| Project Accession: | IBIAP_1000000010 |
| Title: | Histopathological images of thyroid lesions |
| Representative Image: | 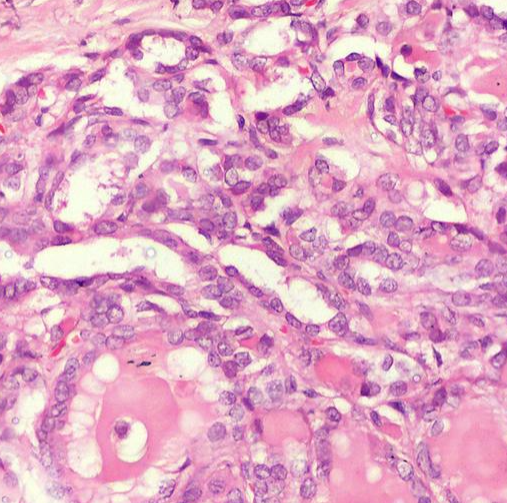 |
| Description: | The dataset comprises 154,498 images derived from 134 slides, representing 125 thyroid nodules from 118 patients. These images encompass six types of thyroid lesions: NIFTP, HTN, FA, IEFVPTC, IFSPTC, and CPTC. |
| Publications: | https://link.springer.com/article/10.1007/s12022-025-09865-0 |
| Associated Codes (URL only): | N/A |
| Funding agency: | DBT |
| Grant Number: | BT/PR34211/AI/133/1/2019 |
| Ethics Statement: | N/A |
| Any Other Information : | N/A |
| Additional File: | N/A |
| Acknowledgments: | This work was supported by the Department of Biotechnology (DBT) [grant numbers BT/PR34211/AI/133/1/2019 and BT/PR40151/BTIS/137/5/2021]. We would like to express our gratitude to the patients who participated in the study, as well as to the AIIMS and ICGEB for providing the necessary infrastructure for conducting the study. We acknowledge Chhaya, Ashish, Divya and Dhairya's crucial assistance in image QC data curation. |
| Sr.No | First name | Last name | Organization | Designation | |
|---|---|---|---|---|---|
| 1 | Shweta | Birla | sbirla84@gmail.com | International Centre for Genetic Engineering Biotechnology (ICGEB) | Postdoctoral Researcher |
| 2 | Nimisha | Tiwari | nimi.tiwari08@gmail.com | International Centre for Genetic Engineering Biotechnology (ICGEB) | Research Scholar |
| 3 | Pragati | Shyamal | pragatishyamal1@gmail.com | All India Institute of Medical Sciences (AIIMS) | Unspecified |
| 4 | Arundhati | Sharma | arundhatisharma1@gmail.com | All India Institute of Medical Sciences (AIIMS) | Co-Investigator |
| 5 | Shipra | Agarwal | drshipra0902@gmail.com | All India Institute of Medical Sciences (AIIMS) | Principal Investigator |
| 6 | Dinesh | Gupta | dinesh@icgeb.res.in | International Centre for Genetic Engineering Biotechnology (ICGEB) | Co-Principal Investigator |
| Study Accession: | HISTOS_1000000014 |
| Title: | Utility of Artificial Intelligence in differentiating Non- invasive Follicular Thyroid Neoplasm with Papillary like Nuclear Features from other follicular-patterned thyroid benign and malignant lesions |
| Imaging Type: | Histopathology (HISTO) |
| Imaging Sub-type: | Diagnostic Pathology |
| Summary: | The introduction of the term non-invasive follicular thyroid neoplasm with papillary-like nuclear features (NIFTP) in 2016 marked a pivotal shift in the classification of encapsulated follicular variants of papillary thyroid carcinoma (eFVPTC) lacking invasive features. While this reclassification significantly reduced overtreatment, the histopathological diagnosis of NIFTP remains challenging due to overlapping features with other thyroid lesions and inter-observer variability. This study presents a novel deep learning (DL)-based, three-stage diagnostic pipeline for distinguishing NIFTP from a wide spectrum of thyroid lesions, including benign and malignant mimics. By replicating the diagnostic strategy of histopathologists, the algorithm evaluates architectural patterns and nuclear features with high precision. Our approach has a potential to enhance diagnostic accuracy in a cost-effective and scalable manner, complementing existing diagnostic methods and thus optimizing clinical decision-making and improving the management of patients with thyroid neoplasms. |
| Keywords: | Histopathology; Deep-Learning; Differential Diagnosis; Thyroid Cancers |
| Additional / Any Other Information: | N/A |
| Release Date: | June 2, 2025 |
| Access Licence Type: | Open Access |
Table 1. The sample types registered under this study are as follows:
Table 2. The samples registered under this study are as follows:
| Sample Type ID | Organism | Taxon ID | Biological Entity | Laterality | Source Tissue | Source Cell/Cell-line | Cell Organelle |
|---|---|---|---|---|---|---|---|
| HISTOSMT_10000000038 | Homo sapiens | 9606 | Thyroid Gland | Not Applicable | Thyroid gland tissue | N/A | N/A |
Table 2. The samples registered under this study are as follows:
| Sample Type ID | Sample ID | Method used for Sample Collection | Cell Phenotype Studied | ICD-11 Code (patient health condition) | Presentation / Clinical Diagnosis |
|---|---|---|---|---|---|
| HISTOSMT_10000000038 | HISTOSM_10000227867 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227868 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227869 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227870 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227871 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227872 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227873 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227874 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227876 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227877 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227878 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227879 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227880 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227881 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227882 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227883 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227884 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227885 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227886 | N/A | N/A | XH0Q59 | IEFVPTC |
