Overview of IBDC
Welcome to the data archiving services provided by the Indian Biological Data Centre (IBDC). This guide is intended to help users understand the standard operating procedures (SOPs) for the submission of animal phenotyping data. Before initiating data submission, users are encouraged to familiarize themselves with the structure, scope, and mandate of the IBDC and its specialized data repositories.
IBDC serves as a national platform for the submission, archival, and dissemination of diverse biological data types through a suite of dedicated, data type–specific portals. At present, IBDC provides active archival services for nucleotide, phenome, metabolome, image, and proteome data.
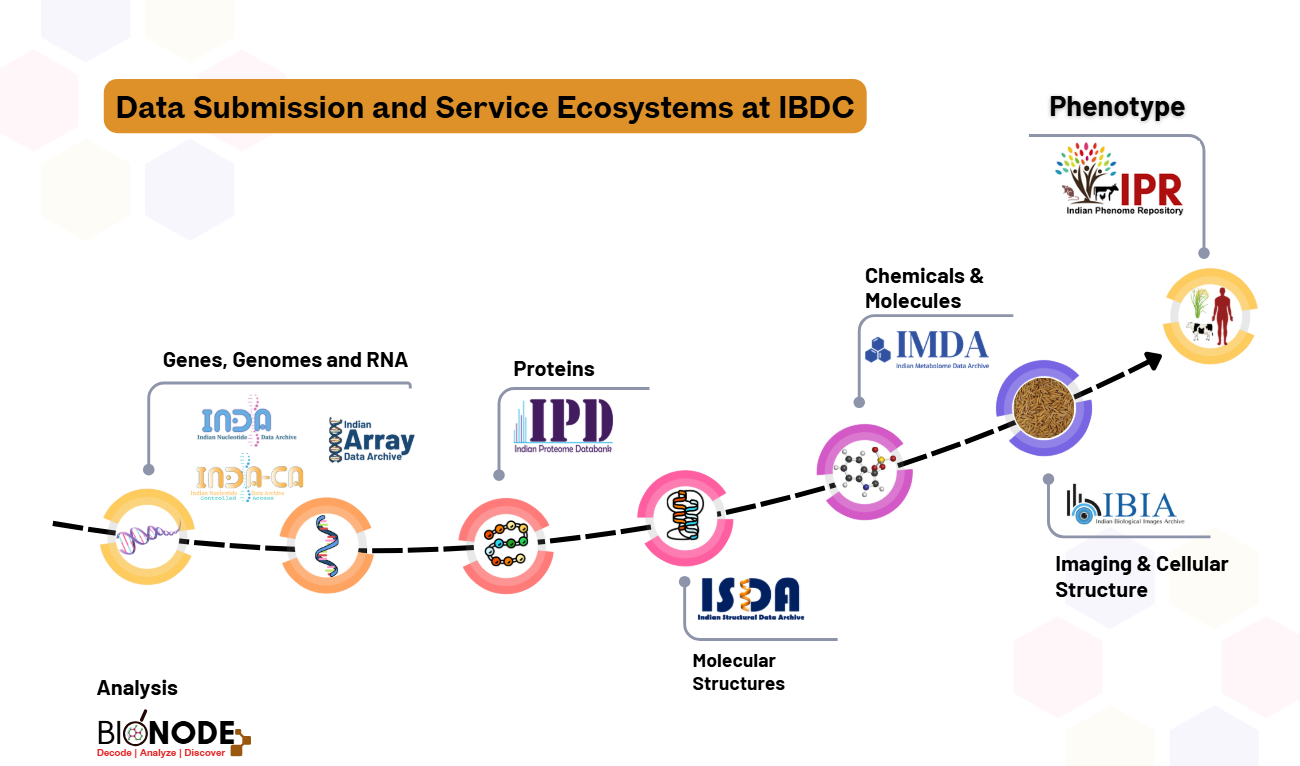
Figure 1. The diversity of data portals at Indian Biological Data Centre (IBDC)
All sequencing-based nucleotide data submissions are handled through the Indian Nucleotide Data Archive (INDA and INDA-CA), while array-based nucleotide data are submitted to the Indian Array Data Archive (IADA). Metabolome, biological image, and proteome data are archived via the Indian Metabolome Data Archive (IMDA), Indian Biological Images Archive (IBIA), and Indian Proteome Databank (IPD), respectively.
Phenomic data are archived through the Indian Phenome Repository (IPR), which comprises four specialized sister repositories: the Indian Crop Phenome Database (ICPD), the Indian Animal Phenome Database (IAPD), and upcoming repositories for human and microorganism phenome data.
This manual provides detailed standard operating procedures for the submission of animal phenotyping data to the Indian Animal Phenome Database (IAPD).
Overview of the Indian Phenome Repository (IPR)
The Indian Phenome Repository (IPR) is a national initiative established under the Indian Biological Data Centre (IBDC) to enable systematic submission, long-term archival, and broad access to high-quality phenome data across agriculture, animal sciences, human health, and microbiology.
Aligned with FAIR (Findable, Accessible, Interoperable, Reusable) and CARE (Collective Benefit, Authority to Control, Responsibility, Ethics) data principles, IPR ensures responsible, ethical, and reusable phenotypic data stewardship, with respect for community and Indigenous data sovereignty.
