Search Count: 4,844
All
Selected
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2026-05-06 Classification: HYDROLASE |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-05-06 Classification: HYDROLASE |
 |
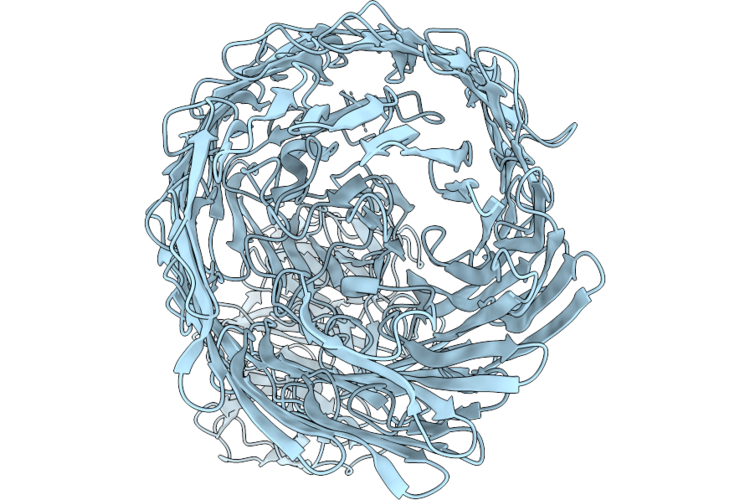
Chap Domain Of Staphylococcus Aureus-Specific Lysin L1 Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA, SCN |
 |
Staphylococcus Aureus-Specific Lysin L1-3 (Lysm-Chap) Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA |
 |
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TOXIN |
 |
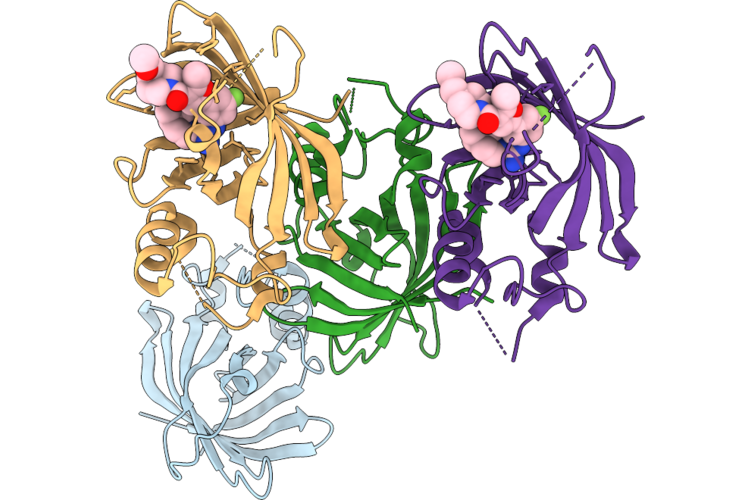
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-05-06 Classification: LIGASE Ligands: A1JCC |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-05-06 Classification: LIGASE Ligands: A1JCD |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-06 Classification: LIGASE Ligands: A1JCE |
 |
