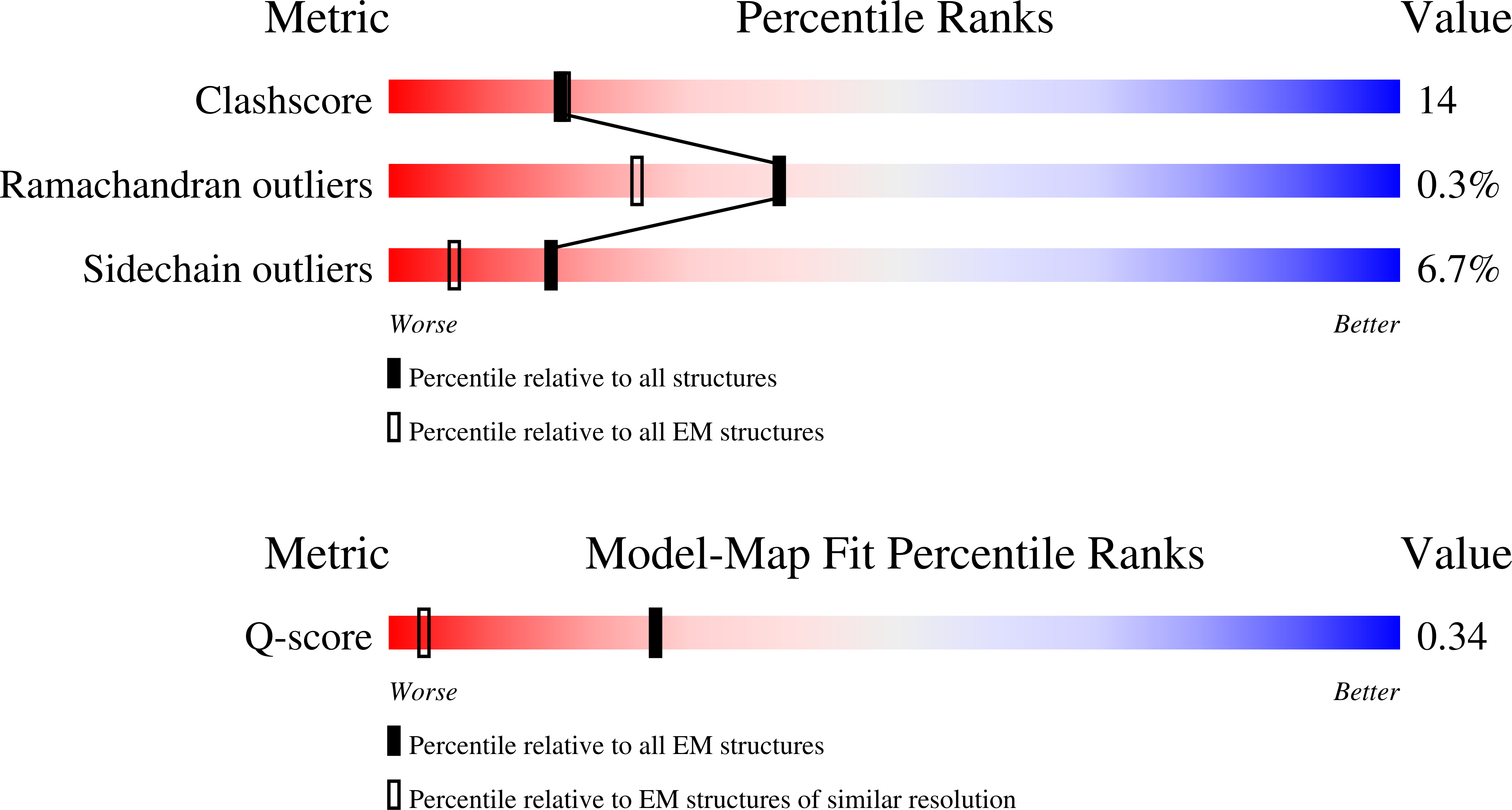
Deposition Date
2024-08-16
Release Date
2026-02-04
Last Version Date
2026-03-18
Entry Detail
PDB ID:
9D72
Keywords:
Title:
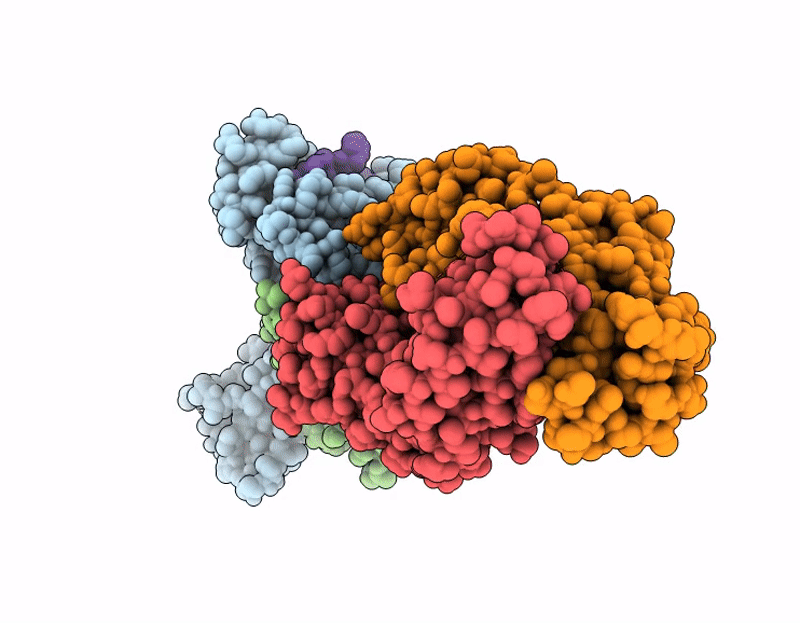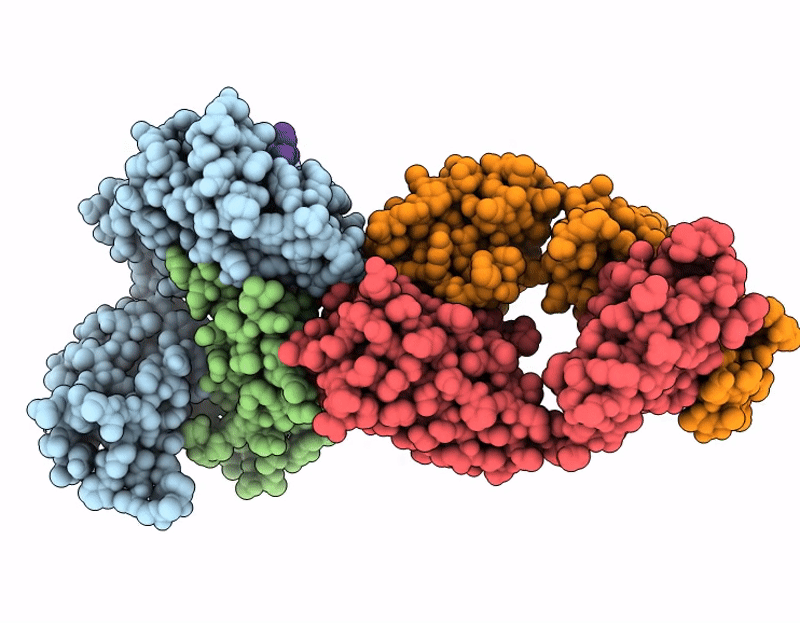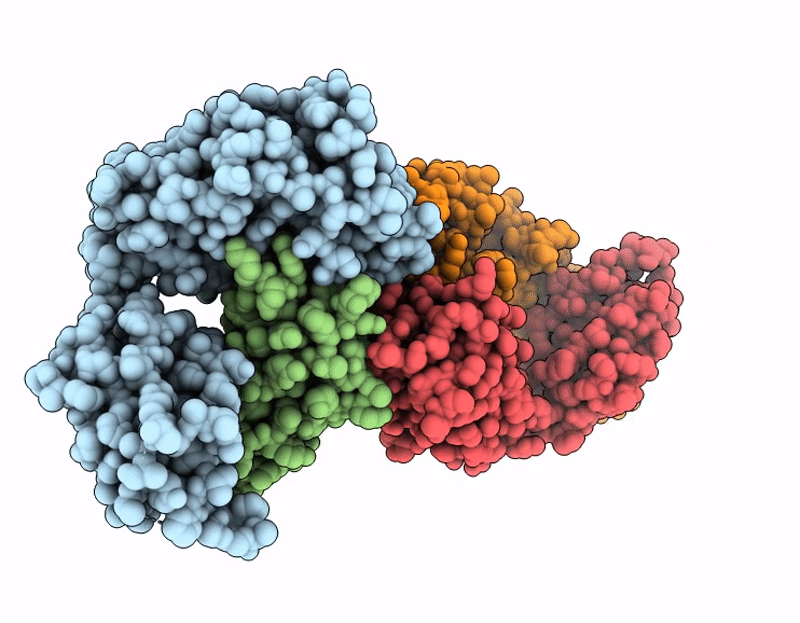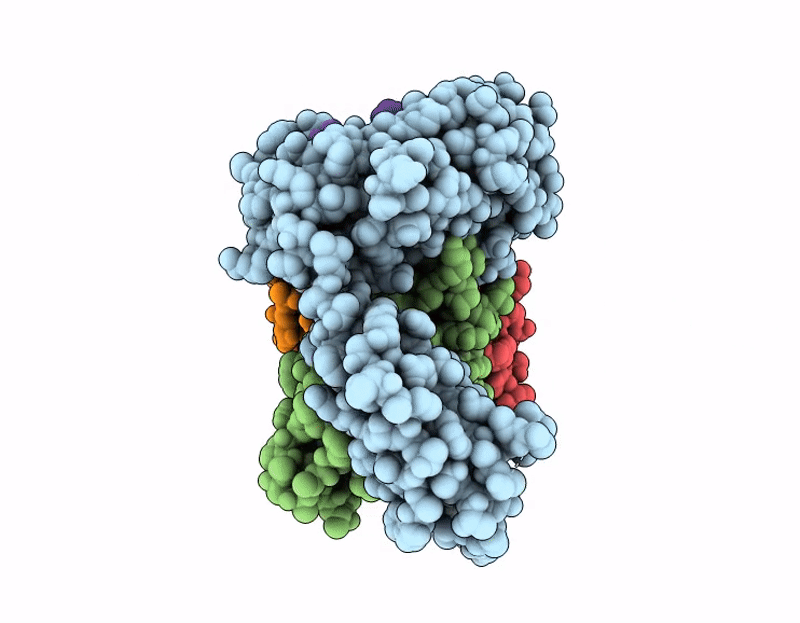
CryoEM structure of anti-MHC-I Fab M1/42 complex with H2-Dd
Biological Source:
Source Organism(s):
Mus musculus (Taxon ID: 10090)
Rattus norvegicus (Taxon ID: 10116)
synthetic construct (Taxon ID: 32630)
Rattus norvegicus (Taxon ID: 10116)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.67 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE