Search Count: 1,220
All
Selected
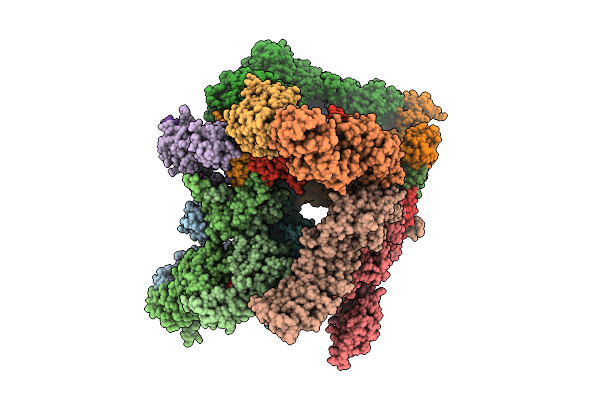 |
Structure Of Human 26S Proteasome Complexed With Midnolin, 19S Proteasome With Ubl And Catch Domain Resolved
Organism: Escherichia coli k-12, Homo sapiens, Pseudotamlana agarivorans
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: HYDROLASE Ligands: ADP, ATP, ZN, MG |
 |
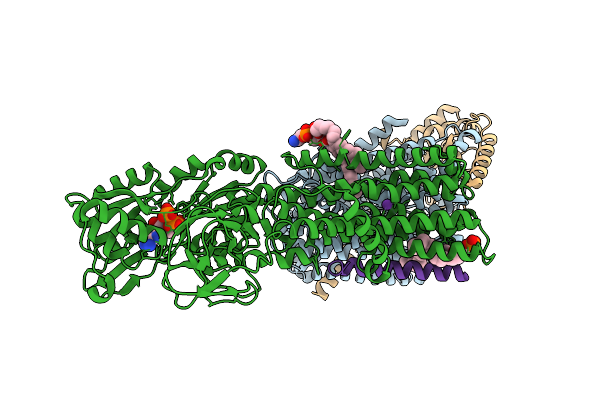
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1 State In Lipid Nanodisc
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.34 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1-Atp State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.28 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, ATP |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.25 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, MG |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.58 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2 State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.72 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
Organism: Coprinopsis cinerea okayama7#130
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2026-03-18 Classification: TOXIN |
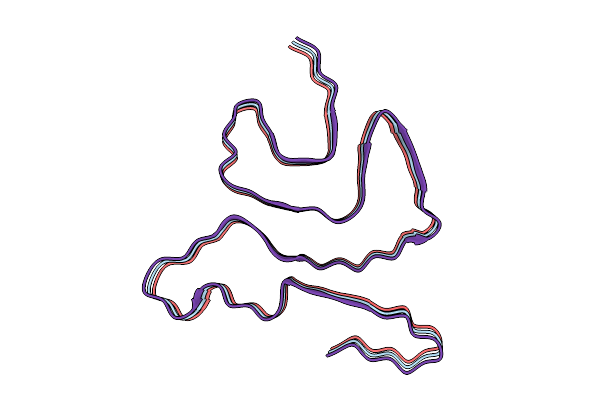 |
The Cryo-Em Structure Of In Situ Amplified (Isa) Alpha-Synuclein Fibrils From Pd Homogenate
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: PROTEIN FIBRIL |
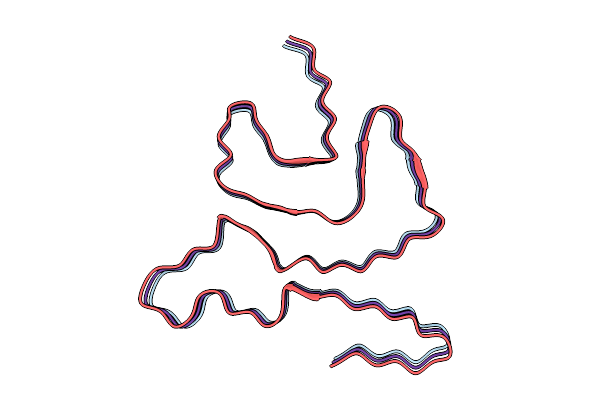 |
The Cryo-Em Structure Of In Situ Amplified (Isa) Alpha-Synuclein Fibrils From Dlb Homogenate
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: PROTEIN FIBRIL |
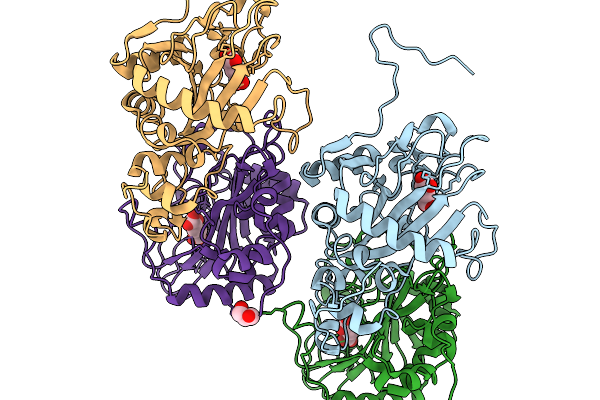 |
Organism: Penicillium simplicissimum
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-03-04 Classification: ANTIBIOTIC Ligands: CO, AKG, GOL |
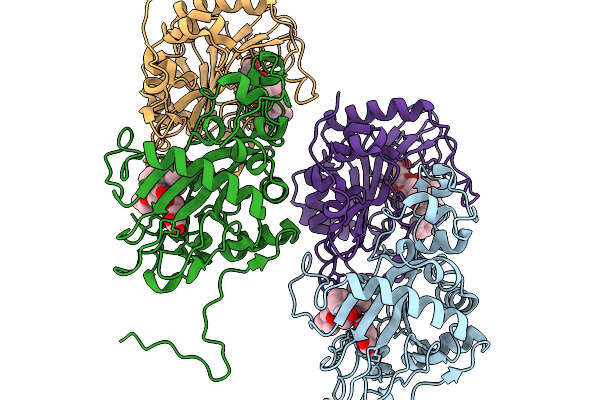 |
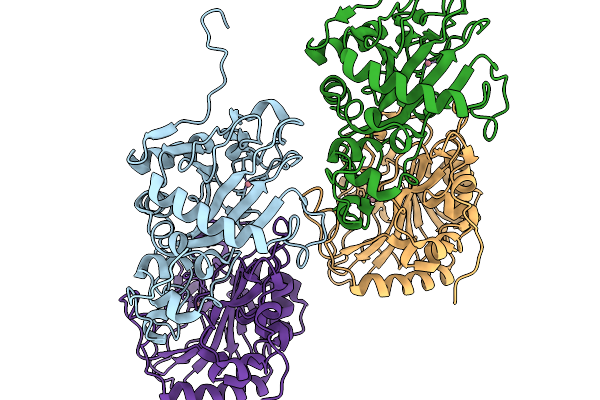
Crystal Structure Of Okae In Complex With A-Ketoglutarate And Okaramine A At 2.5 Angstroms Resolution.
Organism: Penicillium simplicissimum
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CO, A1EMZ, AKG |
 |
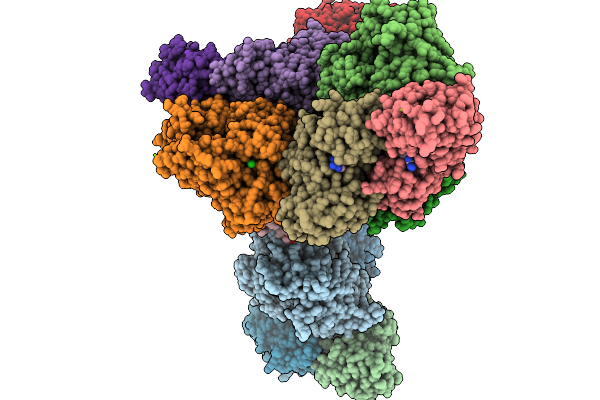
Organism: Penicillium simplicissimum
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CO |
 |
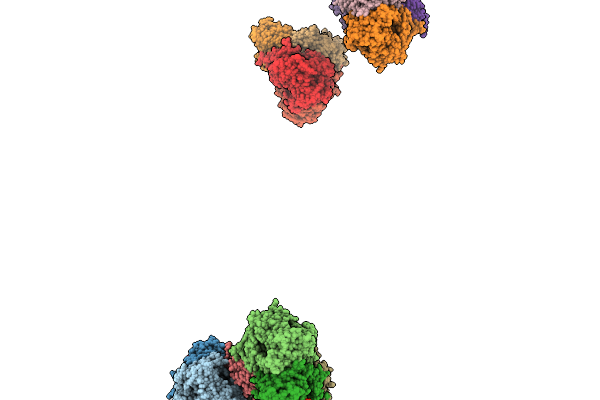
Organism: Thauera chlorobenzoica
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-02-18 Classification: OXIDOREDUCTASE Ligands: SF4, CL, GOL, K, MG, ATP, ADP |
 |
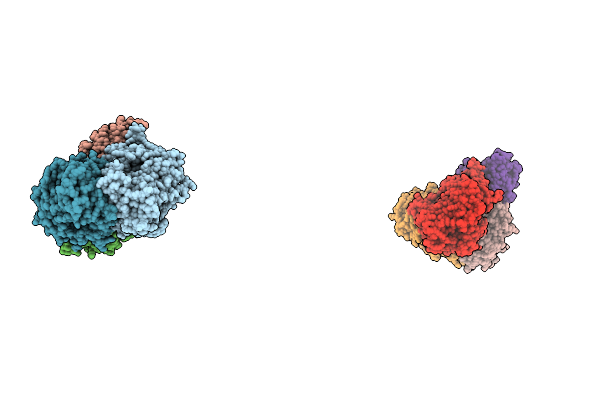
3-Methylbenzoyl-Coa Reductase From Thauera Chlorobenzoica (Subunits Mbdon ) + Adp
Organism: Thauera chlorobenzoica
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-02-18 Classification: OXIDOREDUCTASE Ligands: SF4, SO4, ATP, MG, ADP |
 |
3-Methylbenzoyl-Coa Reductase From Thauera Chlorobenzoica (Mbdonpq) + Amppnp
Organism: Thauera chlorobenzoica
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-02-18 Classification: OXIDOREDUCTASE Ligands: SF4, ATP, MG, ANP, GOL |
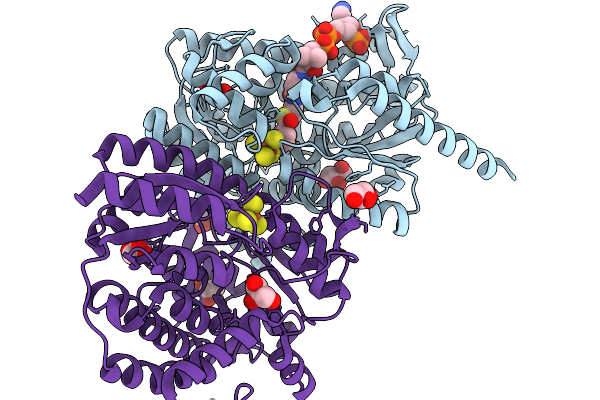 |
3-Methylbenzoyl-Coa Reductase From Thauera Chlorobenzoica (Subunits Mbdon )
Organism: Thauera chlorobenzoica
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE Ligands: SF4, BYC, GOL, CL |
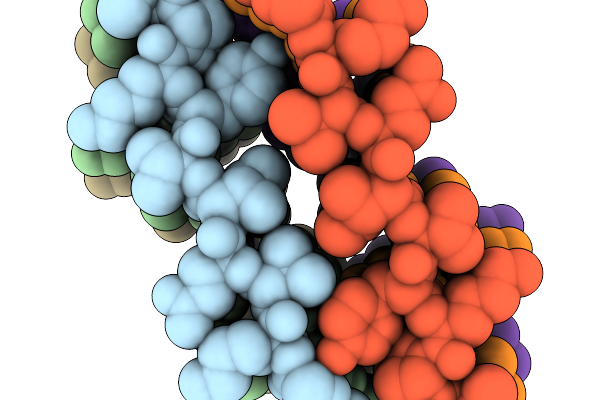 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: PROTEIN FIBRIL |
 |
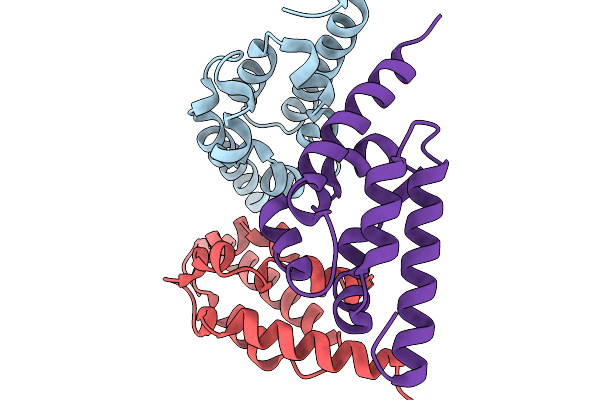
Crystal Structure Of Anti-Crispr Acrie7 Determined By Experimental Mad Phasing With Hg
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: HG |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-02-04 Classification: UNKNOWN FUNCTION Ligands: CL |
 |
Organism: Hydrogenivirga sp. 128-5-r1-1
Method: ELECTRON MICROSCOPY Resolution:2.91 Å Release Date: 2026-02-04 Classification: TRANSFERASE Ligands: A1C3G |