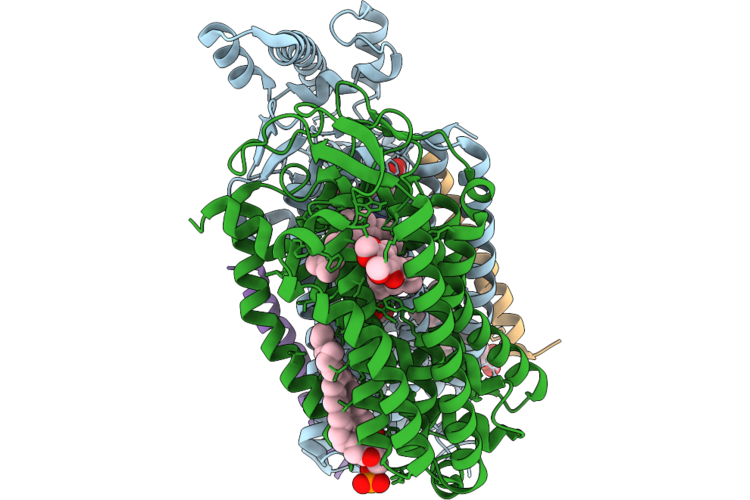
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, A1JN4, MQ9, LPP, UQ8, OXY, PGT, POV |
 |
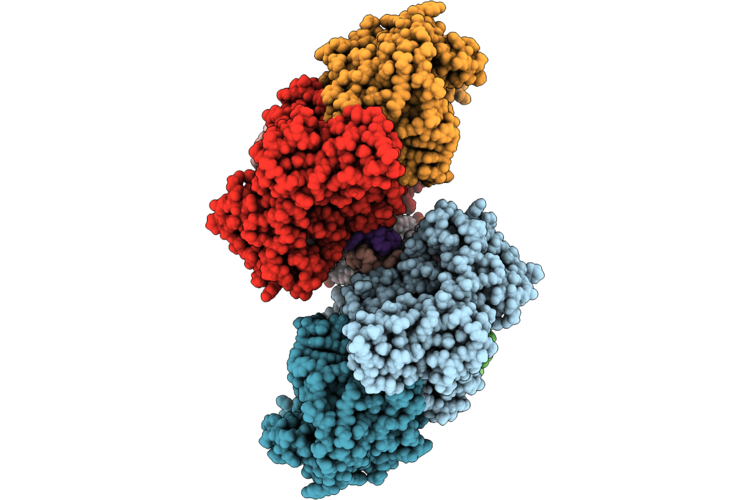
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1JN4, HEM, LPP, OXY, UQ8 |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, LPP, UQ8, 0NI, OXY, A1JN4, PGT, POV |
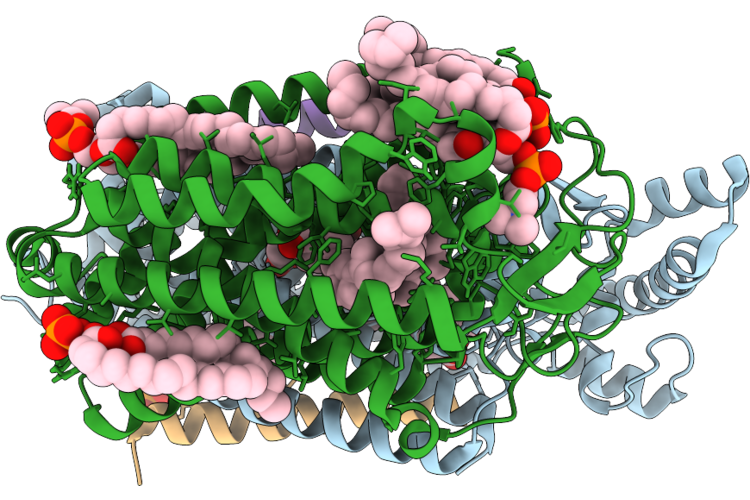 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1JN4, HEM, LPP, OXY, POV, MQ9 |
 |
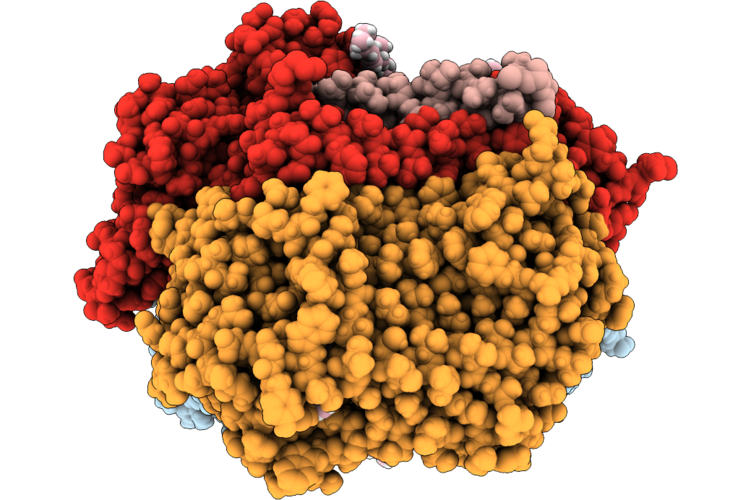
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1JN4, HEM, LPP, OXY, UQ8 |
 |
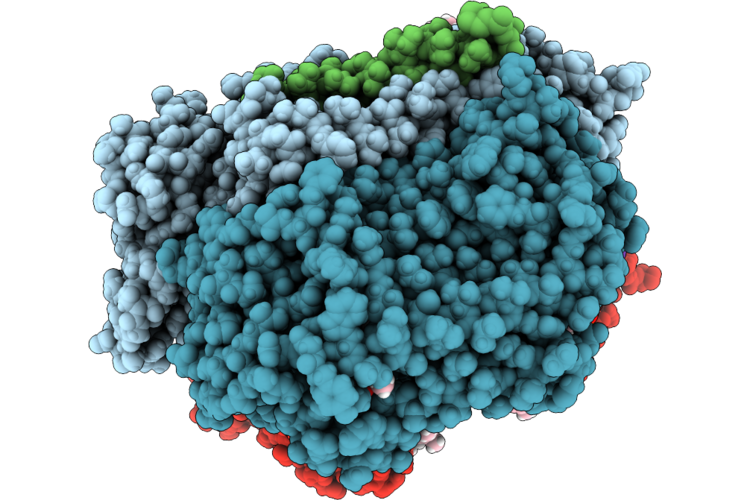
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, A1JN4, LPP, UQ8, MQ8, OXY |
 |
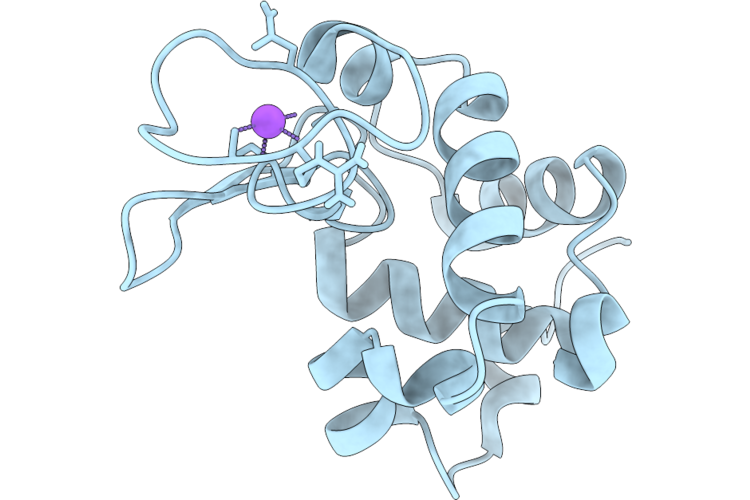
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: LPP, UQ8, HEM, MQ8, A1JN4, OXY |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CL, NA |