Due to the complexity and heterogeneity of phenome data, IPR adopts a modular, domain-specific repository architecture that supports tailored metadata standards, workflows, and access policies while maintaining a common underlying infrastructure..
Repository Structure
- Indian Crop Phenome Database (ICPD): ICPD digitizes multi-crop phenome data generated from agricultural trials and field experiments. It provides a unified platform for archiving, sharing, and reusing crop phenotype datasets in compliance with FAIR principles. All accepted submissions are assigned unique and persistent IBDC accession identifiers.
- Indian Animal Phenome Database (IAPD): IAPD captures phenotypic traits of indigenous, domesticated, and economically important animal species. It supports research and innovation in veterinary science, livestock breeding, productivity enhancement, disease resistance, and environmental adaptation. The database integrates traditional knowledge systems with modern phenotyping frameworks to facilitate national and global data harmonization.
- Indian Human Phenome Database (IHPD): IHPD curates high-quality phenotypic data from diverse Indian populations, encompassing clinical, physiological, genetic, and environmental traits. It underpins research in precision medicine, public health, population genetics, and epidemiology, while reflecting India’s rich biological and socio-cultural diversity under appropriate ethical and regulatory oversight.
- Indian Microorganism Phenome Database (IMPD): IMPD documents phenotypic characteristics of microorganisms isolated from varied Indian environments. It supports research in agriculture, biotechnology, industry, environmental sciences, and medicine by enabling standardized representation and reuse of microbial phenome data.

Figure 2. Four branches of IPR.
Mission and Objectives
- Centralized and standardized national platform for phenotypic data
- Domain-specific metadata frameworks for interoperability
- Template-based, low-overhead data submission
- User-friendly tools for querying, visualization, and download
Indian Animal Phenome Database (IAPD)
The Indian Animal Phenome Database (IAPD) is a specialized repository under the Indian Phenome Repository, developed to support the systematic submission, curation, integration, and sharing of animal phenotypic data. IAPD primarily focuses on livestock and indigenous animal species of India, with the goal of strengthening research, breeding programs, conservation efforts, and evidence-based policymaking.
Much of the animal trial and phenotyping data generated across institutions— particularly for livestock—remains unpublished, inaccessible to the wider research community, or is eventually lost over time. IAPD addresses this challenge by serving as a single, user-friendly platform for the long-term archival, organization, analysis, and sharing of multi-animal phenome data. All data submitted to IAPD are assigned unique and persistent IBDC accession identifiers and are managed in accordance with the FAIR (Findable, Accessible, Interoperable, and Reusable) data principles.
Figure 3. The screenshot of IAPD Homepage
IAPD aligns closely with national priorities in biotechnology, agriculture, and biodiversity conservation. By bringing together traditional livestock knowledge systems and modern phenotyping approaches, the platform enables data-driven decision-making and encourages collaborative research aimed at building sustainable and resilient animal production systems.
Objectives
The objectives of the Indian Animal Phenome Database are to:
- Establish a centralized national repository for animal phenotypic data.
- Enable standardization and harmonization of phenotypic data across studies, institutions, and regions.
- Support biotechnology-led research and innovation in animal breeding, genomics, and phenomics.
- Facilitate genetic improvement and conservation of indigenous and endangered animal breeds.
- Provide a robust scientific evidence base for policy development and implementation.
- Promote data sharing, collaboration, and capacity building among stakeholders.
Scope
The scope of IAPD encompasses:
- Collection and management of morphological, physiological, production, reproductive, behavioural, and health-related traits.
- Coverage of livestock species and native Indian animal breeds, including region-specific populations.
- Integration of phenotypic data generated through field studies, experimental research, surveys, and high-throughput phenotyping platforms.
- Support for multi-tiered data submission, including project, study, organism, trait, and dataset-level information.
- Access for research institutions, universities, breeders, conservation agencies, and policy bodies.
Key Features
IAPD offers the following key features to support users throughout the data submission and access lifecycle:
- Standardized Data Submission Framework: Clearly defined templates and SOPs to ensure data quality, consistency, and interoperability.
- Data Harmonization and Curation: Use of controlled vocabularies (Uberon, ATOL etc), validation checks, and expert curation workflows to enhance reliability.
- Secure, Role-Based Access: Compliance with national data sharing, ethical, and governance guidelines.
- User-Centric Dashboard: An integrated interface for managing submissions, tracking progress, and viewing summary statistics.
- Support and Documentation: Access to SOPs, FAQs, discussion forums, and formal mechanisms for data correction or deletion requests.
- Policy and Compliance Alignment: Adherence to national data policies and best practices for responsible data use.
- Facilitation of Collaboration: Enables coordinated research and knowledge exchange across institutions and disciplines.
The IAPD home page provides an overview of the total number of submissions and offers intuitive tools for submitting animal phenome data. Users can also browse, search, and explore existing datasets through user-friendly interfaces, making it easy to discover, access, and analyze valuable phenotypic information. This streamlined design ensures that contributors and data users alike can efficiently engage with and benefit from the repository.
Overview of metadata used in IAPD
All submissions to the Indian Animal Phenome Database (IAPD) are structured using a well-defined set of metadata objects (Figure 4). These metadata objects ensure that animal phenotypic and trial data are described in a consistent, comprehensive, and reusable manner.
