Search Count: 3,229
All
Selected
 |
Organism: Staphylococcus aureus
Method: SOLUTION NMR Release Date: 2026-04-15 Classification: PROTEIN BINDING |
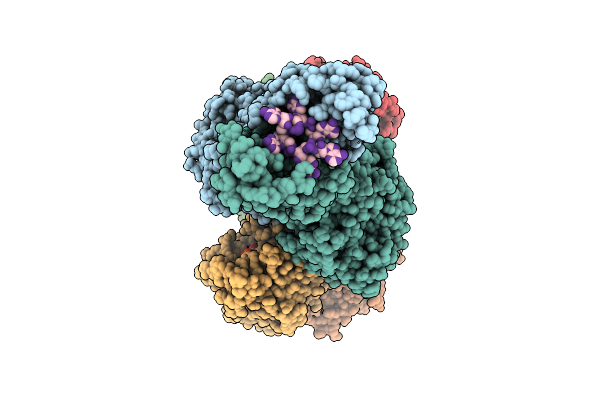 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 3
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, MG, ZN |
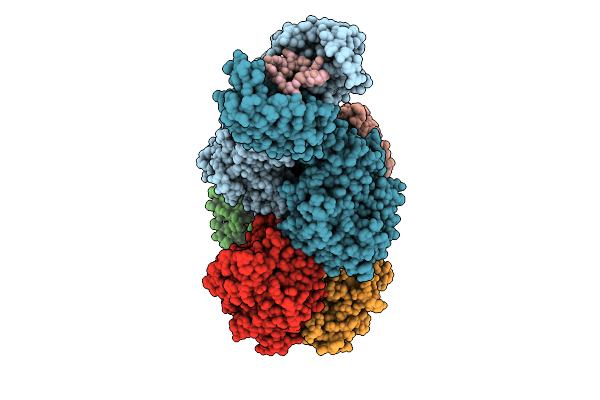 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 4
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
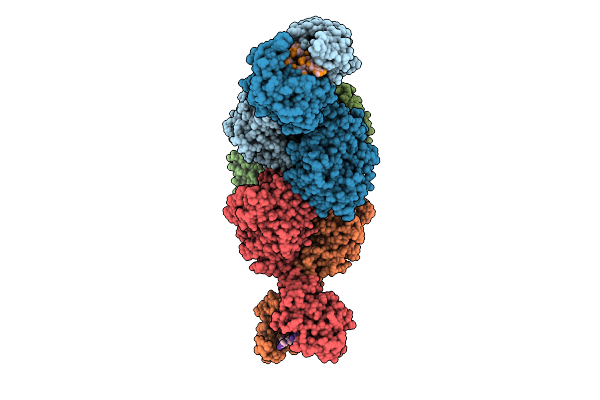 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 5
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
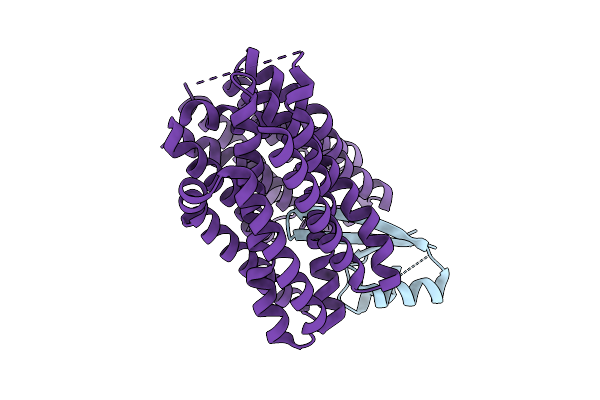 |
Organism: Synthetic construct, Staphylococcus aureus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN |
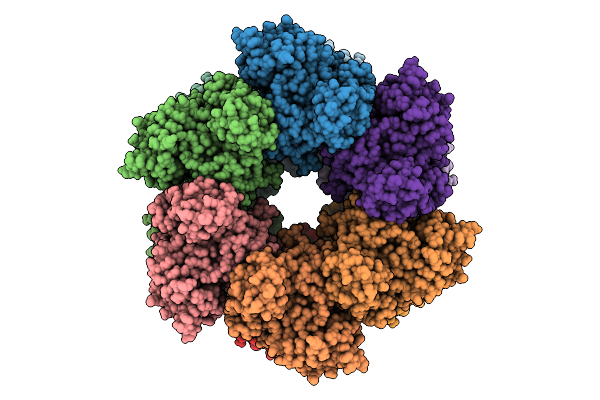 |
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN |
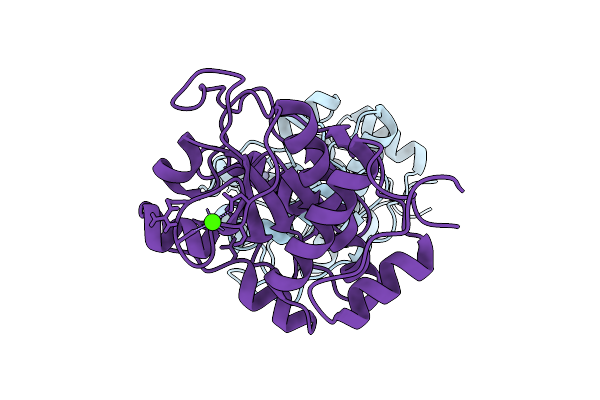 |
Organism: Staphylococcus epidermidis
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: ZN, CA |
