Search Count: 726
All
Selected
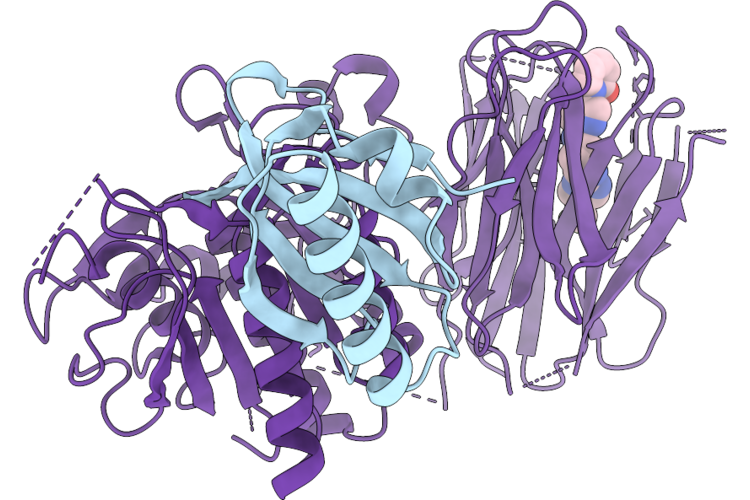 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-05-27 Classification: PROTEIN BINDING Ligands: A1IYR |
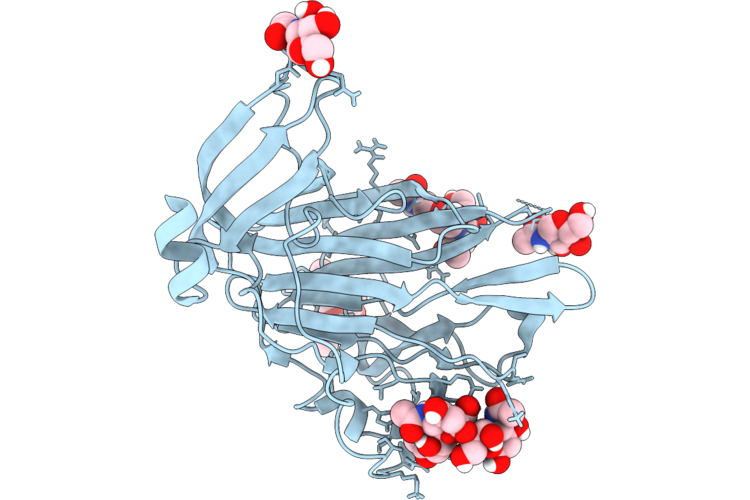 |
Cryo-Em Structure Of Hcov-Oc43-Lab Spike Glycoprotein In Complex With 9O-Acetyl Gd3 Sialoglycan (D1 Domain Local Refine)
Organism: Human coronavirus oc43
Method: ELECTRON MICROSCOPY Resolution:1.70 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: NAG, A1AR1 |
 |
Cryo-Em Structure Of Hcov-Oc43-Lab Spike Glycoprotein In Complex With 9O-Acetyl Gd3 Sialoglycan
Organism: Human coronavirus oc43
Method: ELECTRON MICROSCOPY Resolution:1.68 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: NAG, A1AR1 |
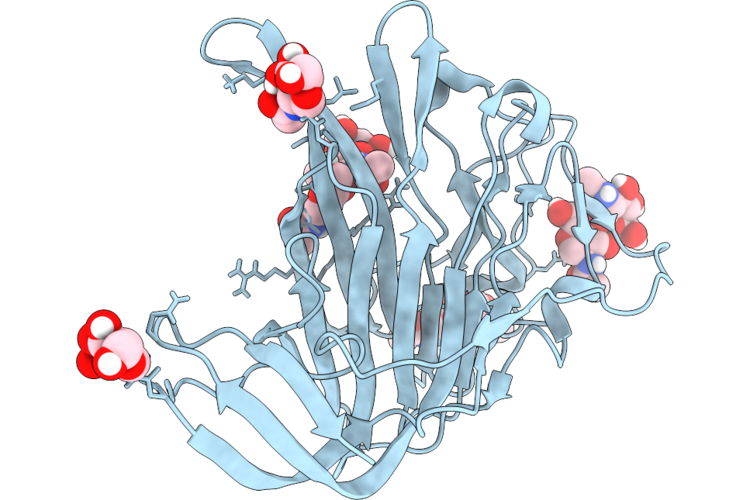 |
Cryo-Em Structure Of Hcov-Oc43-C2 Spike Glycoprotein (Local Refined D1 Domain)
Organism: Human coronavirus oc43