In IAPD, a typical animal phenotyping or trial dataset is organized under a Project (Phenoproject). A phenoproject may include one or more studies, each representing observations made on animals under specific conditions such as species or breed, developmental stage, tissue type, management practices, or environmental treatments.
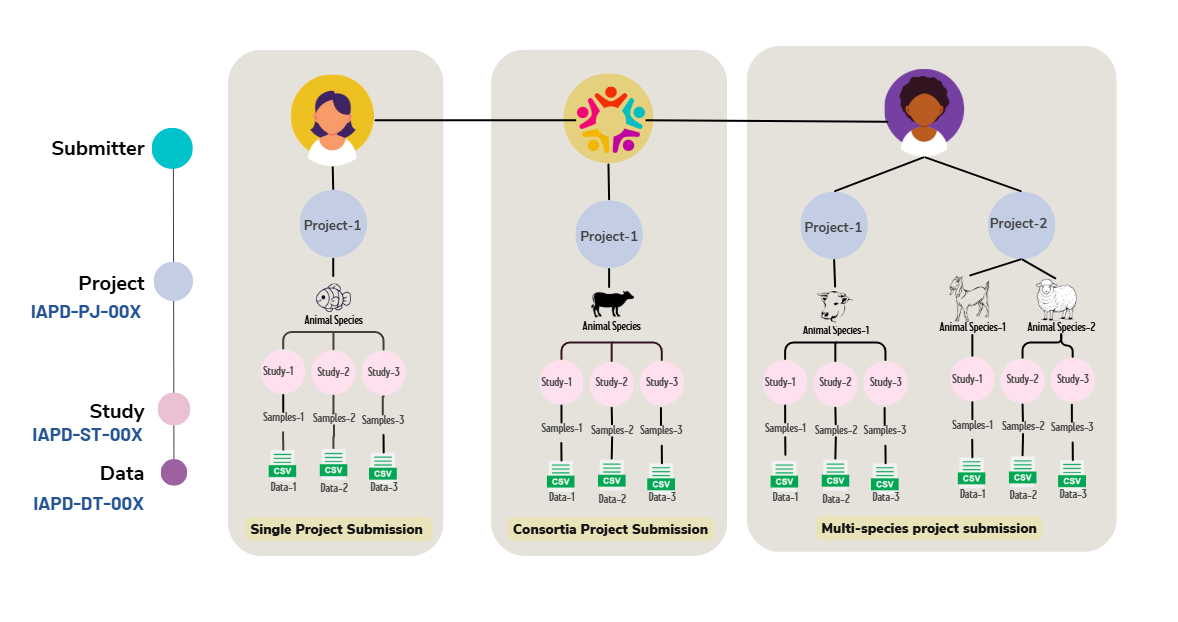
Figure 4. The metadata objects of IAPD.
Each study, in turn, contains observation data for one or more phenotypic traits that are measured and analysed together. The actual measurements and observations are submitted as data files, which are linked to their corresponding studies and projects.
To support this hierarchical organization, the IAPD data model is built around three core modules:
- Project – captures high-level information about the overall research initiative or trial.
- Study – describes individual experimental units or observational studies conducted within a project.
- Data File – contains the submitted phenotypic measurements and associated records.
Each of these core modules is further defined using multiple metadata sub-objects to provide detailed contextual information, enabling accurate interpretation, integration, and reuse of the data.
The structure and definitions of the different IAPD metadata modules are described in detail in the following sections.
Getting Started on Submission
The overview of steps required for the initiate and complete the phenome data submission to IAPD are summarised in Figure 5.
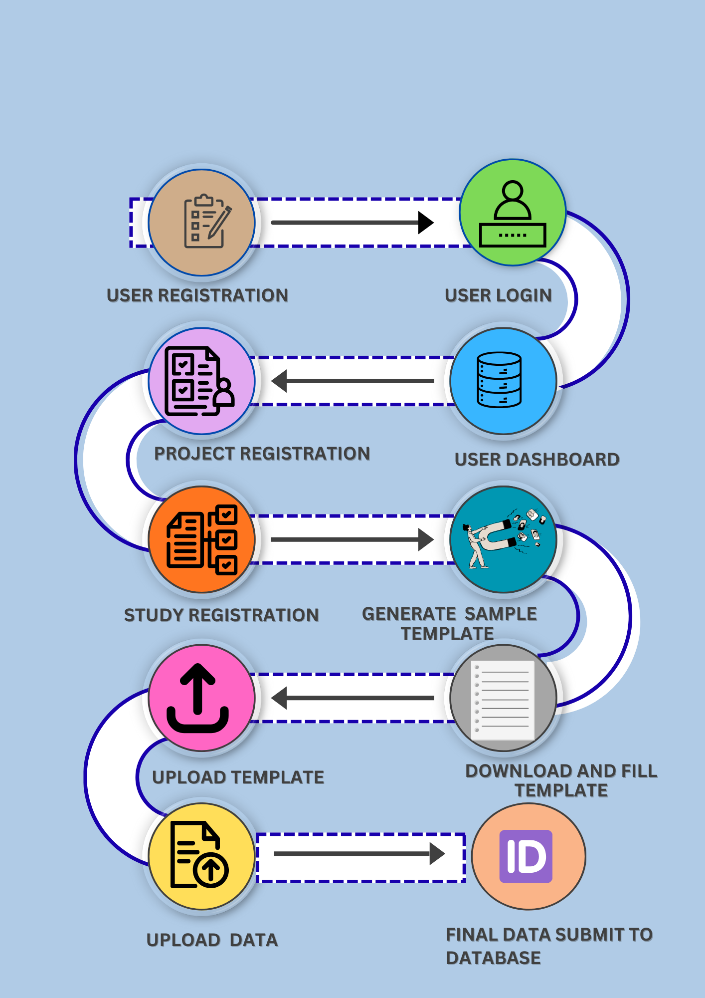
Figure 5. Data submission flow of IAPD.
