Deposition Date
2013-07-18
Release Date
2013-10-09
Last Version Date
2024-06-19
Entry Detail
PDB ID:
4BY9
Keywords:
Title:
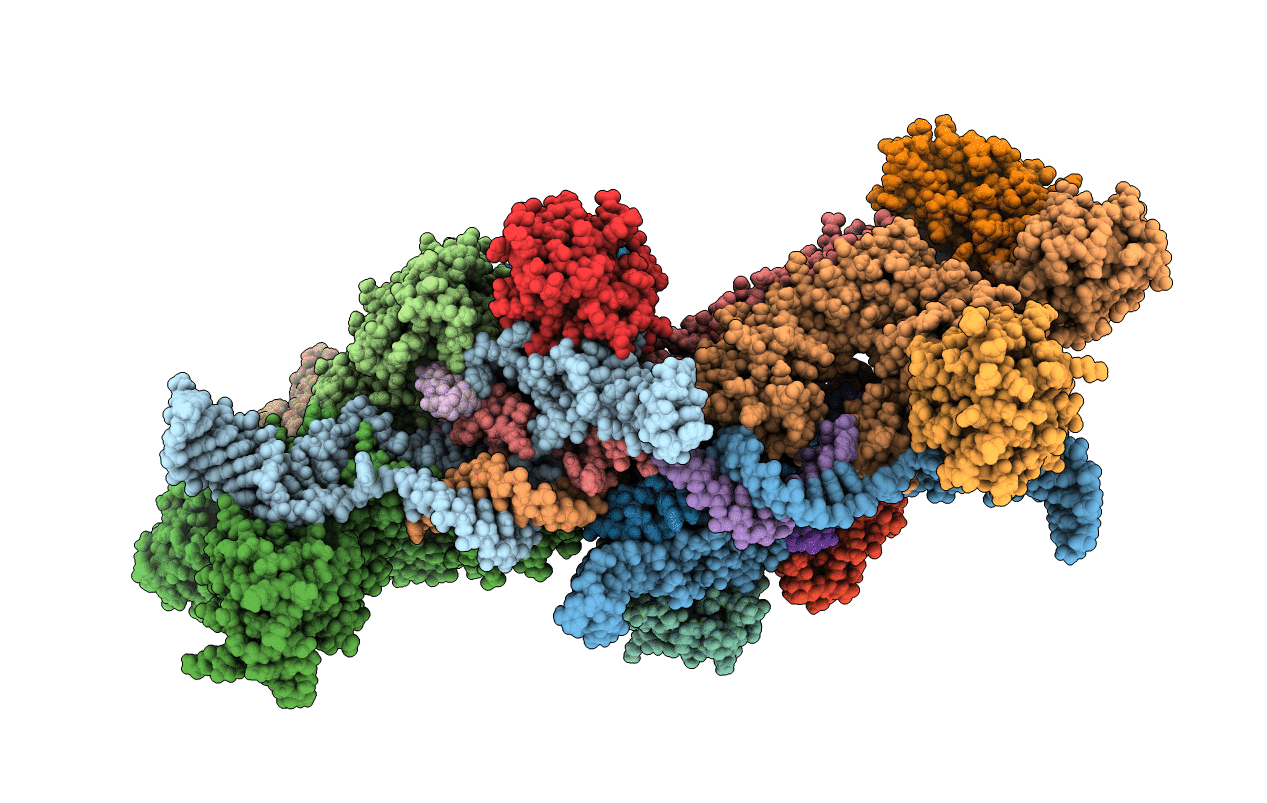
The structure of the Box CD enzyme reveals regulation of rRNA methylation
Biological Source:
Source Organism(s):
PYROCOCCUS FURIOSUS (Taxon ID: 2261)
Expression System(s):
Method Details:
Experimental Method:
Conformers Submitted:
1