Method: ELECTRON MICROSCOPY Resolution:2.05 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Human coronavirus oc43
Method: ELECTRON MICROSCOPY Resolution:1.93 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Hcov-Oc43-C2 Spike Glycoprotein In Complex With 9O-Acetyl Gd3 Sialoglycan (Local Refined D1 Domain)
Organism: Human coronavirus oc43
Method: ELECTRON MICROSCOPY Resolution:1.98 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: NAG, A1AR1 |
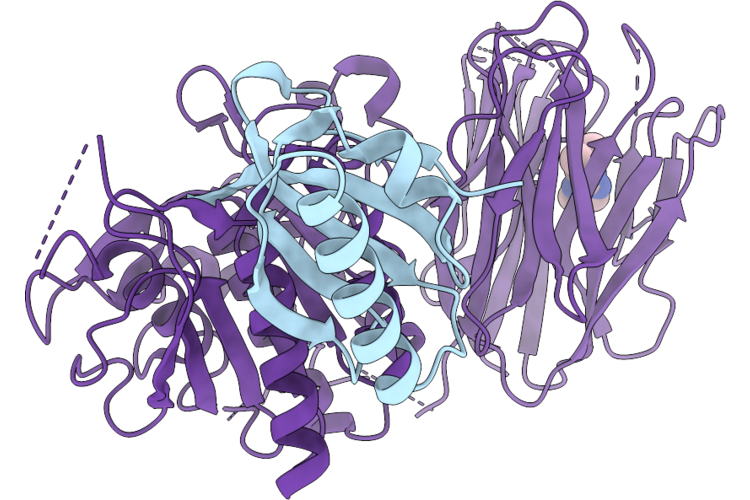 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-05-27 Classification: CHAPERONE Ligands: A1JSM |
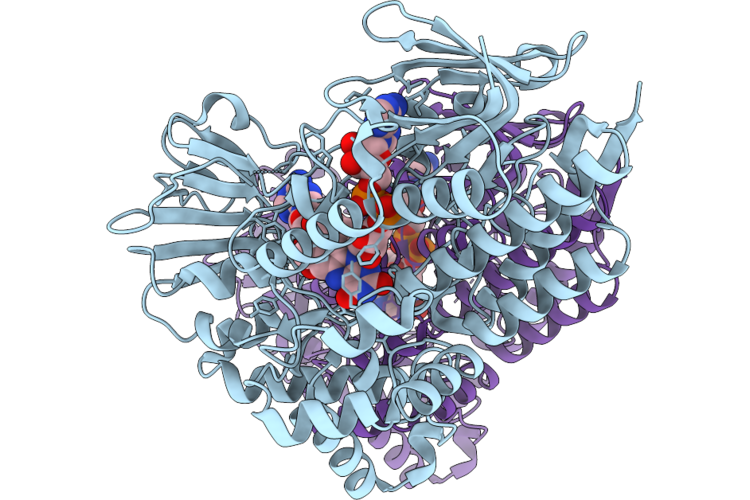 |
Structure Of Cyclohexanone Monooxygenase Mutant From Acinetobacter Calcoaceticus
Organism: Acinetobacter calcoaceticus
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-05-27 Classification: OXIDOREDUCTASE Ligands: FAD, NAP |
 |
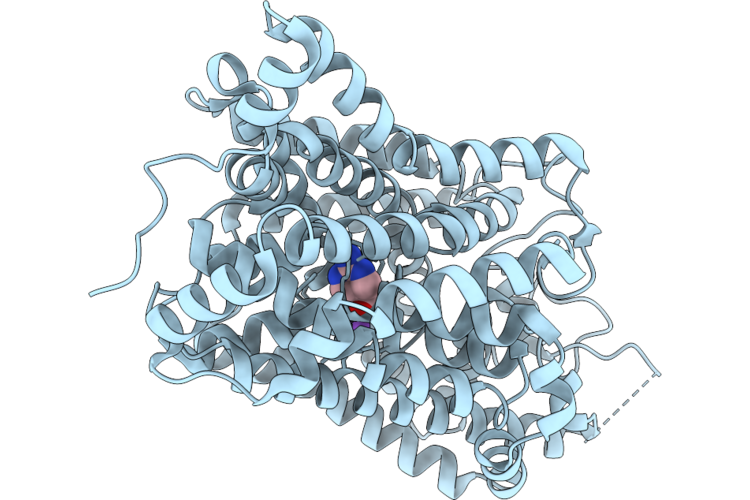