Step A: User Registration
First time users can click on the Get Started button, they are redirected to the Registration Page (Figure 6), where basic user details are requested. Upon successful completion and submission of the registration form, the user will receive the registration confirmation and can now proceed to the next step to log in to the IPR dashboard.
Figure 6. Registration Page.
Step B: Login & User Dashboard
Registered users can access the user dashboard by entering the login credentials created during the registration process. Users must provide the registered username and the corresponding password to log in successfully. The login page also includes the following options to assist users: • Forgot Password: To reset the password if it has been forgotten • Register Now: To create a new user account • Resend Activation Link: To receive the account activation email again if not previously activated These options ensure smooth access and account recovery for users of the portal
Figure 7. Snapshot of Login page of IAPD.
User Dashboard The user dashboard (figure 8) consists of links to submit different parts of data submission on the left hand side of the dashboard and submission stats on the main part of the page. The view project displays the list of details of previously submitted project, study and/or data by the user along with the filtering options in the left (Figure 8b). User may submit their data by clicking on “Submit Data” link. On the right top corner, the options related to user profile are shown along with logout button. On the left panel, tools and user resources are listed. Help section provide users with the SOP, deletion request form and FAQs related to the IAPD data submission. Discussion forum allows users to convey their suggestions and queries regarding the submission process. Data policy lists the rules and regulations followed at IAPD about the data access & usage, privacy & data protection, data contribution and curation, limitation of liability, etc.
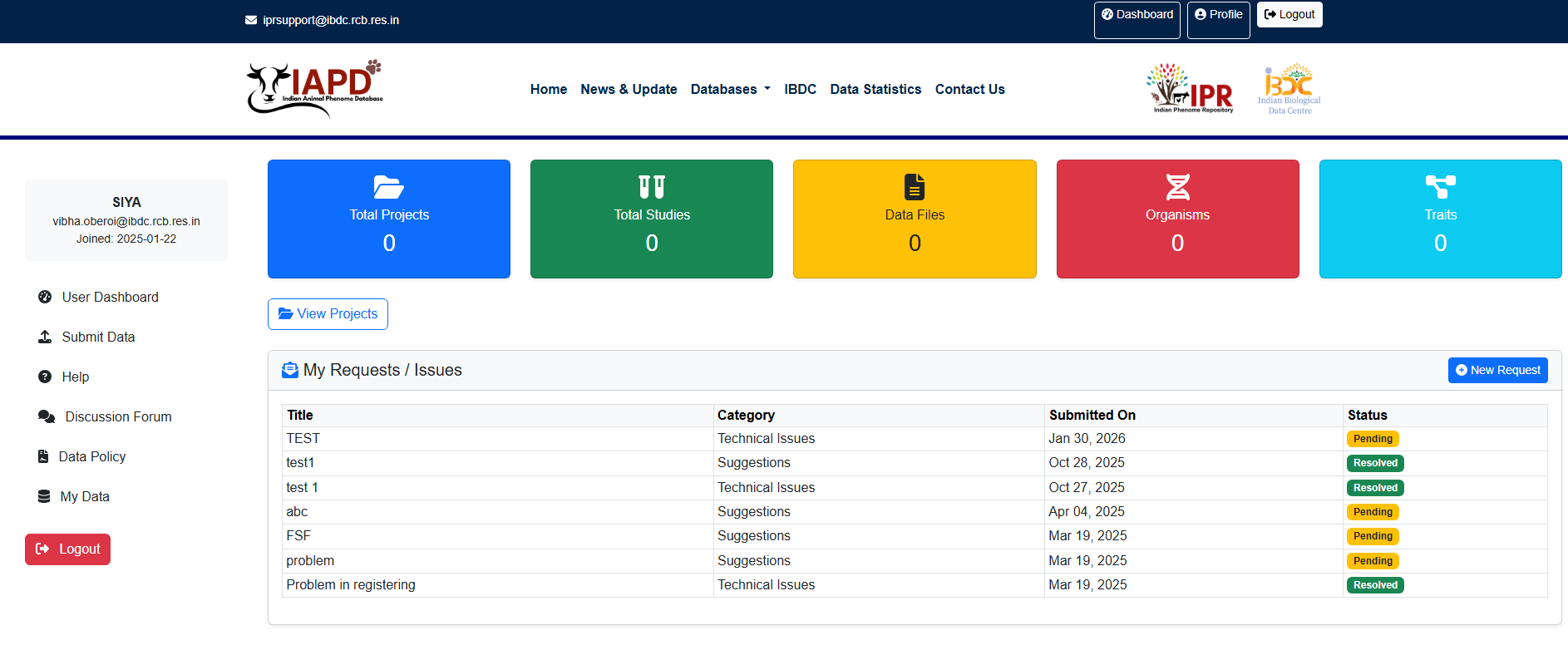
Figure 8a.Screenshot of the IAPD dashboard
Figure 8b. Screenshot of the View Project Page
Step C: Data Submission Process
The Indian Animal Phenome Database (IAPD) recommends users submit data through the online, web-based interactive submission interface, which provides several automated selection fields for capturing project- and study-level metadata mapped to relevant ontology terms. The web-based submission workflow consists of five sequential steps, each of which is described in detail below.