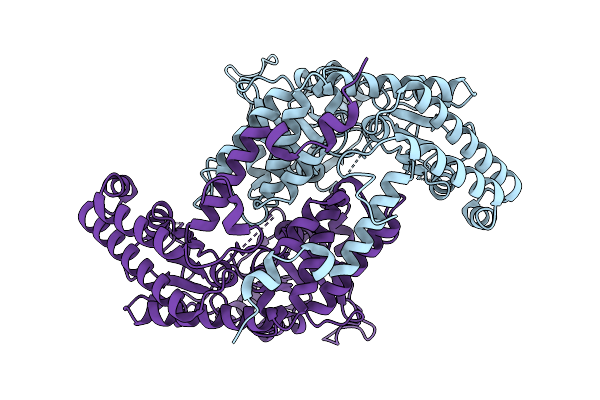 |
Cryo-Em Structure Of The A149T Dimer Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Essex In Complex With Plp
Organism: Glycine max
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE |
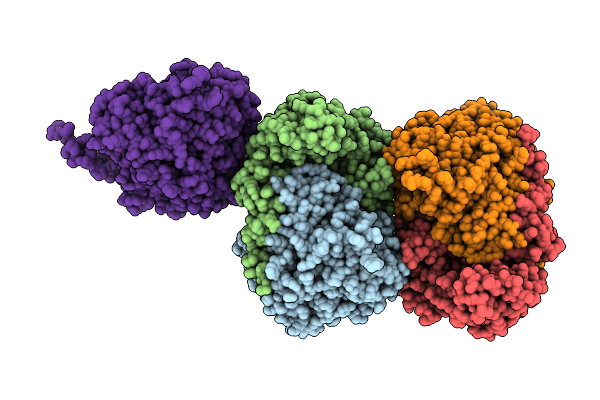 |
Crystal Structure Of A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Essex In Complex With Plp-Glycine
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PLG |
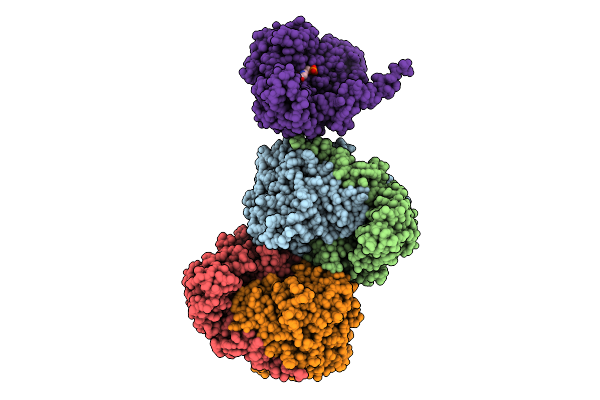 |
Crystal Structure Of The A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Forrest In Complex With Plp-Glycine
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PLG |
 |
Crystal Structure Of The A149T Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Forrest In Complex With Plp
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-08 Classification: TRANSFERASE |