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: A1EVN, CL |
 |
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: A1EMK, CL, NA |
 |
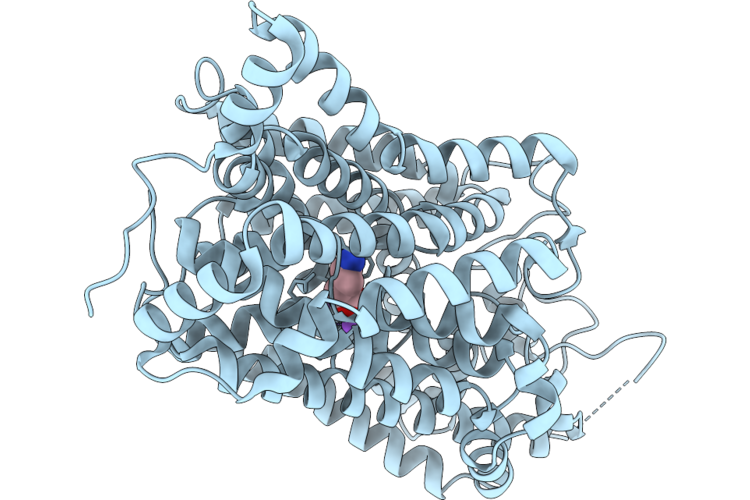
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: BET, NA, CL |
 |
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: ABU, CL, NA |
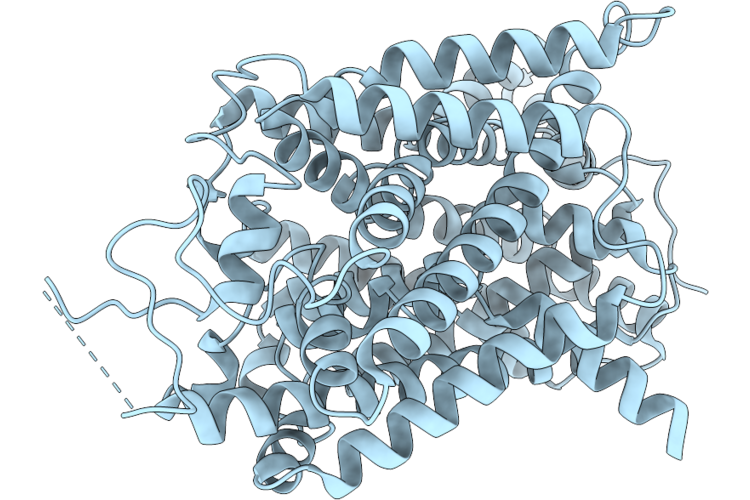 |
Structure Of The Apo State Of Human Betaine/Gaba Transporter 1 In The Inward-Facing Conformation
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: CL |
 |
Structure Of The Apo State Of Human Betaine/Gaba Transporter 1 In The Occluded Conformation
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:3.02 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
 |
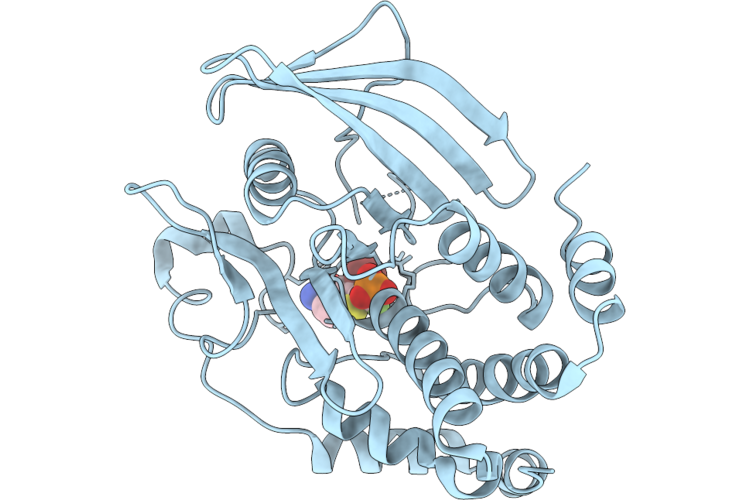
Structure Of Ptp1B Complexed With Difluoromethylphosphonate Inhibitor Compound 2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-05-27 Classification: HYDROLASE/INHIBITOR Ligands: A1C3A |
