Search Count: 99
All
Selected
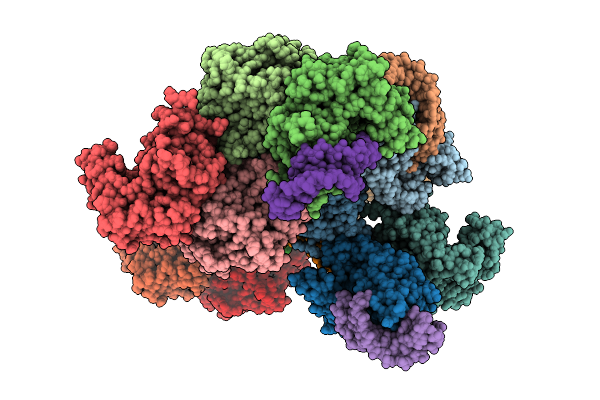 |
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Alanylated Rna Microhelices Analogues Mimicking Ala-Trna-Ala Substrate
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.61 Å Release Date: 2026-03-11 Classification: RNA BINDING PROTEIN |
 |
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: MG, TRT, A1BYH |
 |
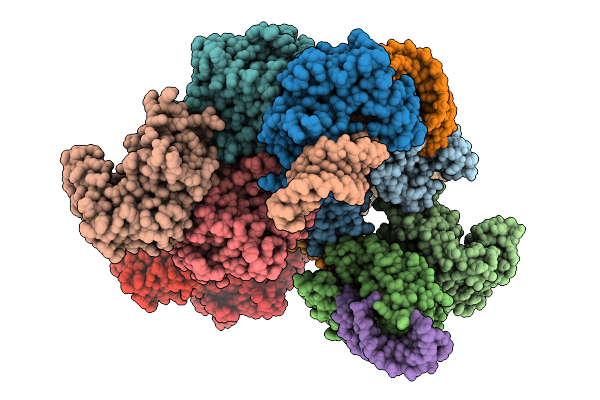
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Ala Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.29 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
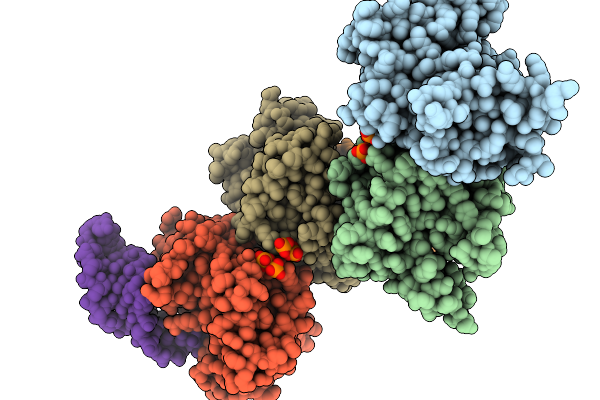
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Glu Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.52 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
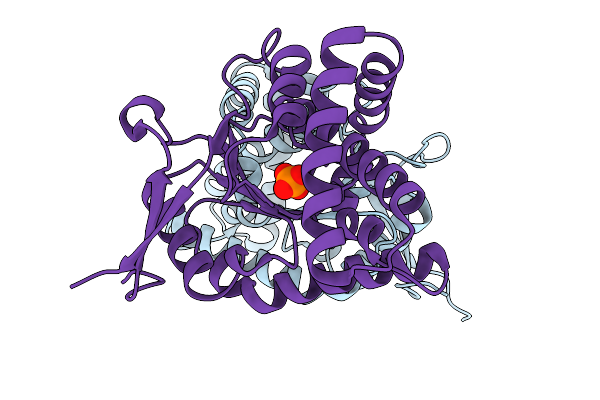
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Glu Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.43 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
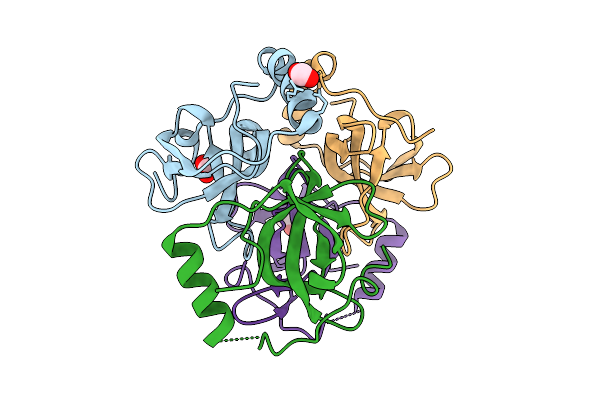
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps)
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-18 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
Organism: Streptococcus intermedius
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-11-05 Classification: CHAPERONE Ligands: FMT, ACY |
 |
