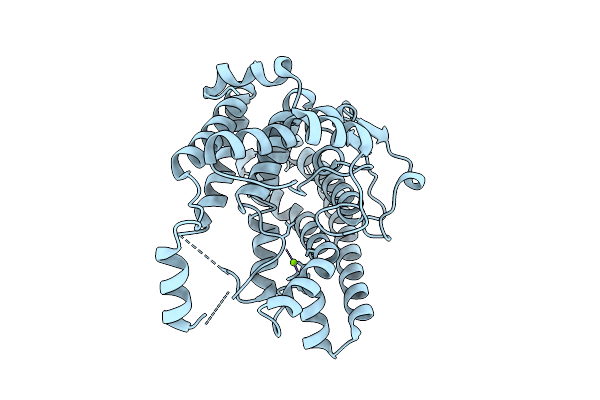 |
Crystal Structure Of The Tc Domain Of A Bifunctional Sesterterpene Synthase
Organism: Aspergillus stellatus
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-03-11 Classification: BIOSYNTHETIC PROTEIN Ligands: MG |
 |
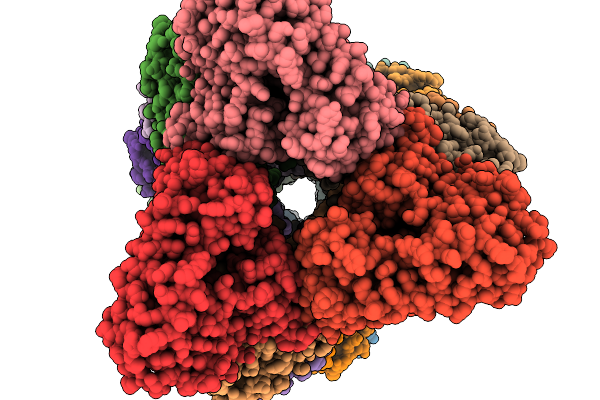
Organism: Aspergillus stellatus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
 |
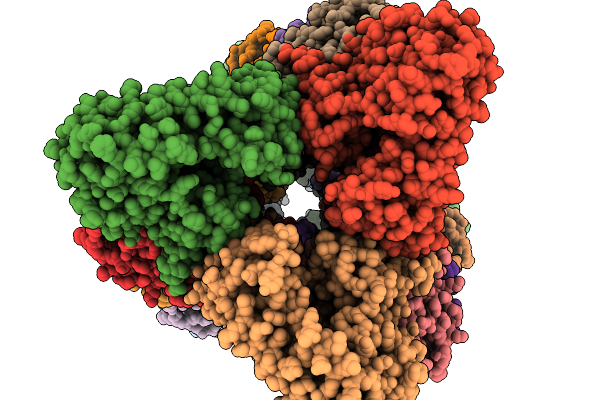
Organism: Aspergillus stellatus
Method: ELECTRON MICROSCOPY Resolution:3.19 Å Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
 |
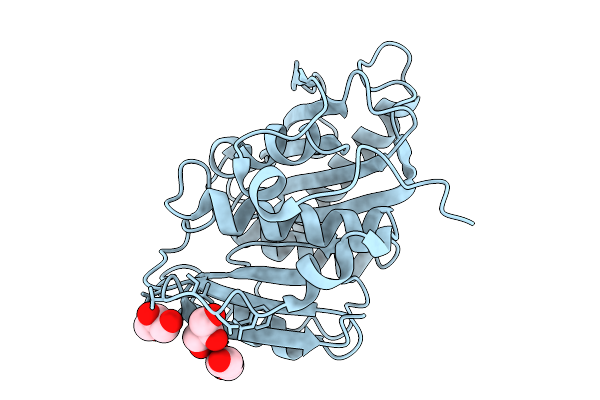
Organism: Aspergillus stellatus
Method: ELECTRON MICROSCOPY Resolution:3.25 Å Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: GOL |
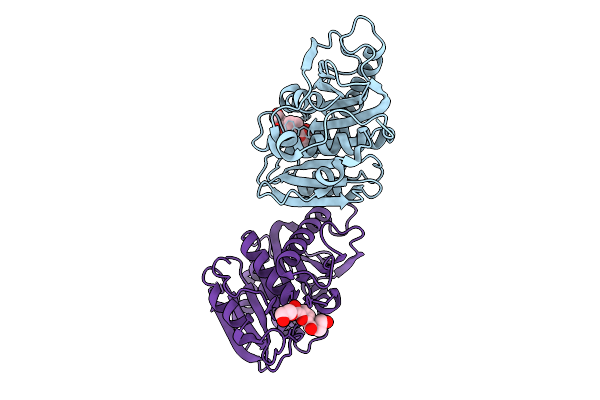 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: A1L8P |
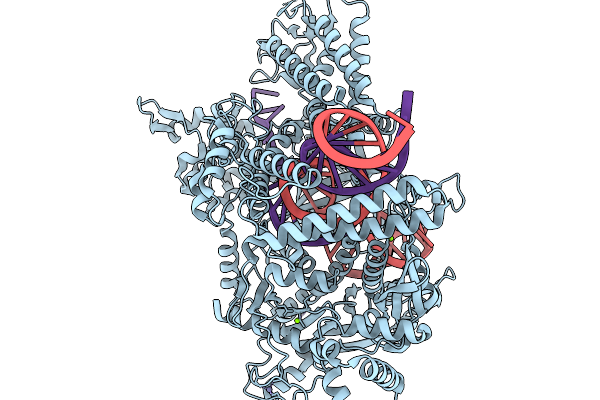 |
Organism: Bacteroidales, Bacteroidales bacterium