 |
Structure Of Ptp1B Complexed With Difluoromethylphosphonate Inhibitor Compound 10
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-05-27 Classification: HYDROLASE/INHIBITOR Ligands: A1C3B |
 |
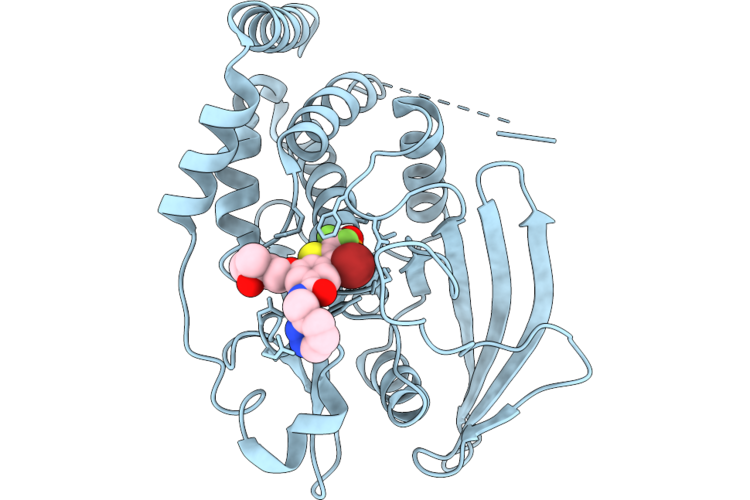
Structure Of Ptp1B Complexed With Difluoromethylphosphonate Inhibitor Compound 15
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-05-27 Classification: HYDROLASE/INHIBITOR Ligands: A1C3C, DMS |
 |
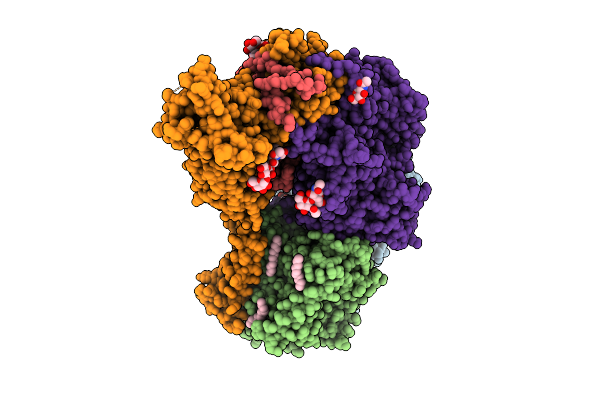
Structure Of Ptp1B Complexed With Difluoromethylphosphonate Inhibitor Compound 30
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2026-05-27 Classification: HYDROLASE/INHIBITOR Ligands: A1C3D |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: Y01, XKP, C14, D12, D10, A1EOT, NAG |
 |
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |