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:3.11 Å Release Date: 2025-10-08 Classification: TRANSFERASE Ligands: MG, TRT, GOL |
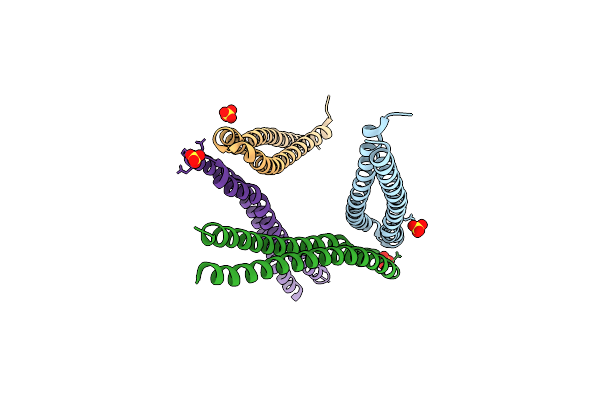 |
Organism: Streptococcus intermedius
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2022-08-24 Classification: PROTEIN BINDING Ligands: SO4 |
 |
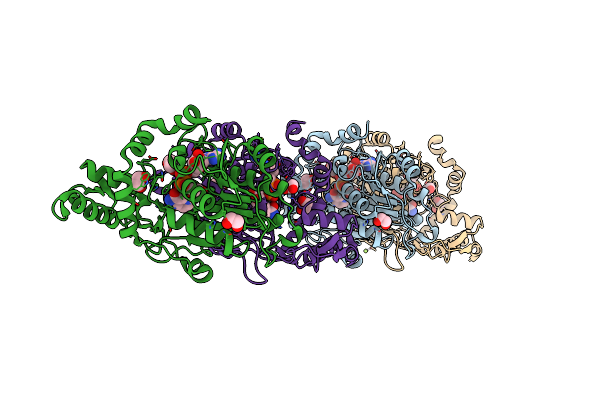
Crystal Structure Of Dna Polymerase Iii Beta Subunit From Elizabethkingia Anophelis
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2022-08-10 Classification: DNA BINDING PROTEIN Ligands: CA, CL, NA |
 |
Crystal Structure Of Nad Bound Dtdp-Glucose 4,6-Dehydratase From Elizabethkingia Anophelis
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2022-08-10 Classification: LYASE Ligands: EDO, NAD, CL, MG |
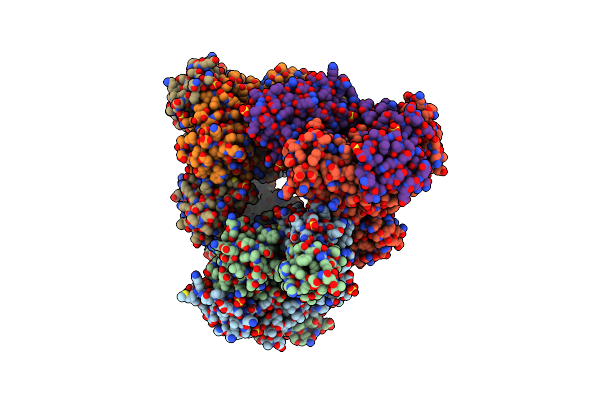 |
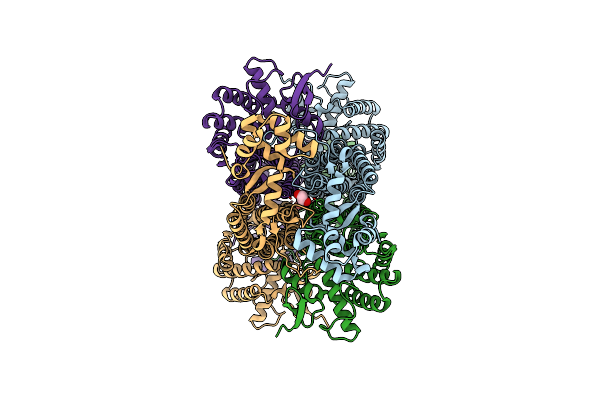
Crystal Structure Of Hydroxymethylglutaryl-Coa Reductase From Elizabethkingia Anophelis Nuhp1
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2021-06-23 Classification: OXIDOREDUCTASE Ligands: CIT, SO4, EDO |
 |
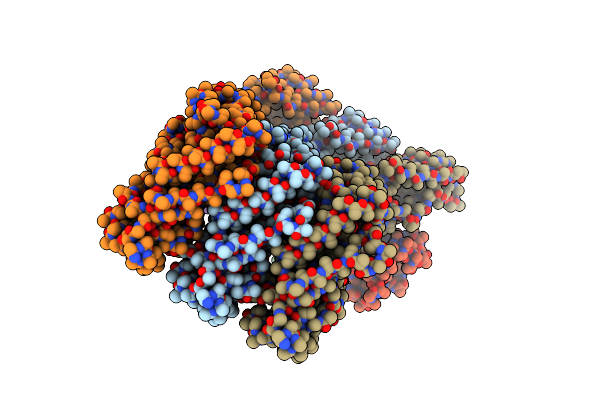
Crystal Structure Of Fumarate Hydratase Class Ii From Elizabethkingia Anophelis Nuhp1
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2021-04-28 Classification: LYASE Ligands: EDO |
 |
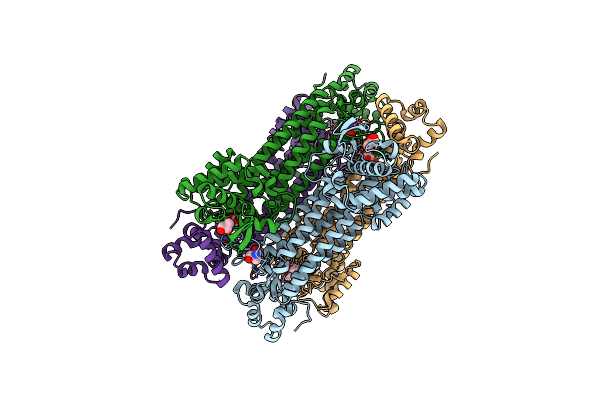
Organism: Streptococcus intermedius, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2020-11-18 Classification: TOXIN |