| HISTOSMT_10000000038 | HISTOSM_10000227887 | N/A | N/A | XH0Q59 | IEFVPTC |
Table 3. The experiment types registered under this study are as follows:
| Experiment Type ID | Instrument Name | Instrument Type | Manufacturer | Model |
|---|---|---|---|---|
| HISTOET_10000000015 | Digital Slide Scanning System | Digital Slide Scanner | Motic | VM1 |
| Experimental Design Summary (HISTOET_10000000015) |
|---|
| Hematoxylin and Eosin (H&E)-stained slides of histopathologically-confirmed cases of FND, HTN,FA, NIFTP, IEFVPTC, IFSPTC, and CPTC patients were retrieved from the archives of the Department of Pathology, All India Institute of Medical Sciences (AIIMS), New Delhi, India, spanning the period from 2015 to 2023. The corresponding slides were further reviewed by an experienced endocrine pathologist to reconfirm the diagnosis and select representative slides for further analysis. The diagnosis adhered to the criteria outlined in the most recent WHO classification of thyroid neoplasms The selected H&E slides were scanned at 40x magnification (equivalent to 0.25 μm/pixel) using the Motic VM1 whole slide scanner to generate whole slide images (WSIs). The slides were than subjected to AI studies. |
| Acquired Images Annotation Description (HISTOET_10000000015) |
|---|
| WHO defining criteria for the various thyroid lesions included in the study. For more details, please refer to "Additional File" of the "Project" section for this record. |
Table 4. The experiments registered under this study are as follows:
| Sample ID | Experiment Type ID | Experiment ID | Image type (Original / Derived / Unknown) | Any Other Information | Staining Type | Images Magnification | Tissue / Tumor Fixative Used | Dataset Split Type (Training / Validation / Test) |
|---|---|---|---|---|---|---|---|---|
| HISTOSM_10000225494 | HISTOET_10000000015 | HISTOE_10000198510 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000225495 | HISTOET_10000000015 | HISTOE_10000198511 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000225496 | HISTOET_10000000015 | HISTOE_10000198512 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000225497 | HISTOET_10000000015 | HISTOE_10000198513 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000225498 | HISTOET_10000000015 | HISTOE_10000198514 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000225499 | HISTOET_10000000015 | HISTOE_10000198515 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000225500 | HISTOET_10000000015 | HISTOE_10000198516 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218657 | HISTOET_10000000015 | HISTOE_10000185612 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218669 | HISTOET_10000000015 | HISTOE_10000185624 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218772 | HISTOET_10000000015 | HISTOE_10000185631 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218784 | HISTOET_10000000015 | HISTOE_10000185643 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218791 | HISTOET_10000000015 | HISTOE_10000185650 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218803 | HISTOET_10000000015 | HISTOE_10000185662 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218810 | HISTOET_10000000015 | HISTOE_10000185669 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218822 | HISTOET_10000000015 | HISTOE_10000185681 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218829 | HISTOET_10000000015 | HISTOE_10000185688 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218905 | HISTOET_10000000015 | HISTOE_10000185700 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218912 | HISTOET_10000000015 | HISTOE_10000185707 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218924 | HISTOET_10000000015 | HISTOE_10000185719 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| HISTOSM_10000218931 | HISTOET_10000000015 | HISTOE_10000185726 | Derived | N/A | Hematoxylin and Eosin (H&E) | 40x | 10% Neutral buffered formalin | Training Dataset |
| Experiment ID | Image File Name (with path) | Image Preview | Image Size |
|---|---|---|---|
| HISTOE_10000129555 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_740.tiff |  Download Image |
40K |
| HISTOE_10000129556 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_741.tiff |  Download Image |
48K |
| HISTOE_10000129557 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_742.tiff |  Download Image |
44K |
| HISTOE_10000129558 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_743.tiff |  Download Image |
48K |
| HISTOE_10000129559 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_744.tiff |  Download Image |
40K |
| HISTOE_10000129560 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_745.tiff |  Download Image |
48K |
| HISTOE_10000129561 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_746.tiff |  Download Image |
48K |
| HISTOE_10000129562 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_747.tiff |  Download Image |
52K |
| HISTOE_10000129563 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_748.tiff |  Download Image |
48K |
| HISTOE_10000129564 | thyroid_lesions/STAGE2_STID/ET_STID/ET_NPTCLF/ET_NPTCLF_749.tiff |  Download Image |
56K |