Step 1 – Register Project
The Register Project module is the first mandatory step for submitting data to the Indian Animal Phenome Database (IAPD). Users must provide basic project details, publication information, data access conditions, ethical compliance, and organism information.
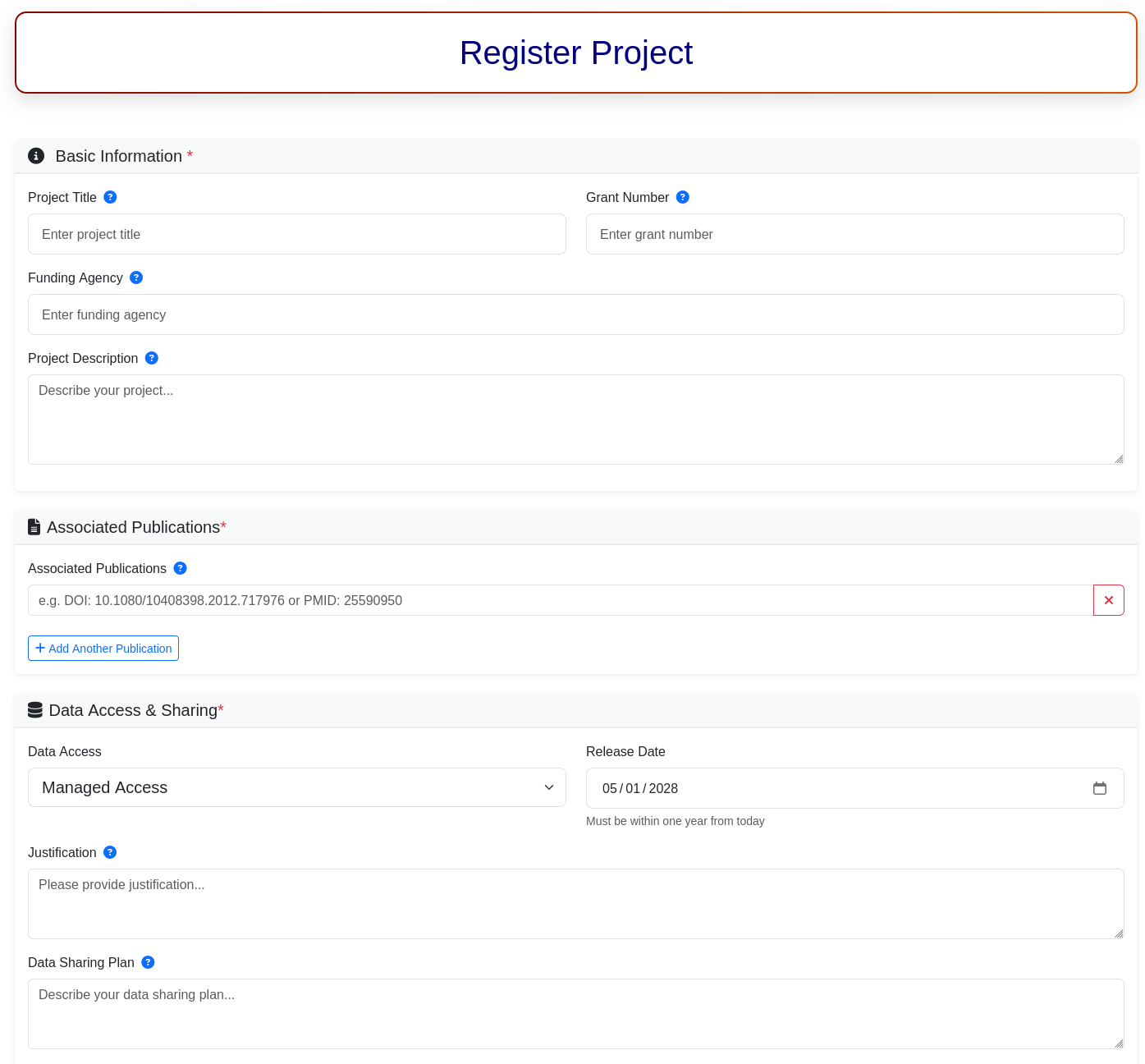
Figure 9. Snapshot of Registration page of Project.
1.1. Basic Information*
Fill in the core details of the research project:
- Project Title: Enter the official title of the research project.
- Grant Number: Provide the funding or grant reference number (if applicable).
- Funding Agency: Specify the name of the funding organization.
- Project Description: Briefly describe the objectives, scope, and relevance of the project.
1.2. Associated Publications*
Link publications related to the project:
- Enter a DOI (e.g., 10.1080/10408398.2012.717976) or PMID.
- Click “Add Another Publication” to include multiple publications.
- Use the delete (✖) option to remove any incorrect entry.
1.3. Data Access & Sharing*
Define how and when the data will be made available:
- Data Access: Select the appropriate access type (e.g., Open, Managed, or No Access).
- Release Date: Choose a release date (must be within one year from the current date).