 |
Crystal Structure Of An Aspartate Ammonia-Lyase From Elizabethkingia Anophelis Nuhp1
Organism: Elizabethkingia anophelis nuh1
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2020-04-29 Classification: LYASE Ligands: TRS, FUM, GOL, CL |
 |
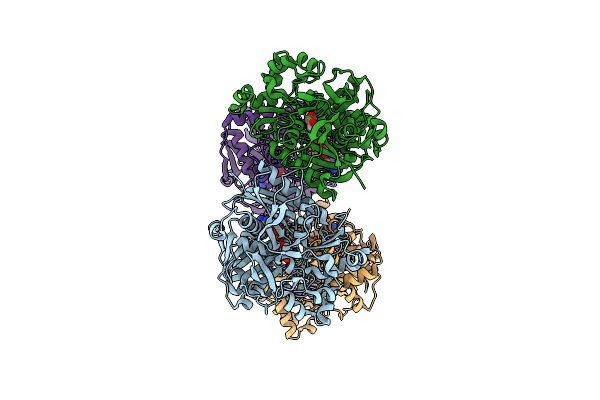
Crystal Structure Of Dihydrolipoyl Dehydrogenase From Elizabethkingia Anophelis Nuhp1
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-01-15 Classification: OXIDOREDUCTASE Ligands: FAD, ZN, CL, EDO |
 |
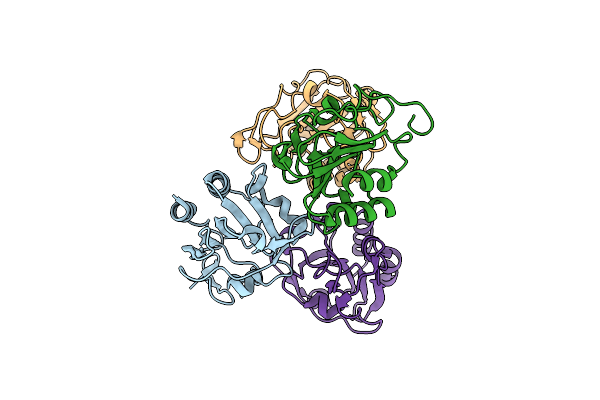
Crystal Structure Of Glycerol Kinase From Elizabethkingia Anophelis Nuhp1 In Complex With Adp And Glycerol
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2019-10-16 Classification: TRANSFERASE Ligands: ADP, GOL, MG |
 |
Crystal Structure Of A Probable Thiol Peroxidase From Elizabethkingia Anophelis Nuhp1
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2019-10-16 Classification: OXIDOREDUCTASE |
 |
Crystal Structure Of Methylglyoxal Synthase From Elizabethkingia Anophelis Nuhp1
Organism: Elizabethkingia anophelis nuhp1
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2019-07-24 Classification: LYASE Ligands: PO4, MPD |
 |
The Structure Of The Variable Domain Of Streptococcus Intermedius Antigen I/Ii (Pas)
Organism: Streptococcus intermedius
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2019-07-17 Classification: CELL ADHESION Ligands: MG |