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: IMMUNE SYSTEM/RNA Ligands: MG, ZN |
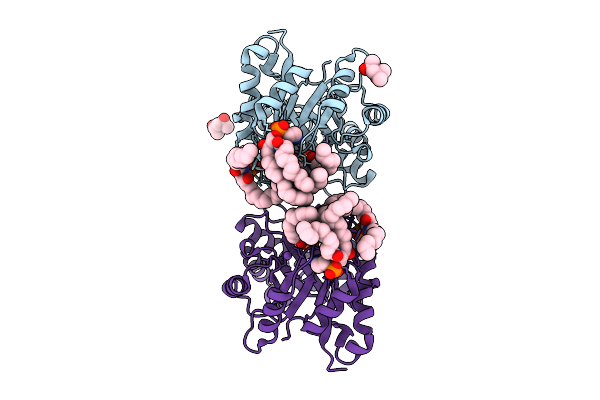 |
Structure Of Phospholipase D Betaib1I From Sicarius Terrosus Venom, H47N Mutant Bound To Product And Substrate Sphingolipids At 2.2 A Resolution From A 2-Day Old Crystal
Organism: Sicarius terrosus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-12-03 Classification: LYASE Ligands: A1A43, A1A44, NA, MG, MPD |
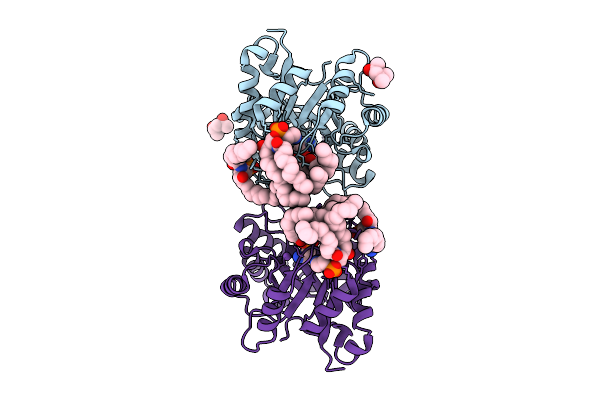 |
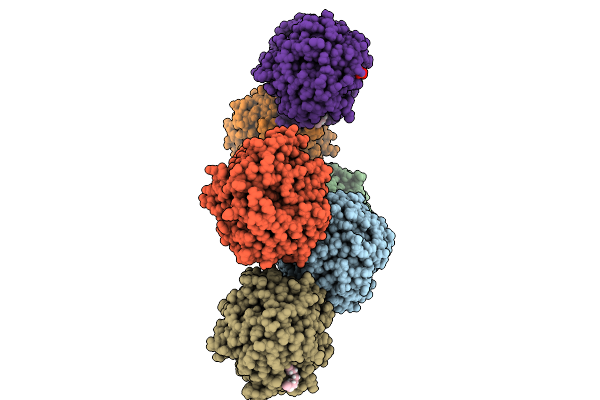
Structure Of Phospholipase D Betaib1I From Sicarius Terrosus Venom, H47N Mutant Bound To Substrate Sphingolipids At 2.60 A Resolution
Organism: Sicarius terrosus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-12-03 Classification: LYASE Ligands: A1A43, NA, MG, MPD |
 |
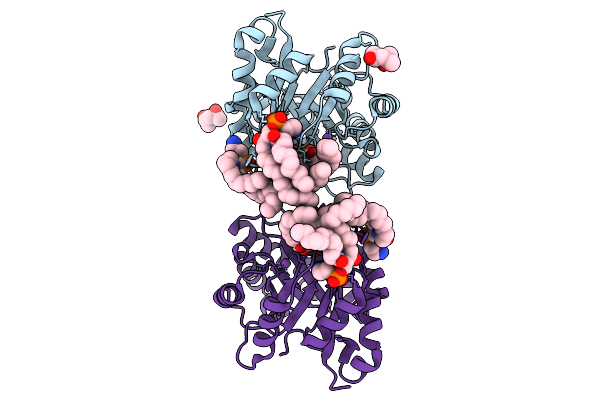
Organism: Mycolicibacterium
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: GOL, MG, 8K6, SO4 |
 |
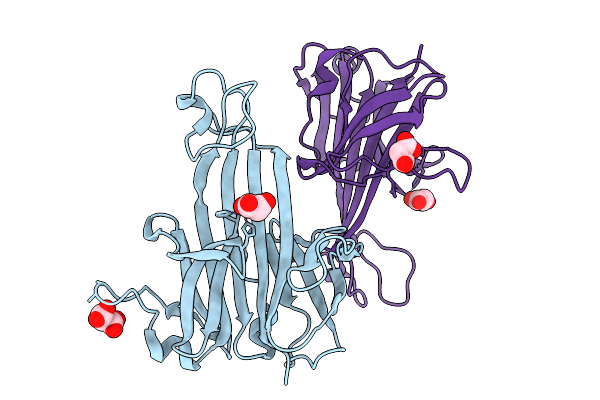
Structure Of Phospholipase D Betaib1I From Sicarius Terrosus Venom, H47N Mutant Bound To Product And Substrate Sphingolipids At 1.85 A Resolution
Organism: Sicarius terrosus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-09-10 Classification: LYASE Ligands: A1A43, A1A44, NA, MG, MPD |
 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2025-09-10 Classification: METAL BINDING PROTEIN Ligands: NI, MLI |
 |
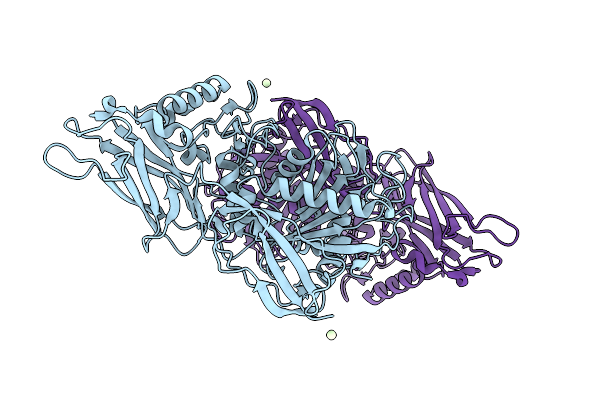
Thomasclavelia Ramosa Immunoglobulin A Protease Middle (Protease) Domain With C-Terminal Domain #1 (Crystallized As Purified)
Organism: Thomasclavelia ramosa
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: ZN, PR |
 |
Thomasclavelia Ramosa Immunoglobulin A Protease Middle (Protease) Domain With C-Terminal Domain #1 (Metal Chelated)
Organism: Thomasclavelia ramosa
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: ZN, PR |
 |
Thomasclavelia Ramosa Immunoglobulin A Protease Middle (Protease) Domain With C-Terminal Domain #1 (Metal Chelated And Zinc Supplemented)