- Justification: Provide a justification for No Access or Managed Access, if applicable.
- Data Sharing Plan: Describe how the data will be shared, accessed, and reused.
1.4. Project Team*
- Project Type: Specify whether the project is an individual project or part of a consortium.
- If the project is a consortium, provide details of collaborating investigators or institutions.
1.5. Ethical Clearance*
- Enable Ethical Clearance Obtained if the project has received approval from the relevant ethics committee.
Note: Ethical clearance is mandatory for animal phenome data submission.
1.6. Organism Information*
Provide details of the organism studied:
- Scientific Name: Enter the scientific name (e.g., Mus musculus).
- NCBI Taxon ID: This field will auto-populate based on the scientific name.
- Click “Add Organism Information” to include multiple organisms, if required.
1.7. Declaration
- Check the confirmation box to acknowledge that you have read and agree to the Biotech-PRIDE Guidelines and FeED Protocols.
1.8. Submit Project
- Click “Register Project” to submit the project details.
- Upon successful submission, the project will be registered and assigned a unique project identifier.
Step 2 – Register Study
After successful project registration, users must register individual studies under the selected project. A study represents a specific experiment, observation, or phenotyping activity conducted on a defined organism.
Users are required to provide the following study details:
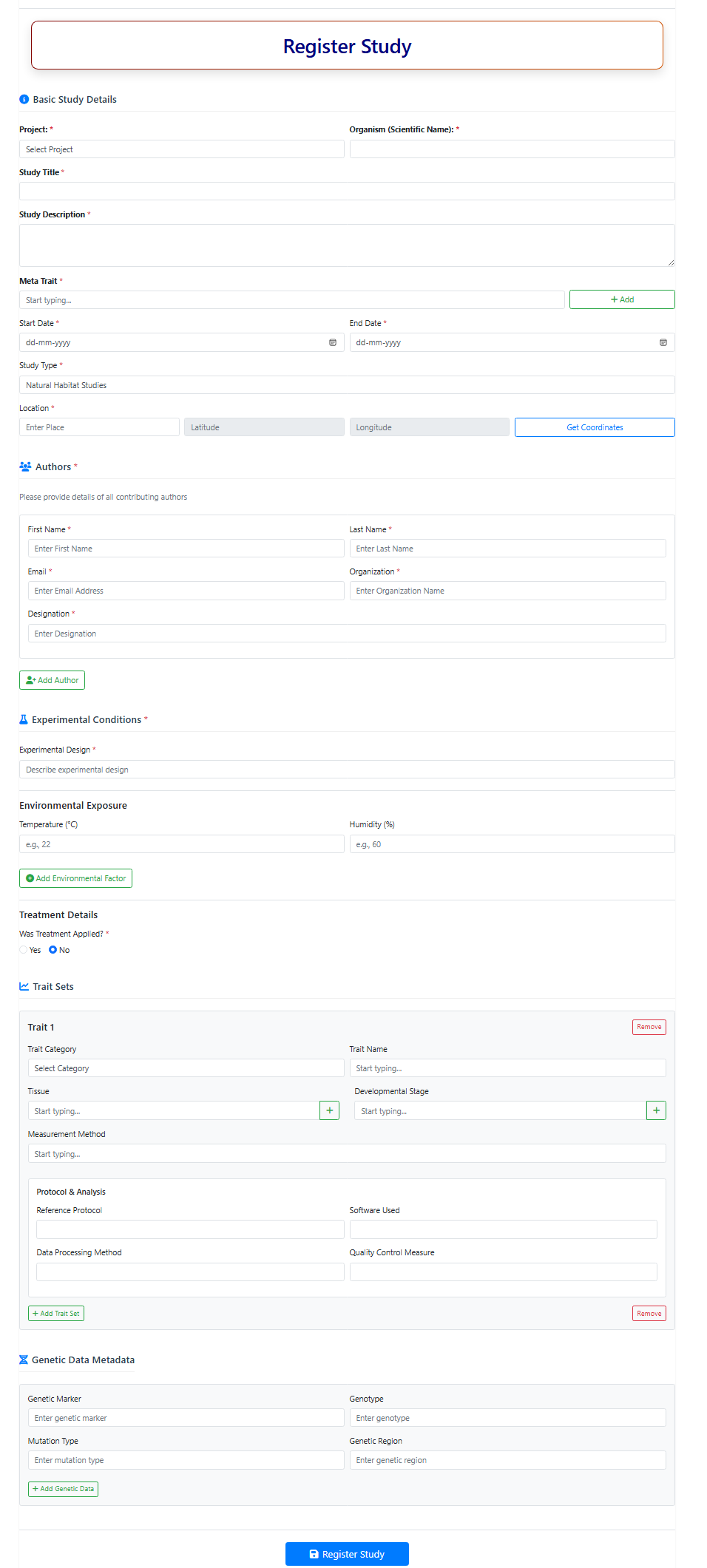
Figure 10. Snapshot of Registration of Study page in IAPD.
2.1. Basic Study Details
-
Project:
Select the relevant registered project from the dropdown list. -
Organism (Scientific Name):
Select the scientific name of the organism used in the study from the dropdown list. -
Study Title*:
Enter a concise and descriptive title for the study. -
Study Description*:
Provide a detailed description of the study, including objectives, experimental design, conditions, and relevance.