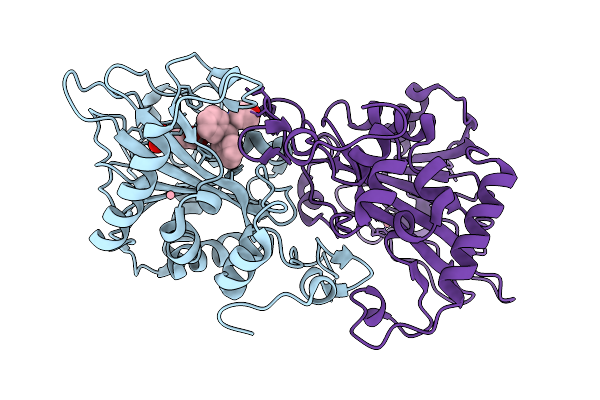 |
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: AKG, CO, A1EM0 |
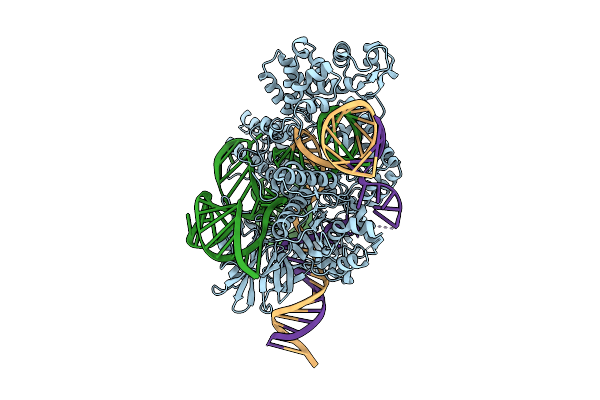 |
Cryo-Em Structure Of Wild-Type Sacas9-Guide Rna-Mismatched Target Dna Complex
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN |
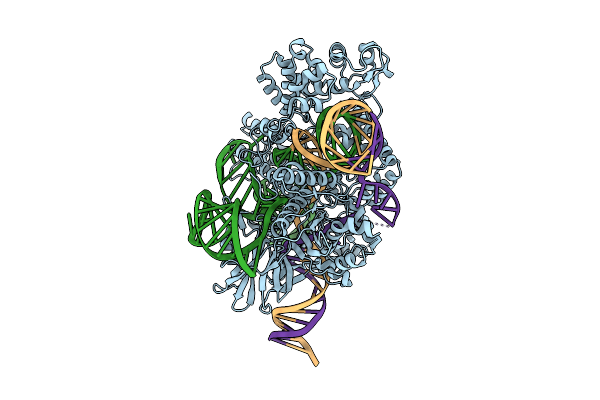 |
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN Ligands: MG |
 |
Organism: Aspergillus fumigatus
Method: ELECTRON MICROSCOPY Resolution:2.76 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: MG, TPP, FAD |
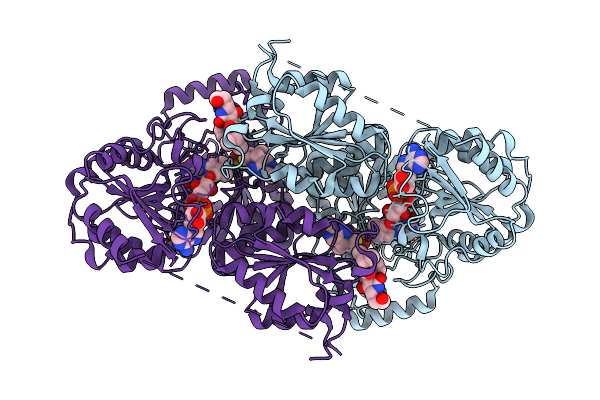 |
Cryoem Structure Of Aspergillus Fumigatus Acetolactate Synthase (Als) In Complex With A Novel Inhibitor
Organism: Aspergillus fumigatus
Method: ELECTRON MICROSCOPY Resolution:2.36 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: MG, TPP, FAD, A1CXH |
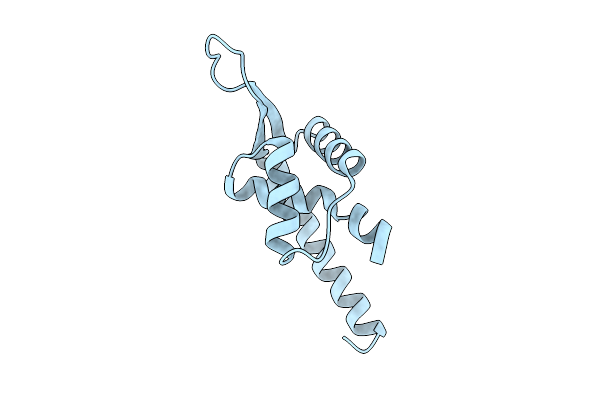 |
Organism: Brachyspira hampsonii