Organism: Thomasclavelia ramosa
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: ZN, EDO |
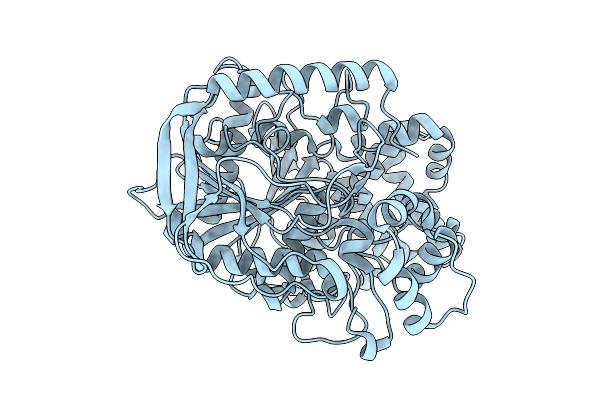 |
Organism: Cochliomyia hominivorax
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2025-08-13 Classification: HYDROLASE |
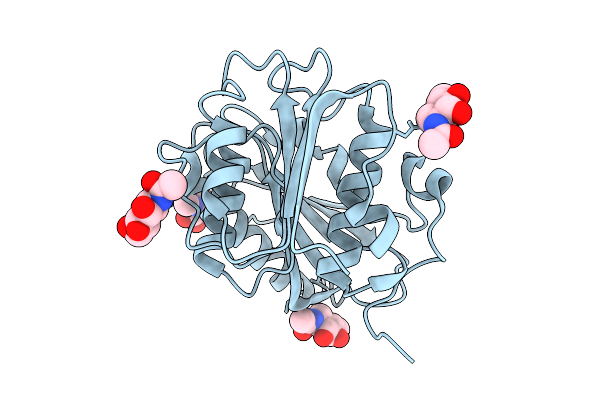 |
Crystal Structure Of Pichia Pastoris-Expressed Fast-Petase-N212A/K233C/S282C Variant
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-07-23 Classification: HYDROLASE Ligands: NAG |
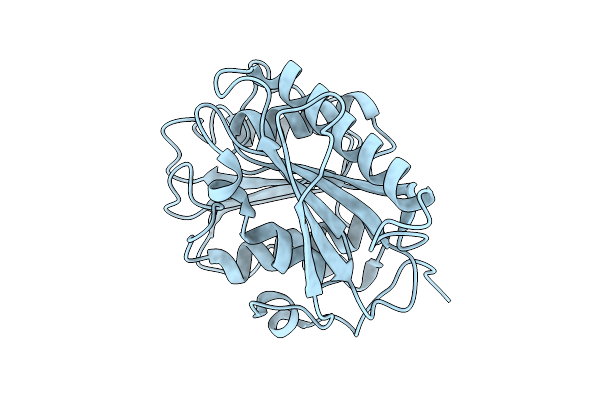 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-07-23 Classification: HYDROLASE |
 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2025-07-23 Classification: HYDROLASE |
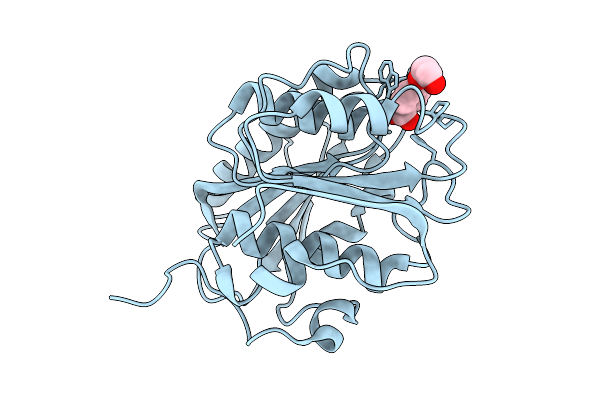 |
Crystal Structure Of Fast-Acc-T140D/R132G/S160A Mutant In Complex With Mono(2-Hydroxyethyl) Terephthalic Acid
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2025-07-23 Classification: HYDROLASE Ligands: C9C |