2.2. Meta Trait Selection
-
Meta Trait:
Select the meta-trait under which traits are recorded. Meta-traits provide high-level categorization (e.g., growth, reproduction, metabolism) and are mapped to Animal Trait Ontology terms. -
Create Meta Trait:
If not available, click ADD to create a new meta-trait by providing trait name, description, and organism. Newly created traits are added automatically. - Enter the project start and end date
- Select the study type from the dropdown
- Provide the study location and click Get Coordinates to auto-detect latitude and longitude
2.3. Authors
Provide details of all authors and contributors:
- First Name*: Author’s given name
- Last Name*: Author’s surname
- Email*: Valid email address
- Organization*: Affiliated institution
- Designation*: Author’s role or position
Click Add Author to include multiple contributors.
2.4. Experimental Conditions
-
Experimental Design*:
Describe controls, replicates, and overall study layout. -
Environmental Exposure:
- Temperature (°C)
- Humidity (%)
Click Add Environmental Factor to include additional conditions.
2.5. Treatment Details
This section captures metadata for any experimental treatment applied.
-
Was Treatment Applied?
- Select Yes if treatment was administered
- Select No for observational or control-only studies
Treatment Information
- Treatment Type*: Case or Control
- Treatment Agent*: Name of treatment (e.g., Dexamethasone)
- Treatment Group*: Treatment / Placebo / Control
- Description: Additional protocol details
Dosage and Duration
- Duration (Minutes)
- Duration (Hours)
- Dose / Concentration (with units)
Route of Administration
- Oral, Intraperitoneal, Intravenous, Topical, etc.
Users may add multiple treatments and define distinct treatment groups.
2.6. Trait Sets
Trait sets capture detailed phenotypic measurements and are mapped to controlled vocabularies (ATOL, Uberon, OBI).
- Trait Category
- Trait Name (ontology-based)
- Tissue (ontology-based or user-defined)
- Developmental Stage
- Measurement Method
- Reference Protocol (PMID / DOI)
- Software Used
- Data Processing Method
- Quality Control Measure
Click Add Trait Set to include more traits. Use Remove to delete unwanted trait sets.
2.7. Genetic Data Metadata (Optional)
- Genetic Marker
- Genotype
- Mutation Type (SNP, insertion, deletion)
- Genetic Region / Locus
Click Add Genetic Data to include multiple entries.
2.8. Completion of Study Registration
- Review all entered information carefully
- Ensure consistency with project-level metadata
- Click Register Study to proceed to the next submission step
Step 3: Sample Template Generation
Once a study has been successfully registered, users can generate and download a pre-filled data structured data submission template by clicking on the Step 3: Generate and Download Template button available on the User Dashboard under the Submit New Study section.
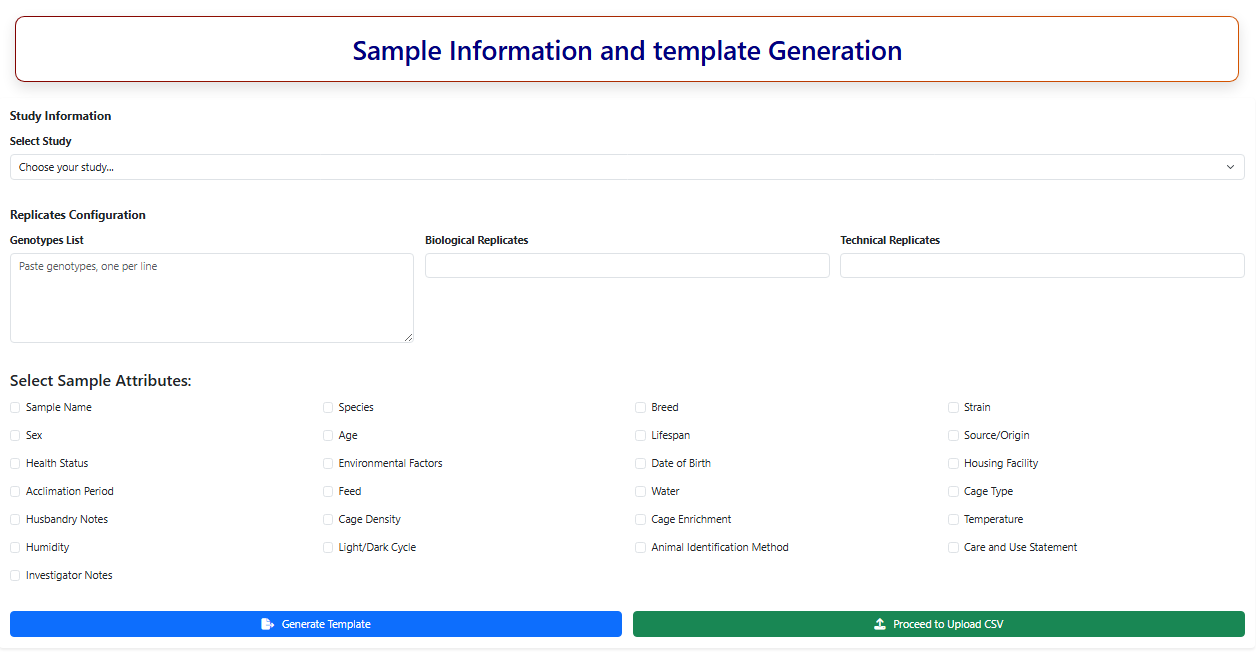
Figure 11. Snapshot of Samplate Template Gneration page of IAPD.
3.1. Study Information
-
Select Study:
Choose the registered study from the dropdown list. The selected study determines available traits, organism details, and validation rules for the sample template. -
Genotypes List:
Paste genotype identifiers for which the phenotypes were recorded, one per line (e.g., strain names, lines, or accessions). -
Biological Replicates:
Enter the number of biological replicates per genotype. -
Technical Replicates:
Enter the number of technical replicates per biological replicate.
3.2. Select Sample Attributes
Select the metadata fields to be included in the sample template by checking the required attributes.
Available sample attributes include (but are not limited to):
- Sample Identification: Sample Name, Animal Identification Method
- Biological Details: Species, Breed, Strain, Sex, Age, Lifespan
- Origin and Health: Source/Origin, Health Status
-
Environmental and Housing Conditions:
- Environmental Factors
- Temperature
- Humidity
- Housing Facility
- Cage Type
- Cage Density
- Cage Enrichment
-
Care and Management:
- Feed
- Water
- Acclimation Period
- Light/Dark Cycle
- Care and Use Statement
-
Administrative Notes:
- Investigator Notes
- Husbandry Notes
- Date of Birth
Select only relevant attributes to keep the template concise and accurate.
- After entering these details, click on the Generate Template CSV button. Users should carefully enter or paste the observation data into the corresponding observation columns, ensure data accuracy and consistency, and then save the completed file for subsequent upload in the next submission step.
Step 4: Upload CSV File
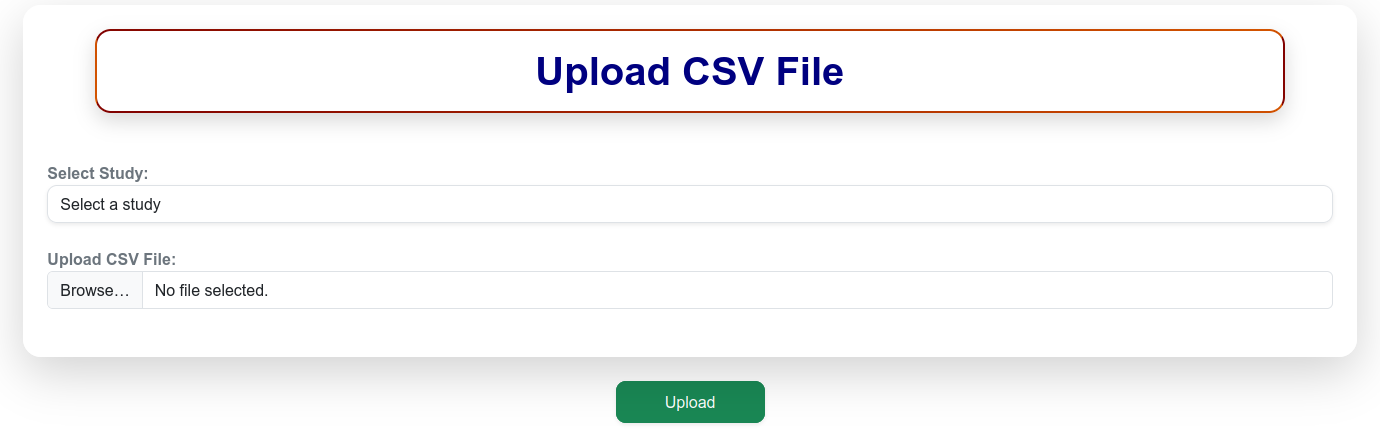
Figure 12. Snapshot of Upload CSV page of IAPD.
After completing the CSV file offline, click “Proceed to Upload CSV”. Select the study accession and browse the complete data file and click upload. Important Note: • Do not edit or change the format of the template file. • The accession list should be unique and no accession should be present in the list multiple times. • Do not leave any observation cell blank in the data file. Alternately use NA or ‘-‘instead of missing or blank values in your data. Upon successful upload, the newly submitted data are validated and updated instantly in the User Dashboard, allowing users to view the submission status and summary details in real time.
Step 5: Associated Data
If a study includes any associated or supplementary data that the user wishes to submit along with the primary phenotyping data, it can be uploaded in this step. To submit associated data, select the relevant Study ID, browse and choose the corresponding file, and click the Submit button to complete the upload. This step is optional and may be skipped without affecting the final completion of the data submission process. Users who do not have associated data to upload can proceed without submitting files in this step.
Figure 13. Snapshot of Upload associated data (if any) page of IAPD.
Browse Data
Users can browse the data available in the Indian Animal Phenome Database (IAPD) through the Browse interface. By default, the page displays all datasets registered in IAPD. However, only datasets categorized under Open Access are available for users to view and download. Data under managed access can be accessed via requesting through data access portal and data will be shared upon approval. Datasets under No Access are not accessible for viewing or download, in accordance with the IAPD Data Sharing Policy. Users can view detailed information for each project by clicking on the corresponding Project ID or Project Title. Selecting a project displays a list of all studies associated with that project, along with their summary details. On the left-hand side of the Browse page, users can refine their search results using multiple filter options, including: Date Range, Organism, Funding Agency, Data Access Type, Trait Category, and Study Type. These filters enable users to efficiently locate datasets relevant to their research interests.