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN |
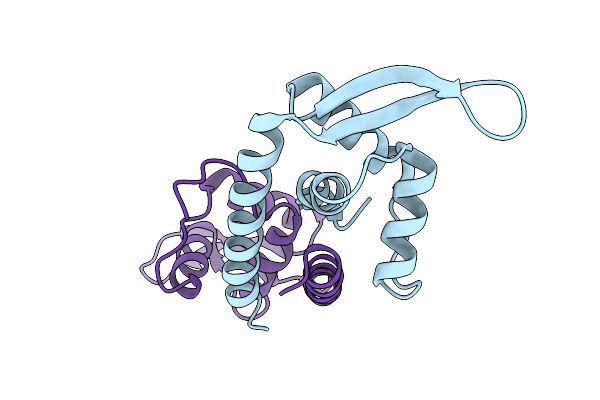 |
Crystal Structure Of Brachyspira Hampsonii Padr With Leu88 Replaced By 3-Aminotyrosine
Organism: Brachyspira hampsonii
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN |
 |
Organism: Staphylococcus aureus
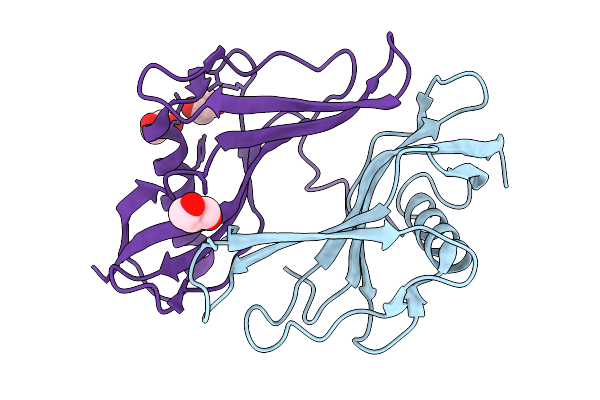
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: PGE, FLC |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: GOL, EDO |
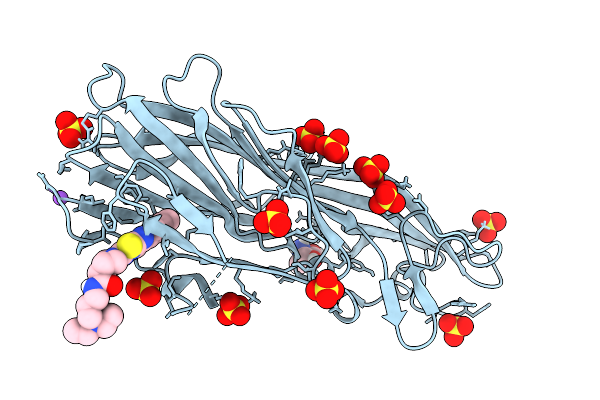 |
Alpha-Hemolysin Of S.Aureus (Monomer) In Complex With The Small Molecule N-(3-(Diethylamino)Propyl)-1-(6-(P-Tolyl)Imidazo[2,1-B][1,3,4]Thiadiazol-2-Yl)Piperidine-4-Carboxamide
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-04 Classification: TOXIN Ligands: A1JN0, TRS, SO4, NA |
 |
Organism: Staphylococcus aureus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: RIBOSOME |
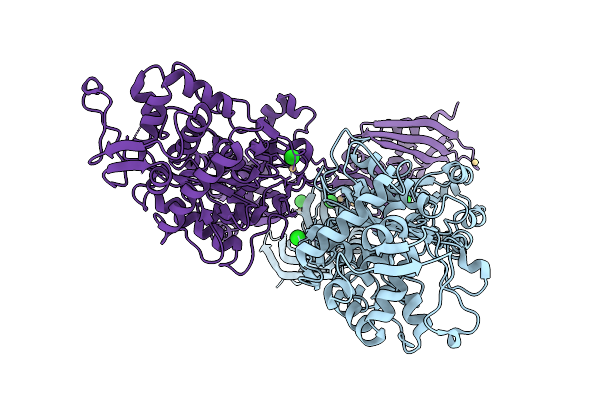 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |

